Summary
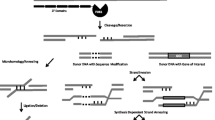
Zinc finger nucleases (ZFNs) are custom-designed molecular scissors, engineered to cut at specific DNA sequences. ZFNs combine the zinc finger proteins (ZFPs) with the nonspecific cleavage domain of the FokI restriction enzyme. The DNA-binding specificity of ZFNs can be easily altered experimentally. This easy manipulation of the ZFN recognition specificity enables one to deliver a targeted double-strand break (DSB) to a genome. The targeted DSB stimulates local gene targeting by several orders of magnitude at that specific cut site via homologous recombination (HR). Thus, ZFNs have become an important experimental tool to make site-specific and permanent alterations to genomes of not only plants and mammals but also of many other organisms. Engineering of custom ZFNs involves many steps. The first step is to identify a ZFN site at or near the chosen chromosomal target within the genome to which ZFNs will bind and cut. The second step is to design and/or select various ZFP combinations that will bind to the chosen target site with high specificity and affinity. The DNA coding sequence for the designed ZFPs are then assembled by polymerase chain reaction (PCR) using oligonucleotides. The third step is to fuse the ZFP constructs to the FokI cleavage domain. The ZFNs are then expressed as proteins by using the rabbit reticulocyte in vitro transcription/translation system and the protein products assayed for their DNA cleavage specificity.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kandavelou, K., M. Mani, S. Durai, and S. Chandrasegaran. (2005). “Magic” scissors for genome surgery. Nat Biotechnol 23:686–687.
Wu, J., K. Kandavelou, and S. Chandrasegaran. (2007). Custom-designed zinc finger nucleases: what is next? Cell Mol Life Sci 64:2933–2944.
Vasquez, K.M., K. Marburger, Z. Intody, and J.H. Wilson. (2001). Manipulating the mammalian genome by homologous recombination. Proc Natl Acad Sci U S A 98:8403–8410.
Kim, C.A., and J.M. Berg. (1996). A 2.2 Å resolution crystal structure of a designed zinc finger protein bound to DNA. Nat Struct Biol 3:940–945.
Pavletich, N.P., and C.O. Pabo. (1991). Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 Å. Science 252:809–817.
Desjarlais, J.R., and J.M. Berg. (1993). Use of a zinc-finger consensus sequence framework and specificity rules to design specific DNA binding proteins. Proc Natl Acad Sci U S A 90:2256–2260.
Shi, Y., and J.M. Berg. (1995). A direct comparison of the properties of natural and designed zinc-finger proteins. Chem Biol 2:83–89.
Beerli, R.R., D.J. Segal, B. Dreier, and C.F. Barbas, III. (1998). Toward controlling gene expression at will: specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc Natl Acad Sci U S A 95:14628–14633.
Kim, J.S., and C.O. Pabo. (1998). Getting a handhold on DNA: design of poly-zinc finger proteins with femtomolar dissociation constants. Proc Natl Acad Sci U S A 95:2812–2817.
Liu, Q., D.J. Segal, J.B. Ghiara, and C.F. Barbas, III. (1997). Design of polydactyl zinc-finger proteins for unique addressing within complex genomes. Proc Natl Acad Sci U S A 94:5525–5530.
Chandrasegaran, S., and J. Smith. (1999). Chimeric restriction enzymes: what is next? Biol Chem 380:841–848.
Kandavelou, K., Mani, M., Durai, S., and Chandrasegaran, S. (2004). Engineering and applications of chimeric nucleases. Springer, Berlin. pp. 413–414.
Kim, Y.G., J. Cha, and S. Chandrasegaran. (1996). Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci U S A 93:1156–1160.
Smith, J., J.M. Berg, and S. Chandrasegaran. (1999). A detailed study of the substrate specificity of a chimeric restriction enzyme. Nucleic Acids Res 27:674–681.
Porteus, M.H., and D. Baltimore. (2003). Chimeric nucleases stimulate gene targeting in human cells. Science 300:763.
Urnov, F.D., J.C. Miller, Y.L. Lee, C.M. Beausejour, J.M. Rock, S. Augustus, A.C. Jamieson, M.H. Porteus, P.D. Gregory, and M.C. Holmes. (2005). Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651.
Wright, D.A., J.A. Townsend, R.J. Winfrey, Jr., P.A. Irwin, J. Rajagopal, P.M. Lonosky, B.D. Hall, M.D. Jondle, and D.F. Voytas. (2005). High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J 44:693–705.
Lloyd, A., C.L. Plaisier, D. Carroll, and G.N. Drews. (2005). Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc Natl Acad Sci U S A 102:2232–2237.
Carroll, D., J.J. Morton, K.J. Beumer, and D.J. Segal. (2006). Design, construction and in vitro testing of zinc finger nucleases. Nat Protoc 1:1329–1341.
Wright, D.A., S. Thibodeau-Beganny, J.D. Sander, R.J. Winfrey, A.S. Hirsh, M. Eichtinger, F. Fu, M.H. Porteus, D. Dobbs, D.F. Voytas, and J.K. Joung. (2006). Standardized reagents and protocols for engineering zinc finger nucleases by modular assembly. Nat Protoc 1:1637–1652.
Smith, J., M. Bibikova, F.G. Whitby, A.R. Reddy, S. Chandrasegaran, and D. Carroll. (2000). Requirements for double-strand cleavage by chimeric restriction enzymes with zinc finger DNA-recognition domains. Nucleic Acids Res 28:3361–3369.
