Abstract
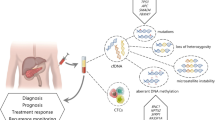
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive tumor and the fourth common cause of cancer death in the Western world. The lack of effective therapeutic strategies is attributed to the late diagnosis of this disease. Methylation markers could improve early detection and help in the surveillance of PDAC after treatment. Analysis of hypermethylation in the tumor tissue and tumor-derived exosomes might help to identify new therapeutic strategies and aid in the understanding of the pathophysiological changes occurring in pancreatic cancer. There are several methods for the detection of methylation events. Whereas methylation-specific PCR (MSP-PCR) is the method of choice, the cost reductions in DNA sequencing enables researchers to add bisulfite sequencing (BSS) to their repertoire if a small number of genes will be tested in a larger set of patients’ samples. During the last years, several techniques to isolate and analyze DNA methylation have been proposed, but DNA modification using sodium bisulfite is still the gold standard.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A (2016) Cancer statistics, 2016. CA Cancer J Clin 66(1):7–30
Schneider G, Siveke JT, Eckel F, Schmid RM (2005) Pancreatic cancer: basic and clinical aspects. Gastroenterology 128(6):1606–1625
Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN, Bruxner TJ, Quinn MC, Nourse C, Murtaugh LC, Harliwong I, Idrisoglu S, Manning S, Nourbakhsh E, Wani S, Fink L, Holmes O, Chin V, Anderson MJ, Kazakoff S, Leonard C, Newell F, Waddell N, Wood S, Xu Q, Wilson PJ, Cloonan N, Kassahn KS, Taylor D, Quek K, Robertson A, Pantano L, Mincarelli L, Sanchez LN, Evers L, Wu J, Pinese M, Cowley MJ, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chantrill LA, Mawson A, Humphris J, Chou A, Pajic M, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Lovell JA, Merrett ND, Toon CW, Epari K, Nguyen NQ, Barbour A, Zeps N, Moran-Jones K, Jamieson NB, Graham JS, Duthie F, Oien K, Hair J, Grutzmann R, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Rusev B, Capelli P, Salvia R, Tortora G, Mukhopadhyay D, Petersen GM, Australian Pancreatic Cancer Genome I, Munzy DM, Fisher WE, Karim SA, Eshleman JR, Hruban RH, Pilarsky C, Morton JP, Sansom OJ, Scarpa A, Musgrove EA, Bailey UM, Hofmann O, Sutherland RL, Wheeler DA, Gill AJ, Gibbs RA, Pearson JV, Waddell N, Biankin AV, Grimmond SM (2016) Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531(7592):47–52
Conroy T, Desseigne F, Ychou M, Bouche O, Guimbaud R, Becouarn Y, Adenis A, Raoul JL, Gourgou-Bourgade S, de la Fouchardiere C, Bennouna J, Bachet JB, Khemissa-Akouz F, Pere-Verge D, Delbaldo C, Assenat E, Chauffert B, Michel P, Montoto-Grillot C, Ducreux M, Groupe Tumeurs Digestives of U, Intergroup P (2011) FOLFIRINOX versus gemcitabine for metastatic pancreatic cancer. N Engl J Med 364(19):1817–1825
Von Hoff DD, Ervin T, Arena FP, Chiorean EG, Infante J, Moore M, Seay T, Tjulandin SA, Ma WW, Saleh MN, Harris M, Reni M, Dowden S, Laheru D, Bahary N, Ramanathan RK, Tabernero J, Hidalgo M, Goldstein D, Van Cutsem E, Wei X, Iglesias J, Renschler MF (2013) Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N Engl J Med 369(18):1691–1703
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, LeBleu VS, Mittendorf EA, Weitz J, Rahbari N, Reissfelder C, Pilarsky C, Fraga MF, Piwnica-Worms D, Kalluri R (2015) Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523(7559):177–182
Madhavan B, Yue S, Galli U, Rana S, Gross W, Muller M, Giese NA, Kalthoff H, Becker T, Buchler MW, Zoller M (2015) Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer 136(11):2616–2627
Yang S, Che SP, Kurywchak P, Tavormina JL, Gansmo LB, Correa de Sampaio P, Tachezy M, Bockhorn M, Gebauer F, Haltom AR, Melo SA, LeBleu VS, Kalluri R (2017) Detection of mutant KRAS and TP53 DNA in circulating exosomes from healthy individuals and patients with pancreatic cancer. Cancer Biol Ther 18(3):158–165
Karpathakis A, Dibra H, Thirlwell C (2013) Neuroendocrine tumours: cracking the epigenetic code. Endocr Relat Cancer 20(3):R65–R82
