Summary
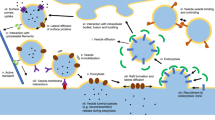
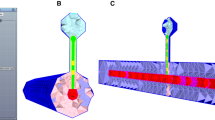
Spatially realistic diffusion-reaction simulations supplement traditional experiments and provide testable hypotheses for complex physiological systems. To date, however, the creation of realistic 3D cell models has been difficult and time-consuming, typically involving hand reconstruction from electron microscopic images. Here, we present a complementary approach that is much simpler and faster, because the cell architecture (geometry) is created directly in silico using 3D modeling software like that used for commercial film animations. We show how a freely available open source program (Blender) can be used to create the model geometry, which then can be read by our Monte Carlo simulation and visualization softwares (MCell and DReAMM, respectively). This new workflow allows rapid prototyping and development of realistic computational models, and thus should dramatically accelerate their use by a wide variety of computational and experimental investigators. Using two self-contained examples based on synaptic transmission, we illustrate the creation of 3D cellular geometry with Blender, addition of molecules, reactions, and other run-time conditions using MCell's Model Description Language (MDL), and subsequent MCell simulations and DReAMM visualizations. In the first example, we simulate calcium influx through voltage-gated channels localized on a presynaptic bouton, with subsequent intracellular calcium diffusion and binding to sites on synaptic vesicles. In the second example, we simulate neurotransmitter release from synaptic vesicles as they fuse with the presynaptic membrane, subsequent transmitter diffusion into the synaptic cleft, and binding to postsynaptic receptors on a dendritic spine.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Tao, L. and Nicholson, C. (2004) Maximum geometrical hindrance to diffusion in brain extracellular space surrounding uniformly spaced convex cells. J. Theor. Biol. 229, 59–68.
Hrabe, J., Hrabetova, S., and Segeth, K. (2004) A model of effective diffusion and tortuosity in the extracellular space of the brain. Biophys. J. 87, 1606–1617.
Tao, A., Tao, L., and Nicholson, C. (2005) Cell cavities increase tortuosity in brain extracellular space. J. Theor. Biol. 234, 525–536.
Stiles, J. R., Van Helden, D., Bartol, T. M., Jr., Salpeter, E. E., and Salpeter, M. M. (1996) Miniature endplate current rise times <100 μs from improved dual recordings can be modeled with passive acetylcholine diffusion from a synaptic vesicle. Proc. Natl. Acad. Sci. USA 93, 5747–5752.
Stiles, J. R., Bartol, T. M., Jr., Salpeter, E. E., and Salpeter, M. M. (1998) Monte Carlo simulation of neurotransmitter release using MCell, a general simulator of cellular physiological processes, in Computational Neuroscience (Bower, J. M., ed.), Plenum, New York, NY, pp. 279–284.
Stiles, J. R., Bartol, T. M., Salpeter, M. M., Salpeter, E. E., and Sejnowski, T. J. (2001) Synaptic variability: new insights from reconstructions and Monte Carlo simulations with MCell, in Synapses (Cowan, W. M., Stevens, C. F., and Sudhof, T. C., eds.), Johns Hopkins University Press, Baltimore, MD, pp. 681–731.
Stiles, J. R. and Bartol, T. M. (2001) Monte Carlo methods for simulating realistic synaptic microphysiology using MCell, in Computational Neuroscience: Realistic Modeling for Experimentalists (De Schutter, E., ed.), CRC Press, Boca Raton, FL, pp. 87–127.
Pawlu, C., DiAntonio, A., and Heckmann, M. (2004) Postfusional control of quantal current shape. Neuron 43, 607–618.
Stiles, J. R., Ford, W. C., Pattillo, J. M., Deerinck, T. E., Ellisman, M. H., Bartol, T. M., and Sejnowski, T. J. (2004) Spatially realistic computational physiology: past, present, and future, in Parallel Computing: Software Technology, Algorithms, Architectures & Applications (Joubert, G. R., Nagel, W. E., Peters, F. J., and Walter, W. V., eds.), Elsevier, Amsterdam, pp. 685–694.
Coggan, J. S., Bartol, T. M., Esquenazi, E., Stiles, J. R., Lamont, S., Martone, M. E., Berg, D. K., Ellisman, M. H., and Sejnowski, T. J. (2005) Evidence for ectopic neurotransmission at a neuronal synapse. Science 309, 446–451.
He, L., Wu, X. S., Mohan, R., and Wu, L. G. (2006) Two modes of fusion pore opening revealed by cell-attached recordings at a synapse. Nature 444, 102–105.
Kerr, R. A., Bartol, T. M., Kaminsky, B., Dittrich, M., Chang, J. J.-C., Baden, S. B., Sejnowski, T. J., and Stiles, J. R. Fast Monte Carlo simulation methods for biological reaction-diffusion systems in solution and on surfaces. SIAM J. Sci. Comput. 30, 3126–3149.
Kerr, R. A., Levine, H., Sejnowski, T. J., and Rappel, W. J. (2006) Division accuracy in a stochastic model of Min oscillations in Escherichia coli. Proc. Natl. Acad. Sci. USA 103, 347–352.
Koh, X., Srinivasan, B., Ching, H. S., and Levchenko, A. (2006) A 3D Monte Carlo analysis of the role of dyadic space geometry in spark generation. Biophys. J. 90, 1999–2014.
www.mcell.psc.edu and pages therein.
www.blender.org.
Synapse Web, Kristen M. Harris, PI, http://synapse-web.org/. The publicly available VRML file that contains the data for the reconstruction can be downloaded from www.synapse-web.org/anatomy/Ca1pyrmd/radiatum/index.stm.
Acknowledgments
The authors thank Aji Janis and Gary Blumenthal for careful reading and testing of the examples in this chapter. This work was supported by NIH R01 GM068630 (JRS), P41 RR06009 (JRS), and F32 GM083473 (MD).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press
About this protocol
Cite this protocol
Czech, J., Dittrich, M., Stiles, J. (2009). Rapid Creation, Monte Carlo Simulation, and Visualization of Realistic 3D Cell Models. In: Maly, I. (eds) Systems Biology. Methods in Molecular Biology, vol 500. Humana Press. https://doi.org/10.1007/978-1-59745-525-1_9
Download citation
DOI: https://doi.org/10.1007/978-1-59745-525-1_9
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-934115-64-0
Online ISBN: 978-1-59745-525-1
eBook Packages: Springer Protocols