Abstract
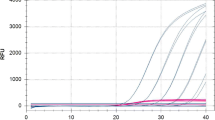
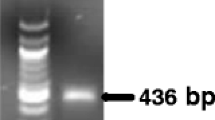
The chapter describes a simple quantitative approach to assess phytoplasma load in samples obtained from “Candidatus Phytoplasma mali”-infected apple plants without the use of external standard curves. The assay is based on the simultaneous detection of a gene of the pathogen and a gene of the host plant in a duplex single-tube real-time PCR reaction using TaqMan chemistry. The quantity of the phytoplasma, relative to its host plant, is determined as the difference between the CT values of the two target genes (ΔCT). A critical data analysis step, affecting the inter-assay reproducibility between different amplification runs, is the setting of the threshold level, which is achieved by the recurrent analysis of a calibrator sample. The relative quantification procedure allows analyzing 45 DNA samples in duplicates on a 96-well reaction plate, in addition to the control and calibrator samples, and thus contributes to a substantial increase of analysis throughput and decrease of reagent/consumable costs per sample.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ravnikar M, Mehle N, Gruden K, Dreo T (2016) Real-time PCR. In: Boonham N, Tomlinson J, Mumford R (eds) Molecular methods in plant disease diagnostics: principles and protocols. CABI Wallingford, Oxfordshire, Boston, pp 28–58
Bustin SA, Nolan T (2004) Chemistries. In: Bustin SA (ed) A–Z of quantitative PCR. International University Line, La Jolla, pp 215–278
Pfaffl MW (2004) Quantification strategies in real-time PCR. In: Bustin SA (ed) A–Z of quantitative PCR. International University Line, La Jolla, pp 87–120
Baric S, Dalla Via J (2004) A new approach to apple proliferation detection: a highly sensitive real-time PCR assay. J Microbiol Methods 57:135–145
Galetto L, Bosco D, Marzachì C (2005) Universal and group-specific real-time PCR diagnosis of flavescence dorée (16Sr-V), bois noir (16Sr-XII) and apple proliferation (16Sr-X) phytoplasmas from field-collected plant hosts and insect vectors. Ann Appl Biol 147:191–201
Pelletier C, Salar P, Gillet J, Cloquemin G, Very P, Foissac X, Malembic-Maher S (2009) Triplex real-time PCR assay for sensitive and simultaneous detection of grapevine phytoplasmas of the 16SrV and 16SrXII-A groups with an endogenous analytical control. Vitis 48:87–95
Linck H, Krüger E, Reineke A (2017) A multiplex TaqMan qPCR assay for sensitive and rapid detection of phytoplasmas infecting Rubus species. PLoS One 12:e0177808
Christensen NM, Nicolaisen M, Hansen M, Schulz A (2004) Distribution of phytoplasmas in infected plants as revealed by real-time PCR and bioimaging. Mol Plant-Microbe Interact 17:1175–1184
Torres E, Bertolini E, Cambra M, Montón C, Martín MP (2005) Real-time PCR for simultaneous and quantitative detection of quarantine phytoplasmas from apple proliferation (16SrX) group. Mol Cell Probes 19:334–340
Saracco P, Bosco D, Veratti F, Marzachì C (2006) Quantification over time of chrysanthemum yellows phytoplasma (16Sr-I) in leaves and roots of the host plant Chrysanthemum carinatum (Schousboe) following inoculation with its insect vector. Physiol Mol Plant Pathol 67:212–219
Bisognin C, Schneider B, Salm H, Grando MS, Jarausch W, Moll E, Seemüller E (2008) Apple proliferation resistance in apomictic rootstocks and its relationship to phytoplasma concentration and simple sequence repeat genotypes. Phytopathology 98:153–158
Baric S, Berger J, Cainelli C, Kerschbamer C, Letschka T, Dalla Via J (2011) Seasonal colonisation of apple trees by ‘Candidatus Phytoplasma mali’ revealed by a new quantitative TaqMan real-time PCR approach. Eur J Plant Pathol 129:455–467
Jawhari M, Abrahamian P, Sater AA, Sobh H, Tawidian P, Abou-Jawdah Y (2015) Specific PCR and real-time PCR assays for detection and quantitation of ‘Candidatus Phytoplasma phoenicium’. Mol Cell Probes 29:63–70
Arratia-Castro AA, Santos-Cervantes ME, Arce-Leal ÁP, Espinoza-Mancillas MG, Rodríguez Negrete EA, Méndez-Lozano J, Arocha-Rosete Y, Leyva-López NE (2016) Detection and quantification of ‘Candidatus Phytoplasma asteris’ and ‘Candidatus Liberibacter asiaticus’ at early and late stages of Huanglongbing disease development. Can J Plant Pathol 38:411–421
Rutledge RG, Côté C (2003) Mathematics of quantitative kinetic PCR and the application of standard curves. Nucleic Acids Res 31:e93
Baric S (2012) Quantitative real-time PCR analysis of ‘Candidatus Phytoplasma mali’ without external standard curves. Erwerbs-Obstbau 54:147–153
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Nolan T, Hands RE, Bustin SH (2006) Quantification of mRNA using real-time RT-PCR. Nat Protoc 1:1559–1582
Hogenhout SA, Oshima K, Ammar ED, Kakizawa S, Kingdom HN, Namba S (2008) Phytoplasmas: bacteria that manipulate plants and insects. Mol Plant Pathol 9:403–423
Gachon CM, Strittmatter M, Müller DG, Kleinteich J, Küpper FC (2009) Detection of differential host susceptibility to the marine oomycete pathogen Eurychasma dicksonii by real-time PCR: not all algae are equal. Appl Environ Microbiol 75:322–328
Liu ZL, Palmquist DE, Ma MG, Liu J, Alexander NJ (2009) Application of a master equation for quantitative mRNA analysis using qRT-PCR. J Biotechnol 143:10–16
Acknowledgments
The author is grateful to J. Dalla Via for critically reading and discussing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Baric, S. (2019). Duplex TaqMan Real-Time PCR for Rapid Quantitative Analysis of a Phytoplasma in Its Host Plant without External Standard Curves. In: Musetti, R., Pagliari, L. (eds) Phytoplasmas. Methods in Molecular Biology, vol 1875. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8837-2_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8837-2_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8836-5
Online ISBN: 978-1-4939-8837-2
eBook Packages: Springer Protocols