Abstract
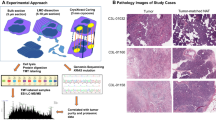
Pure populations of tumor cells are essential for the identification of tumor-associated proteins for the development of targeted therapy. In recent years, laser capture microdissection (LCM) has been used successfully to obtain distinct populations of cells for subsequent molecular analysis. The polycomb group (PcG) protein, enhancer of zeste homolog 2 (EzH2), a methyl-transferase that plays a key role in transcriptional gene repression, is frequently overexpressed in several malignant tumors. High levels of EzH2 are often associated with advanced disease stage in many solid tumors; however, its role in the pathogenesis of pancreatic ductal adeno-carcinoma (PDAC) is poorly understood. Because of the limited sample availability and the absence of in vitro amplification steps for proteins, the use of LCM for proteomics studies largely depends on highly sensitive protein detection methods. Here, we developed a faster and sensitive Western blot protocol and validated it for the detection of EzH2 in ∼2,000 cells. Initially, cultured PANC-1 cells were used to optimize protein electrophoresis and western blotting conditions. Gradient gel electrophoresis in combination with optimized antibody concentrations, and a sensitive chemiluminescent assay provided a strong signal. In order to further confirm the role of EzH2 in PDAC, employing siRNA-mediated gene silencing via long lasting plasmid vectors containing shRNA, we investigated the potential role of EzH2 gene silencing in pancreatic cancer regression. Positive correlation of EzH2 expression was observed with advanced stage, serous histology, and increasing grade in pancreatic cancer patient tissues. Further EzH2 knockdown resulted in decreased cell growth and invasiveness. The findings of this study emphasize that western blotting of a LCM-generated pure population of cancer cells may be a valuable technique for the study of tumor-specific proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Emmert-Buck, M., R. Bonner, et al. (1996). “Laser capture microdissection.” Science 274: 998.
Fend, F. and M. Raffeld (2000). “Laser capture microdissection in pathology.” Journal of Clinical Pathology 53: 666.
Banks, R., M. Dunn, et al. (1999). “The potential use of laser capture microdissection to selectively obtain distinct populations of cells for proteomic analysis-preliminary findings.” Electrophoresis 20: 689–700.
Espina, V., J. Wulfkuhle, et al. (2006). “Laser-capture microdissection.” Nature Protocols 1: 586–603.
Simone, N., A. Remaley, et al. (2000). “Sensitive immunoassay of tissue cell proteins procured by laser capture microdissection.” American Journal of Pathology 156: 445.
Craven, R. and R. Banks (2001). “Laser capture microdissection and proteomics: possibilities and limitation.” Proteomics 1: 1200–1204.
Von Eggeling, F., H. Davies, et al. (2000). “Tissue-specific microdissection coupled with ProteinChipÆ array technologies: applications in cancer research.” Biotechniques 29: 1066–1070.
Cao, R., L. Wang, et al. (2002). “Role of histone H3 lysine 27 methylation in Polycomb-group silencing.” Science 298: 1039.
Kirmizis, A., S. Bartley, et al. (2004). “Silencing of human polycomb target genes is associated with methylation of histone H3 Lys 27.” Genes & Development 18: 1592.
Kleer, C. G., Q. Cao, et al. (2003). “EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells.” Proceedings of the National Academy of Sciences of the United States of America 100: 11606–11611.
Bachmann, I. M., O. J. Halvorsen, et al. (2006). “EZH2 expression is associated with high proliferation rate and aggressive tumor subgroups in cutaneous melanoma and cancers of the endometrium, prostate, and breast.” Journal of Clinical Oncology 24: 268–273.
Balch, C., P. Yan, et al. (2005). “Antimitogenic and chemosensitizing effects of the methylation inhibitor zebularine in ovarian cancer.” Molecular Cancer Therapeutics 4: 1505.
Ohyama, H., X. Zhang, et al. (2000). “Laser capture microdissection-generated target sample for high-density oligonucleotide array hybridization.” Biotechniques 29: 530–536.
Hoffmann, M., J. Gaikwad, et al. (2002). “Analysis of odontoblast gene expression using a novel approach, laser capture microdissection.” Connect Tissue Res 43(2–3): 376–380.
Bonner, R. F., M. Emmert-Buck, et al. (1997). “Laser capture microdissection: molecular analysis of tissue.” Science 278: 1481,1483.
Schutze, K., H. Posl, et al. (1998). “Laser micromanipulation systems as universal tools in cellular and molecular biology and in medicine.” Cell Mol Biol (Noisy-le-grand) 44: 735–746.
Fend, F., M. R. Emmert-Buck, et al. (1999). “Immuno-LCM: laser capture microdissection of immunostained frozen sections for mRNA analysis.” Am J Pathol 154: 61–66.
Namimatsu, S., M. Ghazizadeh, et al. (2005). “Reversing the effects of formalin fixation with citraconic anhydride and heat: a universal antigen retrieval method.” J Histochem Cytochem 53: 3–11.
Uneyama, C., M. Shibutani, et al. (2002). “Methacarn fixation for genomic DNA analysis in microdissected, paraffin-embedded tissue specimens.” J Histochem Cytochem 50: 1237–1245.
Friedl, P. and K. Wolf (2003). “Tumour-cell invasion and migration: diversity and escape mechanisms.” Nature Reviews Cancer 3: 362–374.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Qazi, A.M. et al. (2011). Laser Capture Microdissection of Pancreatic Ductal Adeno-Carcinoma Cells to Analyze EzH2 by Western Blot Analysis. In: Murray, G. (eds) Laser Capture Microdissection. Methods in Molecular Biology, vol 755. Humana Press. https://doi.org/10.1007/978-1-61779-163-5_20
Download citation
DOI: https://doi.org/10.1007/978-1-61779-163-5_20
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-162-8
Online ISBN: 978-1-61779-163-5
eBook Packages: Springer Protocols