Nakhasi HL, Lynch KR, Dolan KP, Unterman RD, Feigelson P (1981) Covalent modification and repressed transcription of a gene in hepatoma cells. Proc Natl Acad Sci U S A 78(2):834–837
Kulis M, Esteller M (2010) DNA methylation and cancer. Adv Genet 70:27–56
Wissmann C, Wild PJ, Kaiser S, Roepcke S, Stoehr R, Woenckhaus M, Kristiansen G, Hsieh J-C, Hofstaedter F, Hartmann A, Knuechel R, Rosenthal A, Pilarsky C (2003) WIF1, a component of the Wnt pathway, is down-regulated in prostate, breast, lung, and bladder cancer. J Pathol 201(2):204–212
Denis H, Ndlovu MN, Fuks F (2011) Regulation of mammalian DNA methyltransferases: a route to new mechanisms. EMBO Rep 12(7):647–656
Hodges E, Smith AD, Kendall J, Xuan Z, Ravi K, Rooks M, Zhang MQ, Ye K, Bhattacharjee A, Brizuela L, McCombie WR, Wigler M, Hannon GJ, Hicks JB (2009) High definition profiling of mammalian DNA methylation by array capture and single molecule bisulfite sequencing. Genome Res 19(9):1593–1605
Dutruel C, Bergmann F, Rooman I, Zucknick M, Weichenhan D, Geiselhart L, Kaffenberger T, Rachakonda PS, Bauer A, Giese N, Hong C, Xie H, Costello JF, Hoheisel J, Kumar R, Rehli M, Schirmacher P, Werner J, Plass C, Popanda O, Schmezer P (2013) Early epigenetic downregulation of WNK2 kinase during pancreatic ductal adenocarcinoma development. Oncogene. https://doi.org/10.1038/onc.2013.312
Sato N, Fukushima N, Hruban RH, Goggins M (2008) CpG island methylation profile of pancreatic intraepithelial neoplasia. Mod Pathol 21(3):238–244
Lofton-Day C, Model F, Devos T, Tetzner R, Distler J, Schuster M, Song X, Lesche R, Liebenberg V, Ebert M, Molnar B, Grützmann R, Pilarsky C, Sledziewski A (2008) DNA methylation biomarkers for blood-based colorectal cancer screening. Clin Chem 54(2):414–423
Tan AC, Jimeno A, Lin SH, Wheelhouse J, Chan F, Solomon A, Rajeshkumar NV, Rubio-Viqueira B, Hidalgo M (2009) Characterizing DNA methylation patterns in pancreatic cancer genome. Mol Oncol 3(5–6):425–438
Murphy PJ, Cipriany BR, Wallin CB, Ju CY, Szeto K, Hagarman JA, Benitez JJ, Craighead HG, Soloway PD (2013) Single-molecule analysis of combinatorial epigenomic states in normal and tumor cells. Proc Natl Acad Sci U S A 110(19):7772–7777
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462(7271):315–322
Stirzaker C, Taberlay PC, Statham AL, Clark SJ (2013) Mining cancer methylomes: prospects and challenges. Trends Genet. https://doi.org/10.1016/j.tig.2013.11.004
Slieker RC, Bos SD, Goeman JJ, Bovée JV, Talens RP, van der Breggen R, Suchiman HED, Lameijer E-W, Putter H, van den Akker EB, Zhang Y, Jukema JW, Slagboom PE, Meulenbelt I, Heijmans BT (2013) Identification and systematic annotation of tissue-specific differentially methylated regions using the Illumina 450k array. Epigenetics Chromatin 6(1):26
Sánchez-Vega F, Gotea V, Petrykowska HM, Margolin G, Krivak TC, Deloia JA, Bell DW, Elnitski L (2013) Recurrent patterns of DNA methylation in the ZNF154, CASP8, and VHL promoters across a wide spectrum of human solid epithelial tumors and cancer cell lines. Epigenetics 8(12)
Morris TJ, Butcher LM, Feber A, Teschendorff AE, Chakravarthy AR, Wojdacz TK, Beck S (2013) ChAMP: 450k chip analysis methylation pipeline. Bioinformatics. https://doi.org/10.1093/bioinformatics/btt684
Wehrum D, Grützmann R, Hennig M, Saeger H-D, Pilarsky C (2008) Recent patents concerning diagnostic and therapeutic applications of aberrantly methylated sequences in pancreatic cancer. Recent Pat DNA Gene Seq 2(2):97–106
Thun M, Jemal A, Desantis C, Blackard B, Ward E (2009) An overview of the cancer burden for primary care physicians. Prim Care 36(3):439–454
Kuroki T, Yendamuri S, Trapasso F, Matsuyama A, Aqeilan RI, Alder H, Rattan S, Cesari R, Nolli ML, Williams NN, Mori M, Kanematsu T, Croce CM (2004) The tumor suppressor gene WWOX at FRA16D is involved in pancreatic carcinogenesis. Clin Cancer Res 10(7):2459–2465
Zhao G, Qin Q, Zhang J, Liu Y, Deng S, Liu L, Wang B, Tian K, Wang C (2013) Hypermethylation of HIC1 promoter and aberrant expression of HIC1/SIRT1 might contribute to the carcinogenesis of pancreatic cancer. Ann Surg Oncol 20(Suppl 3):301–311
