Abstract
Immune cell function is tightly regulated by cellular metabolism, which in turn is strongly linked to the nutrient availability in the microenvironment surrounding the cells. This link is critical for effector CD8+ T cells which, after activation, must migrate from nutrient-rich environments into nutrient-scarce regions such as the tumor microenvironment. Assessing how nutrient availability modulates the metabolism of effector CD8+ T cells is thus key for understanding how harsh environments may impair their proliferation and effector function. Here, we describe an approach to systematically study the impact of the nutrient microenvironment on the metabolism of effector CD8+ T cells, based on performing stable 13C isotope labeling measurements on in vitro-differentiated murine effector CD8+ T cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Parham P (2014) The immune system. Garland Science, New York, NY
Pearce EJ, Pearce EL (2017) Driving immunity: all roads lead to metabolism. Nat Rev Immunol 18(2):81–82
Loftus RM, Finlay DK (2016) Immunometabolism: cellular metabolism turns immune regulator. J Biol Chem 291(1):1–10
O’Neill LAJ, Kishton RJ, Rathmell J (2016) A guide to immunometabolism for immunologists. Nat Rev Immunol 16(9):553–565
Buck MD, Sowell RT, Kaech SM et al (2017) Metabolic instruction of immunity. Cell 169(4):570–586
Wang R, Green DR (2012) Metabolic reprogramming and metabolic dependency in T cells. Immunol Rev 249(1):14–26
Pearce EL, Poffenberger MC, Chang CH et al (2013) Fueling immunity: insights into metabolism and lymphocyte function. Science 342(6155):1242454
Buck MD, O’Sullivan D, Pearce EL (2015) T cell metabolism drives immunity. J Exp Med 212(9):1345–1360
Chang CH, Curtis JD, Maggi LB et al (2013) Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153(6):1239–1251
Renner K, Singer K, Koehl GE et al (2017) Metabolic hallmarks of tumor and immune cells in the tumor microenvironment. Front Immunol 8(8):1–11
Lunt SY, Fendt S-M (2018) Metabolism – a cornerstone of cancer initiation, progression, immune evasion and treatment response. Curr Opin Syst Biol 8:67–72
Cham CM, Driessens G, O’Keefe JP et al (2008) Glucose deprivation inhibits multiple key gene expression events and effector functions in CD8+ T cells. Eur J Immunol 38(9):2438–2450
Chang CH, Qiu J, O’Sullivan D et al (2015) Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162(6):1229–1241
Geiger R, Rieckmann JC, Wolf T et al (2016) L-arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167(3):829–842.e13
Ma EH, Bantug G, Griss T et al (2017) Serine is an essential metabolite for effector T cell expansion. Cell Metab 25:1–13
Fischer K, Hoffmann P, Voelkl S et al (2007) Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood 109(9):3812–3819
Pilon-Thomas S, Kodumudi KN, El-Kenawi AE et al (2016) Neutralization of tumor acidity improves antitumor responses to immunotherapy. Cancer Res 76(6):1381–1390
Kalderon B, Gopher A, Lapidot A (1987) A quantitative analysis of the metabolic pathways of hepatic glucose synthesis in vivo with 13C-labeled substrates. FEBS Lett 213(1):209–214
Katz J, Lee WN, Wals PA et al (1989) Studies of glycogen synthesis and the krebs cycle by mass isotopomer analysis with [U-13C]glucose in rats. J Biol Chem 264(22):12994–13004
Mueller D, Heinzle E (2013) Stable isotope-assisted metabolomics to detect metabolic flux changes in mammalian cell cultures. Curr Opin Biotechnol 24(1):54–59
Buescher JM, Antoniewicz MR, Boros LG et al (2015) A roadmap for interpreting 13C metabolite labeling patterns from cells. Curr Opin Biotechnol 34:189–201
Wittmann C, Heinzle E (1999) Mass spectrometry for metabolic flux analysis. Biotechnol Bioeng 62(6):739–750
Sauer U (2006) Metabolic networks in motion: 13C-based flux analysis. Mol Syst Biol 2:62
Niedenführ S, Wiechert W, Nöh K (2015) How to measure metabolic fluxes: a taxonomic guide for 13C fluxomics. Curr Opin Biotechnol 34:82–90
Tardito S, Oudin A, Ahmed SU et al (2015) Glutamine synthetase activity fuels nucleotide biosynthesis and supports growth of glutamine-restricted glioblastoma. Nat Cell Biol 17(12):1556–1568
Elia I, Broekaert D, Christen S et al (2017) Proline metabolism supports metastasis formation and could be inhibited to selectively target metastasizing cancer cells. Nat Commun 8:15267
Christen S, Lorendeau D, Schmieder R et al (2016) Breast cancer-derived lung metastases show increased pyruvate carboxylase-dependent anaplerosis. Cell Rep 17(3):837–848
Lorendeau D, Rinaldi G, Boon R et al (2017) Dual loss of succinate dehydrogenase (SDH) and complex I activity is necessary to recapitulate the metabolic phenotype of SDH mutant tumors. Metab Eng 43(B):187–197
Berthois Y, Katzenellenbogen JA, Katzenellenbogen BS (1986) Phenol red in tissue culture media is a weak estrogen: implications concerning the study of estrogen-responsive cells in culture. Proc Natl Acad Sci U S A 83(8):2496–2500
Acknowledgments
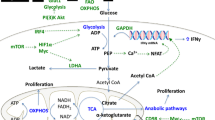
JFG is supported by an FWO (Fonds voor Wetenschappelijk Onderzoek–Vlaanderen, Research Foundation–Flanders) Postdoctoral Fellowship. S-MF acknowledges funding from the European Research Council under the ERC Consolidator Grant Agreement no. 771486 – MetaRegulation, Marie Curie – CIG, FWO – Odysseus II, FWO – Research Grants/Projects, Eugène Yourassowsky Schenking, and KU Leuven – Methusalem Co-Funding. We would like to acknowledge http://www.somersault1824.com for image elements used in Fig. 1 (Creative Commons license CC BY-NC-SA 4.0).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Fernández-García, J., Fendt, SM. (2019). Assessing the Impact of the Nutrient Microenvironment on the Metabolism of Effector CD8+ T Cells. In: Fendt, SM., Lunt, S. (eds) Metabolic Signaling. Methods in Molecular Biology, vol 1862. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8769-6_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8769-6_14
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8768-9
Online ISBN: 978-1-4939-8769-6
eBook Packages: Springer Protocols