Abstract
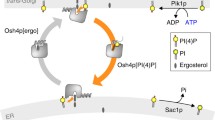
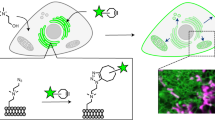
In order to understand how lipids are sorted between cellular compartments, kinetic assays are required to selectively follow the transport of lipid species in cells. We present here a microfluidics-based protocol to follow the transport of phosphatidylserine (PS) in yeast cells from the site of its synthesis, the endoplasmic reticulum (ER), to downstream compartments, primarily the plasma membrane under our conditions. This assay takes advantage of yeast cells lacking Cho1, the enzyme responsible for PS synthesis. Lyso-PS can be added exogenously and is taken up by the cells and converted to PS. Because acylation of lyso-PS to PS appears to occur at the ER, anterograde transport of PS from the ER can then be followed by fluorescent microscopy using the specific PS reporter C2Lact-GFP. We describe the construction of the required cho1Δ yeast strain and the preparation of lyso-PS. We present an example of the use of this assay to follow the activity of the yeast PS transport proteins Osh6 and Osh7.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wong LH, Copic A, Levine TP (2017) Advances on the transfer of lipids by lipid transfer proteins. Trends Biochem Sci 42:516–530
Moser von Filseck J, Vanni S, Mesmin B et al (2015) A phosphatidylinositol-4-phosphate powered exchange mechanism to create a lipid gradient between membranes. Nat Commun 6:6671
Kannan M, Lahiri S, Liu LK et al (2017) Phosphatidylserine synthesis at membrane contact sites promotes its transport out of the ER. J Lipid Res 58:553–562
Georgiev AG, Johansen J, Ramanathan VD et al (2013) Arv1 regulates PM and ER membrane structure and homeostasis but is dispensable for intracellular sterol transport. Traffic 14:912–921
Yeung T, Gilbert GE, Shi J et al (2008) Membrane phosphatidylserine regulates surface charge and protein localization. Science 319:210–213
Andersen MH, Graversen H, Fedosov SN et al (2000) Functional analyses of two cellular binding domains of bovine lactadherin. Biochemistry 39:6200–6206
Del Vecchio K, Stahelin RV (2018) Investigation of the phosphatidylserine binding properties of the lipid biosensor, Lactadherin C2 (LactC2), in different membrane environments. J Bioenerg Biomembr 50:1–10
Platre MP, Noack LC, Doumane M et al (2018) A combinatorial lipid code shapes the electrostatic landscape of plant endomembranes. Dev Cell 45:465–480 e411
Maeda K, Anand K, Chiapparino A et al (2013) Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins. Nature 501:257–261
Moser von Filseck J, Copic A, Delfosse V et al (2015) INTRACELLULAR TRANSPORT. Phosphatidylserine transport by ORP/Osh proteins is driven by phosphatidylinositol 4-phosphate. Science 349:432–436
Fairn GD, Hermansson M, Somerharju P et al (2011) Phosphatidylserine is polarized and required for proper Cdc42 localization and for development of cell polarity. Nat Cell Biol 13:1424–1430
Atkinson K, Fogel S, Henry SA (1980) Yeast mutant defective in phosphatidylserine synthesis. J Biol Chem 255:6653–6661
Yamaoka Y, Yu Y, Mizoi J et al (2011) PHOSPHATIDYLSERINE SYNTHASE1 is required for microspore development in Arabidopsis thaliana. Plant J 67:648–661
Natarajan P, Wang J, Hua Z et al (2004) Drs2p-coupled aminophospholipid translocase activity in yeast Golgi membranes and relationship to in vivo function. Proc Natl Acad Sci U S A 101:10614–10619
Riekhof WR, Wu J, Gijon MA et al (2007) Lysophosphatidylcholine metabolism in Saccharomyces cerevisiae: the role of P-type ATPases in transport and a broad specificity acyltransferase in acylation. J Biol Chem 282:36853–36861
Longtine MS, McKenzie A 3rd, Demarini DJ et al (1998) Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14:953–961
Janke C, Magiera MM, Rathfelder N et al (2004) A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21:947–962
Gietz D, St Jean A, Woods RA et al (1992) Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res 20:1425
Looke M, Kristjuhan K, Kristjuhan A (2011) Extraction of genomic DNA from yeasts for PCR-based applications. BioTechniques 50:325–328
Acknowledgments
We thank Jackson-Verbavatz and Leon labs for support, A-C Gavin for sharing yeast strains, and the ImagoSeine facility at the Institut Jacques Monod, member of the France-Bio-Imaging national research structure (ANR-10-INBS-04). This work is funded by the French National Research Agency grant ExCHANGE (ANR-16-CE13-0006) and by the CNRS.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
D’Ambrosio, J.M., Albanèse, V., Čopič, A. (2019). Following Anterograde Transport of Phosphatidylserine in Yeast in Real Time. In: Drin, G. (eds) Intracellular Lipid Transport. Methods in Molecular Biology, vol 1949. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-9136-5_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9136-5_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-9135-8
Online ISBN: 978-1-4939-9136-5
eBook Packages: Springer Protocols