Abstract
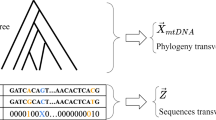
This chapter introduces the problem of ancestral sequence reconstruction: given a set of extant orthologous DNA genomic sequences (or even whole-genomes), together with a phylogenetic tree relating these sequences, predict the DNA sequence of all ancestral species in the tree. Blanchette et al. (1) have shown that for certain sets of species (in particular, for eutherian mammals), very accurate reconstruction can be obtained. We explain the main steps involved in this process, including multiple sequence alignment, insertion and deletion inference, substitution inference, and gene arrangement inference. We also describe a simulation-based procedure to assess the accuracy of the reconstructed sequences. The whole reconstruction process is illustrated using a set of mammalian sequences from the CFTR region.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Blanchette, M., Green, E. D., Webb, M., and Haussler, D. (2004) Reconstructing large regions of an ancestral mammalian genome in silico. Genome Res. 14, 2412–2423.
International Human Genome Sequencing Consortium, Lander, E., et al. (2001) Initial sequencing and analysis of the human genome. Nature 5, 409(6822), 860–921 (PMID: 12466850).
International Mouse Genome Sequencing Consortium, Waterston, R. H., Lindblad-Toh, K., Birney, E., et al. (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 5, 420(6915), 520–562 (PMID: 12466850).
Rat Genome Sequencing Project Consortium, Gibbs, R. A., Weinstock, G. M., Metzker, M. L., et al. (2004) Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521.
Margulies, E. H., Blanchette, M., NISC Comparative Sequencing Program, Haussler, D., and Green, E. (2003) Identification and characterization of multi-species conserved sequences. Genome Res. 13(12), 2507–2518 (PMID: 14656959).
Cooper, G. M., Brudno, M., Green, E. D., Batzoglou, S., and Sidow, A. (2003) Quantitative estimates of sequence divergence for comparative analyses of mammalian genomes. Genome Res. 13(5), 813–820.
Bejerano, G., Pheasant, M., Makunin, I., et al. (2004) Ultraconserved elements in the human genome. Science 304(5675), 1321–1325.
Goodman, M., Barnabas, J., Matsuda, G., and Moore, G. W. (1971) Molecular evolution in the descent of man. Nature 233, 604–613.
Enard, W., Przeworski, M., Fisher, S. E., et al. (2002) Molecular evolution of FOXP2, a gene involved in speech and language. Nature 418(6900), 869–872.
Eizirik, E., Murphy, W. J., and O’Brien, S. J. (2001) Molecular dating and biogeography of the early placental mammal radiation. J. Hered. 92(2), 212–219 (PMID: 11396581).
Springer, M. S., Murphy, W. J., Eizirik, E., and O’Brien, S. J. (2003). Placental mammal diversification and the Cretaceous-Tertiary boundary. Proc. Natl Acad. Sci. U. S A 4, 100(3), 1056–1060 (PMID: 12552136).
Thomas, J., Touchman, J. W., Blakesley, R. W., et al. (2003) Comparative analyses of multi-species sequences from targeted genomic regions. Nature 424, 788–793.
Karolchick, D., Baertsch, R., Diekhans, M., et al. (2003) The UCSC genome browser database. Nucleic Acids Res. 31, 51–54.
Maddison, D. R. and Schulz K.-S. (ed.) (2004) The Tree of Life Web Project. http://tolweb.org
Felsenstein, J. (1989) PHYLIP-Phylogeny inference package (Version 3.2). Cladistics 5, 164–166.
Swofford, D. L. (2003) PAUP: Phylogenetic Analysis Using Parsimony. Sinauer, Sunderland, MA.
Huelsenbeck, J. P. and Ronquist, F. (2001) MrBayes: Bayesian inference of phylogeny. Bioinformatics 17, 754–755.
Bray, N. and Pachter, L. (2004) MAVID: constrained ancestral alignment of multiple sequences. Genome Res. 14, 693–699.
Cooper, G. M., Stone, E. A., Asimenos, G., et al. (2005) Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15(7), 901–913.
Blanchette, M., Kent, W. J., Riemer, C., et al. (2004) Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14(4), 708–715 (PMID: 15060014).
Schwartz, S., Kent, W. J., Smith, A., et al. (2003) Human-mouse alignments with BLASTZ. Genome Res. 13(1), 103–107.
Chindelevitch, L., Li, Z., Blais, E., and Blanchette, M. (2006) On the inference of parsimonious indel evolutionary scenarios. J. Bioinformatics Comput. Biol. in press.
Fredslund, J., Hein, J., and Scharling, T. (2003) A large version of the small parsimony problem. Lecture Notes in Bioinformatics, Proceedings of WABI’03. 2812, 417–432.
Yang, Z., Kumar, S., and Nei, M. (1995) A new method of inference of ancestral nucleotide and amino acid sequences. Genetics 141, 1641–1650.
Siepel, A. and Haussler, D. (2003) Combining phylogenetic and hidden Markov models in biosequence analysis. Proceedings of the 7th Annual International. Conference on Research in Computational Molecular Biology. pp. 277–286.
Bourque, G. and Pevzner, P. (2002) Genome-scale evolution: reconstructing gene orders in the ancestral species. Genome Res. 12(1), 26–36.
Stoye, J., Evers, D., and Meyer, F. (1997) Generating benchmarks for multiple sequence alignments and phylogenetic reconstructions. Proc. Int. Conf. Intell. Syst. Mol. Biol. 5, 303–204 (PMID: 9322053).
Hasegawa, M., Kishino, H., and Yano, T. (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 22(2), 160–174.
Kent, J., Baertsch, R., Hinrichs, A., Miller, W., and Haussler, D. (2003). Evolution’s cauldron: duplication, deletion and rearrangement in the mouse and human genomes, Proc. Natl Acad. Sci. USA 100(20), 11,848–11,489.
Jurka, J. (2002) Repbase update: a database and an electronic journal of repetitive elements. Trends Genet. 16(9), 418–420 (PMID: 10973072).
Smit, A. and Green, P. (1999) RepeatMasker, http://ftp.genome.washington.edu/RM/RepeatMasker.html
Hoeffding, W. (1963) Probability inequalities for sums of bounded random variables. J. Am. Stat. Assoc. 58, 13–27.
Le Cam, L. (1986) Asymptotic Methods in Statistical Decision Theory, Springer, New York.
Lucena, B. and Haussler, D. (2005) Counterexample to a claim about the reconstruction of an ancestral character states. Syst Biol. 54(4), 693–695.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Humana Press Inc., Totowa, NJ
About this protocol
Cite this protocol
Blanchette, M., Diallo, A.B., Green, E.D., Miller, W., Haussler, D. (2008). Computational Reconstruction of Ancestral DNA Sequences. In: Murphy, W.J. (eds) Phylogenomics. Methods in Molecular Biology™, vol 422. Humana Press. https://doi.org/10.1007/978-1-59745-581-7_11
Download citation
DOI: https://doi.org/10.1007/978-1-59745-581-7_11
Publisher Name: Humana Press
Print ISBN: 978-1-58829-764-8
Online ISBN: 978-1-59745-581-7
eBook Packages: Springer Protocols