Abstract
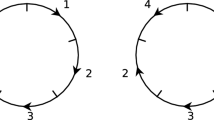
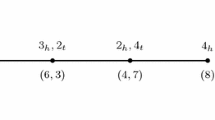
The Double Cut and Join is an operation acting locally at four chromosomal positions without regard to chromosomal context. This chapter discusses its application and the resulting menu of operations for genomes consisting of arbitrary numbers of circular chromosomes, as well as for a general mix of linear and circular chromosomes. In the general case the menu includes: inversion, translocation, transposition, formation and absorption of circular intermediates, conversion between linear and circular chromosomes, block interchange, fission, and fusion. This chapter discusses the well-known edge graph and its dual, the adjacency graph, recently introduced by Bergeron et al. Step-by-step procedures are given for constructing and manipulating these graphs. Simple algorithms are given in the adjacency graph for computing the minimal DCJ distance between two genomes and finding a minimal sorting; and use of an online tool (Mauve) to generate synteny blocks and apply DCJ is described.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Nadeau, J. H., Taylor, B. A. (1984) Lengths of chromosomal segments conserved since divergence of man and mouse. Proc Natl Acad Sci U S A 81, 814–818.
Pevzner, P. A. (2000) Computational Molecular Biology: An Algorithmic Approach. MIT Press, Cambridge, MA.
Yancopoulos, S., Attie, O., Friedberg, R. (2005) Efficient sorting of genomic permutations by translocation, inversion and block interchange. Bioinformatics 21, 3340–3346
Bergeron, A., Mixtacki, J., Stoye, J. (2006) A unifying view of genome rearrangements. WABI 2006, 163–173.
Christie, D. A. (1996) Sorting permutations by block interchanges. Inform Proc Lett 60,165–169.
Lin, Y. C., et al. (2005) An efficient algorithm for sorting by block-interchanges and its application to the evolution of Vibrio species. J Comput Biol 12, 102112.
Meidanis, J., Dias, Z. (2001) Genome rearrangements distance by fusion, fission, and transposition is easy. In Proceedings of SPIRE'2001—String Processing and Information Retrieval, Laguna de San Rafael, Chile.
Kececioglu, J., Sankoff, D. (1995) Exact and approximation algorithms for sorting by reversals, with application to genome rearrangement. Algorithmica 13, 180–210.
Bergeron, A., private communication.
Bafna, V., Pevzner, P.A. (1993) Genome rearrangements and sorting by reversals. In Proceedings of the 34th Annual IEEE Symposium on Foundations of Computer Science, IEEE Press, Los Alamitos, CA.
Hannenhalli, S., Pevzner, P. A. (1995) Transforming men into mice (polynomial algorithm for genomic distance problem). In Proceedings of the 36th Annual IEEE Symposium on Foundations of Computer Science, Milwaukee, WI.
Darling, A. C. E., Mau, B., Blattner, F. R., et al. (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14, 1394–1403.
Darling, A. E., Treangen, T. J., Zhang, L., et al. (2006) Procrastination leads to efficient filtration for local multiple alignment. Lecture Notes in Bioinformatics. Springer-Verlag, New York.
Darling, A. E., Treangen, T. J., Messeguer, X., et al. (2007) Analyzing patterns of microbial evolution using the Mauve genome alignment system, in (Bergman, ed.), Comparative Genomics. Humana Press, Totowa, NJ, in press.
Acknowledgments
The authors are grateful to David Sankoff for his advice and encouragement; Anne Bergeron for communicating the idea of the adjacency graph in advance of publication; Mike Tsai for his online implementation of the DCJ; and Betty Harris for invaluable logistic support and encouragement. S.Y. thanks Nicholas Chiorazzi for his enthusiasm, encouragement, and support; and A.E.D is supported by NSF grant DBI-0630765.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Friedberg, R., Darling, A.E., Yancopoulos, S. (2008). Genome Rearrangement by the Double Cut and Join Operation. In: Keith, J.M. (eds) Bioinformatics. Methods in Molecular Biology™, vol 452. Humana Press. https://doi.org/10.1007/978-1-60327-159-2_18
Download citation
DOI: https://doi.org/10.1007/978-1-60327-159-2_18
Publisher Name: Humana Press
Print ISBN: 978-1-58829-707-5
Online ISBN: 978-1-60327-159-2
eBook Packages: Springer Protocols