Mandell, J.G., and C.F. Barbas, III. (2006). Zinc finger tools: custom DNA-binding domains for transcription factors and nucleases. Nucleic Acids Res 34:W516–W523.
Liu, P.Q., E.J. Rebar, L. Zhang, Q. Liu, A.C. Jamieson, Y. Liang, H. Qi, P.X. Li, B. Chen, M.C. Mendel, X. Zhong, Y.L. Lee, S.P. Eisenberg, S.K. Spratt, C.C. Case, and A.P. Wolffe. (2001). Regulation of an endogenous locus using a panel of designed zinc finger proteins targeted to accessible chromatin regions. Activation of vascular endothelial growth factor A. J Biol Chem 276:11323–11334.
Liu, Q., Z. Xia, X. Zhong, and C.C. Case. (2002). Validated zinc finger protein designs for all 16 GNN DNA triplet targets. J Biol Chem 277:3850–3856.
Dreier, B., R.R. Beerli, D.J. Segal, J.D. Flippin, and C.F. Barbas, III. (2001). Development of zinc finger domains for recognition of the 5′-ANN-3′ family of DNA sequences and their use in the construction of artificial transcription factors. J Biol Chem 276:29466–29478.
Zhang, L., S.K. Spratt, Q. Liu, B. Johnstone, H. Qi, E.E. Raschke, A.C. Jamieson, E.J. Rebar, A.P. Wolffe, and C.C. Case. (2000). Synthetic zinc finger transcription factor action at an endogenous chromosomal site. Activation of the human erythropoietin gene. J Biol Chem 275:33850–33860.
Mani, M., K. Kandavelou, F.J. Dy, S. Durai, and S. Chandrasegaran. (2005). Design, engineering, and characterization of zinc finger nucleases. Biochem Biophys Res Commun 335:447–457.
Ruminy, P., C. Derambure, S. Chandrasegaran, and J.P. Salier. (2001). Long-range identification of hepatocyte nuclear factor-3 (FoxA) high and low-affinity binding sites with a chimeric nuclease. J Mol Biol 310:523–535.
Bibikova, M., D. Carroll, D.J. Segal, J.K. Trautman, J. Smith, Y.G. Kim, and S. Chandrasegaran. (2001). Stimulation of homologous recombination through targeted cleavage by chimeric nucleases. Mol Cell Biol 21:289–297.
Bibikova, M., K. Beumer, J.K. Trautman, and D. Carroll. (2003). Enhancing gene targeting with designed zinc finger nucleases. Science 300:764.
Bibikova, M., M. Golic, K.G. Golic, and D. Carroll. (2002). Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161:1169–1175.
Morton, J., M.W. Davis, E.M. Jorgensen, and D. Carroll. (2006). Induction and repair of zinc-finger nuclease-targeted double-strand breaks in Caenorhabditis elegans somatic cells. Proc Natl Acad Sci U S A 103:16370–16375.
Alwin, S., M.B. Gere, E. Guhl, K. Effertz, C.F. Barbas, III, D.J. Segal, M.D. Weitzman, and T. Cathomen. (2005). Custom zinc-finger nucleases for use in human cells. Mol Ther 12:610–617.
Beumer, K., G. Bhattacharyya, M. Bibikova, J.K. Trautman, and D. Carroll. (2006). Efficient gene targeting in Drosophila with zinc-finger nucleases. Genetics 172:2391–2403.
Porteus, M.H. (2006). Mammalian gene targeting with designed zinc finger nucleases. Mol Ther 13:438–446.
Moehle, E.A., J.M. Rock, Y.L. Lee, Y. Jouvenot, R.C. DeKelver, P.D. Gregory, F.D. Urnov, and M.C. Holmes. (2007). Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc Natl Acad Sci U S A 104:3055–3060.
Porteus, M.H., and D. Carroll. (2005). Gene targeting using zinc finger nucleases. Nat Biotechnol 23:967–973.
Wilson, J.H. (2003). Pointing fingers at the limiting step in gene targeting. Nat Biotechnol 21:759–760.
Porteus, M.H., J.P. Connelly, and S.M. Pruett. (2006). A look to future directions in gene therapy research for monogenic diseases. PLoS Genet 2:e133.
Miller, J.C., M.C. Holmes, J. Wang, D.Y. Guschin, Y.L. Lee, I. Rupniewski, C.M. Beausejour, A.J. Waite, N.S. Wang, K.A. Kim, P.D. Gregory, C.O. Pabo, and E.J. Rebar. (2007). An improved zinc-finger nuclease architecture for highly specific genome editing. Nat Biotechnol 25:778–785.
Szczepek, M., V. Brondani, J. Buchel, L. Serrano, D.J. Segal, and T. Cathomen. (2007). Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat Biotechnol 25:786–793.
Acknowledgments
We thank Dr. Sundar Durai for drawing the figures. The research on ZFNs in our lab has been supported by various grants from National Institutes of Health, USA, during the past 13 years; it is currently being funded by the research grant GM077291 from NIGMS/NIH. Our work on ZFN-mediated gene targeting in human stem cells is partially supported by a grant from the Maryland Stem Cell Research Fund.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Humana Press, a part of Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Kandavelou, K., Chandrasegaran, S. (2009). Custom-Designed Molecular Scissors for Site-Specific Manipulation of the Plant and Mammalian Genomes. In: Foote, R., Lee, J. (eds) Micro and Nano Technologies in Bioanalysis. Methods in Molecular Biology™, vol 544. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-59745-483-4_40
Download citation
DOI: https://doi.org/10.1007/978-1-59745-483-4_40
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-934115-40-4
Online ISBN: 978-1-59745-483-4
eBook Packages: Springer Protocols