Abstract
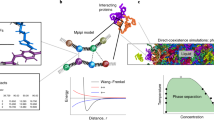
Globular proteins are roughly spherical biomolecules with attractive and highly directional interactions. This microscopic observation motivates describing these proteins as patchy particles: hard spheres with attractive surface patches. Mapping a biomolecule to a patchy model requires simplifying effective protein–protein interactions, which in turn provides a microscopic understanding of the protein solution behavior. The patchy model can indeed be fully analyzed, including its phase diagram. In this chapter, we detail the methodology of mapping a given protein to a patchy model and of determining the phase diagram of the latter. We also briefly describe the theory upon which the methodology is based, provide practical information, and discuss potential pitfalls. Data and scripts relevant to this work have been archived and can be accessed at https://doi.org/10.7924/r4ww7bs1p.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Change history
01 July 2020
The acknowledgement section text has been updated in the chapter.
References
Chen V, Davis I, Richardson D (2009) KiNG (Kinemage, next generation): a versatile interactive molecular and scientific visualization program. Protein Sci 18(11):2403–2409
Berendsen H, van der Spoel D, van Drunen R (1995) Gromacs: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91(1):43–56
Case D, Cerutti D, Cheatham T III, Darden T, Duke R, Giese T, Gohlke H, Goetz A, Greene D, Homeyer N, Izadi S, Kovalenko A, Lee T, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Mermelstein D, Merz K, Monard G, Nguyen H, Omelyan I, Onufriev A, Pan F, Qi R, Roe D, Roitberg A, Sagui C, Simmerling C, Botello-Smith W, Swails J, Walker R, Wang J, Wolf R, Wu X, Xiao L, York D, PA K (2017) Amber 2017. Tech. rep., University of California, San Francisco
Frenkel D, Smit B (2001) Understanding molecular simulation: from algorithms to applications. Academic, Orlando
Rovigatti L, Russo J, Romano F (2018) How to simulate patchy particles. Eur Phys J E 41(5):59
Rovigatti L, Romano F, Russo J (2018) lorenzo-rovigatti/patchyparticles v1.0.1. https://doi.org/10.5281/zenodo.1171695
Fusco D, Headd J, De Simone A, Wang J, Charbonneau P (2014) Characterizing protein crystal contacts and their role in crystallization: rubredoxin as a case study. Soft Matter 10(2):290–302
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28(1):235–242
Bönisch H, Schmidt C, Bianco P, Ladenstein R (2005) Ultrahigh-resolution study on Pyrococcus abyssi rubredoxin. I. 0.69 Å X-ray structure of mutant W4L/R5S. Acta Crystallogr D 61(7):990–1004
Kästner J (2011) Umbrella sampling. Wiley Interdiscip Rev Comput Mol Sci 1(6):932–942
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372(3):774–797
Rostkowski M, Olsson M, Søndergaard C, Jensen J (2011) Graphical analysis of pH-dependent properties of proteins predicted using PROPKA. BMC Struct Biol 11(1):1
Hub J, De Groot B, Van Der Spoel D (2010) g_wham – a free weighted histogram analysis implementation including robust error and autocorrelation estimates. J Chem Theory Comput 6(12):3713–3720
Kern N, Frenkel D (2003) Fluid–fluid coexistence in colloidal systems with short-ranged strongly directional attraction. J Chem Phys 118(21):9882–9889
Fusco D, Charbonneau P (2016) Soft matter perspective on protein crystal assembly. Colloids Surf B 137:22–31
Sear R (1999) Phase behavior of a simple model of globular proteins. J Chem Phys 111(10):4800–4806
Wentzel N, Gunton J (2008) Effect of solvent on the phase diagram of a simple anisotropic model of globular proteins. J Phys Chem B 112(26):7803–7809
Dixit N, Zukoski C (2002) Crystal nucleation rates for particles experiencing anisotropic interactions. J Chem Phys 117(18):8540–8550
Vega C, Sanz E, Abascal J, Noya E (2008) Determination of phase diagrams via computer simulation: methodology and applications to water, electrolytes and proteins. J Phys Condens Matter 20(15):153101
Romano F, Sanz E, Sciortino F (2010) Phase diagram of a tetrahedral patchy particle model for different interaction ranges. J Chem Phys 132(18):184501
Frenkel D, Ladd A (1984) New Monte Carlo method to compute the free energy of arbitrary solids. Application to the FCC and HCP phases of hard spheres. J Chem Phys 81(7):3188–3193
Riley KF, Hobson MP, Bence SJ (2006) Mathematical methods for physics and engineering: a comprehensive guide. Cambridge University Press, Cambridge
Kofke D (1993) Gibbs-Duhem integration: a new method for direct evaluation of phase coexistence by molecular simulation. Mol Phys 78(6):1331–1336
Kofke D (1993) Direct evaluation of phase coexistence by molecular simulation via integration along the saturation line. J Chem Phys 98(5):4149–4162
Panagiotopoulos A (1987) Direct determination of phase coexistence properties of fluids by Monte Carlo simulation in a new ensemble. Mol Phys 61(4):813–826
Noro MG, Frenkel D (2000) Extended corresponding-states behavior for particles with variable range attractions. J Chem Phys 113(8):2941–2944
Blote H, Luijten E, Heringa J (1995) Ising universality in three dimensions: a Monte Carlo study. J Phys A 28(22):6289
Fusco D, Charbonneau P (2013) Crystallization of asymmetric patchy models for globular proteins in solution. Phys Rev E 88(1):012721
Tang Z, Zhang Z, Wang Y, Glotzer S, Kotov N (2006) Self-assembly of CdTe nanocrystals into free-floating sheets. Science 314(5797):274–278
Ye X, Chen J, Engel M, Millan J, Li W, Qi L, Xing G, Collins J, Kagan C, Li J et al. (2013) Competition of shape and interaction patchiness for self-assembling nanoplates. Nat Chem 5(6):466
Glotzer S, Solomon M (2007) Anisotropy of building blocks and their assembly into complex structures. Nat Mater 6(8):557
Bianchi E, Capone B, Kahl G, Likos C (2015) Soft-patchy nanoparticles: modeling and self-organization. Faraday Discuss 181:123–138
de las Heras D, da Gama M (2016) Temperature (de)activated patchy colloidal particles. J Phys Condens Matter 28(24):244008
Wilber AW, Doye JP, Louis AA, Noya EG, Miller MA, Wong P (2007) Reversible self-assembly of patchy particles into monodisperse icosahedral clusters. J Chem Phys 127(8):08B618
Acknowledgments
We thank Diana Fusco for guiding discussions. The authors acknowledge support from National Science Foundation Grant no. NSF DMR-1749374. This work used the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number ACI-1548562, as well as the Duke Compute Cluster.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Altan, I., Charbonneau, P. (2019). Obtaining Soft Matter Models of Proteins and their Phase Behavior. In: McManus, J. (eds) Protein Self-Assembly. Methods in Molecular Biology, vol 2039. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9678-0_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9678-0_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9677-3
Online ISBN: 978-1-4939-9678-0
eBook Packages: Springer Protocols