Abstract
The continuous improvement of gene editing tools has allowed a major revolution in biological sciences. Although a variety of gain and loss-of-function approaches have been widely used for the last decades, some limitations arose from non-specific targeting or lack of complete inhibition of the gene of interest. CRISPR/Cas9 editing technology introduced new and significant advantages because it can directly modify the gene of interest and completely blocks its expression.
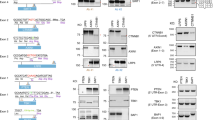
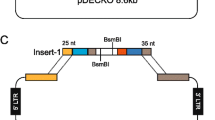
In the context of cancer studies, the heterogeneity of the tumor microenvironment requires comprehensive approaches to unveil the contribution of multiple genes. For example, a deeper understanding of the biology of proteases such as ADAMTS (a disintegrin and metalloproteinase with thrombospondin type 1 motifs) will improve our perspective of complex phenomena affected by extracellular matrix remodeling, including embryonic development, angiogenesis, immune infiltration, metastasis, and tumor plasticity. Here, we present a method using CRISPR/Cas9 technology to inhibit the expression of the representative ADAMTS1 in cancer cells. Following the first steps of gene edition, we pursue further selection of silenced cells and provide a detailed description of sequence analysis and validation assays. This method leads to inactivation of ADAMTS1 in cancer cells, providing a relevant biological tool that will allow subsequent in vivo and in vitro ADAMTS1 functional analysis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hall B, Limaye A, Kulkarni AB (2009) Overview: generation of gene knockout mice. In: Current Protocols in Cell Biology. Wiley, Hoboken, NJ, pp 19.12.1–19.12.17
Grimm D (2006) MOUSE GENETICS: a mouse for every gene. Science 312:1862–1866. https://doi.org/10.1126/science.312.5782.1862
Kim J-S (2016) Genome editing comes of age. Nat Protoc 11:1573–1578. https://doi.org/10.1038/nprot.2016.104
Sakuma T, Yamamoto T (2015) CRISPR/Cas9: the leading edge of genome editing technology. In: Target. Genome Ed. Using site-specific nucleases. Springer, Tokyo, pp 25–41
Ochiai H, Yamamoto T (2015) Genome editing using zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs). In: Target. Genome Ed. Using site-specific nucleases. Springer, Tokyo, pp 3–24
Turk B (2006) Targeting proteases: successes, failures and future prospects. Nat Rev Drug Discov 5:785–799. https://doi.org/10.1038/nrd2092
Lu P, Takai K, Weaver VM, Werb Z (2011) Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb Perspect Biol 3:a005058. https://doi.org/10.1101/cshperspect.a005058
Bix G, Iozzo RV (2005) Matrix revolutions: “tails” of basement-membrane components with angiostatic functions. Trends Cell Biol 15:52–60. https://doi.org/10.1016/j.tcb.2004.11.008
Kuno K, Kanada N, Nakashima E et al (1997) Molecular cloning of a gene encoding a new type of metalloproteinase-disintegrin family protein with thrombospondin motifs as an inflammation associated gene. J Biol Chem 272:556–562. https://doi.org/10.1074/jbc.272.1.556
Vázquez F, Hastings G, Ortega MA et al (1999) METH-1, a human ortholog of ADAMTS-1, and METH-2 are members of a new family of proteins with angio-inhibitory activity. J Biol Chem 274:23349–23357. https://doi.org/10.1074/jbc.274.33.23349
Rodríguez-Manzaneque JC, Fernández-Rodríguez R, Rodríguez-Baena FJ, Iruela-Arispe ML (2015) ADAMTS proteases in vascular biology. Matrix Biol 44–46:38–45. https://doi.org/10.1016/j.matbio.2015.02.004
Cal S, López-Otín C (2015) ADAMTS proteases and cancer. Matrix Biol 44–46:77–85. https://doi.org/10.1016/j.matbio.2015.01.013
Apte SS (2009) A disintegrin-like and metalloprotease (reprolysin-type) with thrombospondin type 1 motif (ADAMTS) superfamily: functions and mechanisms. J Biol Chem 284:31493–31497. https://doi.org/10.1074/jbc.R109.052340
Mittaz L, Russell DL, Wilson T et al (2004) Adamts-1 is essential for the development and function of the urogenital system. Biol Reprod 70:1096–1105. https://doi.org/10.1095/biolreprod.103.023911
Shindo T, Kurihara H, Kuno K et al (2000) ADAMTS-1: a metalloproteinase-disintegrin essential for normal growth, fertility, and organ morphology and function. J Clin Invest 105:1345–1352. https://doi.org/10.1172/JCI8635
Fernández-Rodríguez R, Rodríguez-Baena FJ, Martino-Echarri E et al (2016) Stroma-derived but not tumor ADAMTS1 is a main driver of tumor growth and metastasis. Oncotarget 7:34507–34519. https://doi.org/10.18632/oncotarget.8922
Casal C, Torres-Collado AX, Plaza-Calonge MCDC et al (2010) ADAMTS1 contributes to the acquisition of an endothelial-like phenotype in plastic tumor cells. Cancer Res 70:4676–4686. https://doi.org/10.1158/0008-5472.CAN-09-4197
Rodríguez-Manzaneque JC, Milchanowski AB, Dufour EK et al (2000) Characterization of METH-1/ADAMTS1 processing reveals two distinct active forms. J Biol Chem 275:33471–33479. https://doi.org/10.1074/jbc.M002599200
Acknowledgments
Work in the author’s laboratory has been supported by grants from the Ministerio de Economía y Competitividad and Instituto de Salud Carlos III from Spain, co-financed by FEDER (PI16/00345 to JCRM). OS is supported by a scholarship from Doctoral and Post-doctoral Program, National Secretary of Science and Technology of Panama (#270-2015-011).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Peris-Torres, C., Serrano, O., Plaza-Calonge, M.d.C., Rodríguez-Manzaneque, J.C. (2020). Inhibition of ADAMTS1 Expression by Lentiviral CRISPR/Cas9 Gene Editing Technology. In: Apte, S. (eds) ADAMTS Proteases. Methods in Molecular Biology, vol 2043. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9698-8_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9698-8_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9697-1
Online ISBN: 978-1-4939-9698-8
eBook Packages: Springer Protocols