Summary
Cervical cancer, a potentially preventable disease, remains the second most common malignancy in women worldwide. Human papillomavirus is the single most important etiological agent in cervical cancer. HPV contributes to neoplastic progression through the action of two viral oncoproteins E6 and E7, which interfere with critical cell cycle pathways, p53, and retinoblastoma. However, evidence suggests that HPV infection alone is insufficient to induce malignant changes and other host genetic variations are important in the development of cervical cancer. Advances in molecular biology and high throughput gene expression profiling technologies have heralded a new era in biomarker discovery and identification of molecular targets related to carcinogenesis. These advancements have improved our understanding of carcinogenesis and will facilitate screening, early detection, management, and personalised targeted therapy.
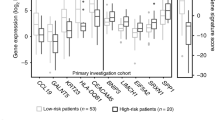
In this chapter, we have described the use of high density microarrays to assess gene expression profiles in cervical cancer. Using this approach we have identified a number of novel genes which are differentially expressed in cervical cancer, including several genes involved in cell cycle regulation. These include p16ink4a, MCM 3 and 5, CDC6, Geminin, Cyclins A-D, TOPO2A, CDCA1, and BIRC5. We have validated expression of mRNA using real-time PCR and protein by immunohistochemistry.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ferlay, J., et al. Globocan 2002. Cancer incidence, mortality and prevalence worldwide. IARC CancerBase number 5. Version 2.0. IARC Press Lyon 2004. Available at http://www-dep-iarc.fr. Accessed 24 October, 2006.
Bray, F., Carstensen, B., Moller, H., et al. (2005) Incidence trends of adenocarcinoma of the cervix in 13 European countries. Cancer Epidemiol Biomarkers Prev 14, 2191–2199.
Schorge, J.O., Knowles, L.M., and Lea, J.S. (2004) Adenocarcinoma of the cervix. Curr Treat Options Oncol 5, 119–127.
Tjalma, W.A., Van Waes, T.R., Van den Eeden, L.E., and Bogers, J.J. (2005) Role of human papillomavirus in the carcinogenesis of squa-mous cell carcinoma and adenocarcinoma of the cervix. Best Pract Res Clin Obstet Gynaecol 19, 469–483.
Duensing, S. and Munger, K. (2004) Mechanisms of genomic instability in human cancer: insights from studies with human papillomavi-rus oncoproteins. Int J Cancer 109, 157–162.
Nanda, K., McCrory, D.C., Myers, E.R., et al. (2000) Accuracy of the Papanicolaou test in screening for and follow-up of cervical cyto-logic abnormalities: a systematic review. Ann Intern Med 132, 810–819.
Shim, C., Zhang, W., Rhee, C.H., and Lee. J.H. (1998) Profiling of differentially expressed genes in human primary cervical cancer by complementary DNA expression array. Clin Cancer Res 4, 3045–3050.
Cheng, Q., Lau, W.M., Chew, S.H., Ho, T.H., Tay, S.K., and Hui, K.M (2002). Identification of molecular markers for the early detection of human squamous cell carcinoma of the uterine cervix. Br J Cancer 86, 274–281.
Santin, A.D., Zhan, F., Bignotti, E., et al. (2005) Gene expression profiles of primary HPV16- and HPV18-infected early stage cervical cancers and normal cervical epithelium: identification of novel candidate molecular markers for cervical cancer diagnosis and therapy. Virology 331, 269–291.
Zhai, Y., Kuick, R., Nan, B., Ota, I., Weiss, S.J., Trimble, C.L., Fearon, E.R., and Cho, K.R. (2007) Gene expression analysis of prein-vasive and invasive cervical squamous cell carcinomas identifies HOXC10 as a key mediator of. Cancer Res 67, 10163–10172.
Chao, A., Wang, T.H., Lee, Y.S., Hsueh, S., Chao, A.S., Chang, T.C., Kung, W.H., Huang, S.L., Chao, F.Y., Wei, M.L., and Lai, C.H. (2006) Molecular characterization of adenocarcinoma and squamous carcinoma of the uterine cervix using microarray analysis of gene expression. Int J Cancer 119, 91–98.
Contag, S.A., Gostout, B.S., Clayton, A.C., Dixon, M.H., McGovern, R.M., and Calhoun, E.S. (2004) Comparison of gene expression in squamous cell carcinoma and adenocarcinoma of the uterine cervix. Gynecol Oncol 95, 610–617.
Hudelist, G., Czerwenka, K., Singer, C., Pischinger, K., Kubista, E., and Manavi, M. (2005) cDNA array analysis of cytobrush-col-lected normal and malignant cervical epithelial cells: a feasibility study. Cancer Genet Cytogenet 158, 35–42.
Manavi, M., Hudelist, G., Fink-Retter, A., Gschwandtler-Kaulich, D., Pischinger, K., and Czerwenka, K. (2007) Gene profiling in Pap-cell smears of high-risk human papillomavirus-positive squamous cervical carcinoma. Gynecol Oncol 105, 418–426.
Bachtiary, B., Boutros, P.C., Pintilie, M., Shi, W., Bastianutto, C., Li, J.H., Schwock, J., Zhang, W., Penn, L.Z., Jurisica, I., Fyles, A., and Liu, F.F. (2006) Gene expression profiling in cervical cancer: an exploration of intratumor heterogeneity. Clin Cancer Res 12, 5632– 5640.
Gius, D., Funk, M.C., Chuang, E.Y., Feng, S., Huettner, P.C., Nguyen, L., Bradbury, C.M., Mishra, M., Gao, S., Buttin, B.M., Cohn, D.E., Powell, M.A., Horowitz, N.S., Whitcomb, B.P., and Rader, J.S. (2007) Profiling microdissected epithelium and stroma to model genomic signatures for cervical carcino-genesis accommodating for covariates, Cancer Res 67, 7113–7123.
Murphy, N., Ring, M., Heffron, C.C., et al. (2005) p16INK4A, CDC6, and MCM5: predictive biomarkers in cervical preinvasive neoplasia and cervical cancer. J Clin Pathol 58, 525–534.
Murphy, N., Ring, M., Heffron, C.C., et al. (2005) Quantitation of CDC6 and MCM5 mRNA in cervical intraepithelial neoplasia and invasive squamous cell carcinoma of the cervix. Mod Pathol 18, 844–849.
Martin, C.M., Kehoe, L., Spillane, C.O., and O'Leary, J.J. (2007) Gene discovery in cervical cancer: towards diagnostic and therapeutic biomarkers. Mol Diagn Ther 11, 277–290.
Martin, C.M., Astbury, K., and O'Leary, J.J. (2006) Molecular profiling of cervical neopla-sia. Expert Rev Mol Diagn 6, 217–229.
Martin, C.M., Sheils, O., and O'Leary, J.J. (2005) Real Time TaqMan® PCR Technology. In The Science of Laboratory Diagnosis, second edition (editors Crocker, J. Burnett, D.), 495–504. Wiley, New York.
Livak, K.J. and Schmittgen, T.D. (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2(-Delta Delta C(T) ) Method. Methods 25, 402–408.
Barbacioru, C.C., Wang, Y., Canales, R.D., Sun, Y.A., Keys, D.N., Chan, F., Poulter, K.A., Samaha, R.R. (2006) Effect of various normalization methods on Applied Biosystems expression array system data. BMC Bioinfor-matics (7), 533–547.
Acknowledgments
The authors wish to thank other members of the Cervical Cancer Research Group at the Coombe Women's and Infants University Hospital and Trinity College, Dublin and Cerviva; The Irish Cervical Screening Research Consortium. This work and the group are funded by The Coombe Women's and Infants University Hospital, the Royal City of Dublin Trust, The Meath Foundation, The Health Research Board Ireland, and The Irish Cancer Society.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Martin, C.M., Astbury, K., McEvoy, L., O'Toole, S., Sheils, O., O'Leary, J.J. (2009). Gene Expression Profiling in Cervical Cancer: Identification of Novel Markers for Disease Diagnosis and Therapy. In: Kozlov, S.V. (eds) Inflammation and Cancer. Methods in Molecular Biology™, vol 511. Humana Press. https://doi.org/10.1007/978-1-59745-447-6_15
Download citation
DOI: https://doi.org/10.1007/978-1-59745-447-6_15
Publisher Name: Humana Press
Print ISBN: 978-1-934115-14-5
Online ISBN: 978-1-59745-447-6
eBook Packages: Springer Protocols