Yao F, Sun M, Dong M, Jing F, Chen B, Xu H, Wang S (2013) NPTX2 hypermethylation in pure pancreatic juice predicts pancreatic neoplasms. Am J Med Sci 346(3):175–180
Yang L, Yang H, Li J, Hao J, Qian J (2013) ppENK gene methylation status in the development of pancreatic carcinoma. Gastroenterol Res Pract 2013:130927
Yamamura A, Miura K, Karasawa H, Morishita K, Abe K, Mizuguchi Y, Saiki Y, Fukushige S, Kaneko N, Sase T, Nagase H, Sunamura M, Motoi F, Egawa S, Shibata C, Unno M, Sasaki I, Horii A (2013) Suppressed expression of NDRG2 correlates with poor prognosis in pancreatic cancer. Biochem Biophys Res Commun 441(1):102–107
Wang P, Chen L, Zhang J, Chen H, Fan J, Wang K, Luo J, Chen Z, Meng Z, Liu L (2013) Methylation-mediated silencing of the miR-124 genes facilitates pancreatic cancer progression and metastasis by targeting Rac1. Oncogene. https://doi.org/10.1038/onc.2012.598
Thu KL, Radulovich N, Becker-Santos DD, Pikor LA, Pusic A, Lockwood WW, Lam WL, Tsao M-S (2013) SOX15 is a candidate tumor suppressor in pancreatic cancer with a potential role in Wnt/β-catenin signaling. Oncogene. https://doi.org/10.1038/onc.2012.595
Park JS, Park YN, Lee KY, Kim JK, Yoon DS (2013) P16 hypermethylation predicts surgical outcome following curative resection of mid/distal bile duct cancer. Ann Surg Oncol 20(8):2511–2517
Zhao L, Cui Q, Lu Z, Chen J (2012) Aberrant methylation of RASSF2A in human pancreatic ductal adenocarcinoma and its relation to clinicopathologic features. Pancreas 41(2):206–211
Zhang L, Gao J, Li Z, Gong Y (2012) Neuronal pentraxin II (NPTX2) is frequently down-regulated by promoter hypermethylation in pancreatic cancers. Dig Dis Sci 57(10):2608–2614
Wu Y, Li J, Sun CY, Zhou Y, Zhao YF, Zhang SJ (2012) Epigenetic inactivation of the canonical Wnt antagonist secreted frizzled-related protein 1 in hepatocellular carcinoma cells. Neoplasma 59(3):326–332
Li M, Zhao ZW (2012) Clinical implications of mismatched repair gene promoter methylation in pancreatic cancer. Med Oncol 29(2):970–976
Hong S-M, Omura N, Vincent A, Li A, Knight S, Yu J, Hruban RH, Goggins M (2012) Genome-wide CpG island profiling of intraductal papillary mucinous neoplasms of the pancreas. Clin Cancer Res 18(3):700–712
Heichman KA, Warren JD (2012) DNA methylation biomarkers and their utility for solid cancer diagnostics. Clin Chem Lab Med 50(10):1707–1721
Giovannetti E, Erozenci A, Smit J, Danesi R, Peters GJ (2012) Molecular mechanisms underlying the role of microRNAs (miRNAs) in anticancer drug resistance and implications for clinical practice. Crit Rev Oncol Hematol 81(2):103–122
Park JK, Ryu JK, Yoon WJ, Lee SH, Lee GY, Jeong K-S, Kim Y-T, Yoon YB (2012) The role of quantitative NPTX2 hypermethylation as a novel serum diagnostic marker in pancreatic cancer. Pancreas 41(1):95–101
Mishra NK, Guda C (2017) Genome-wide DNA methylation analysis reveals molecular subtypes of pancreatic cancer. Oncotarget. https://doi.org/10.18632/oncotarget.15993
Thompson MJ, Rubbi L, Dawson DW, Donahue TR, Pellegrini M (2015) Pancreatic cancer patient survival correlates with DNA methylation of pancreas development genes. PLoS One 10(6):e0128814
Biewusch K, Heyne M, Grützmann R, Pilarsky C (2012) DNA methylation in pancreatic cancer: protocols for the isolation of DNA and bisulfite modification. Methods Mol Biol 863:273–280
Resnick RM, Cornelissen MT, Wright DK, Eichinger GH, Fox HS, ter Schegget J, Manos MM (1990) Detection and typing of human papillomavirus in archival cervical cancer specimens by DNA amplification with consensus primers. J Natl Cancer Inst 82(18):1477–1484
Acknowledgments
Thanks to Alfred E. Neumann for fruitful discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Liu, B., Pilarsky, C. (2018). Analysis of DNA Hypermethylation in Pancreatic Cancer Using Methylation-Specific PCR and Bisulfite Sequencing. In: Dumitrescu, R., Verma, M. (eds) Cancer Epigenetics for Precision Medicine . Methods in Molecular Biology, vol 1856. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8751-1_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8751-1_16
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8750-4
Online ISBN: 978-1-4939-8751-1
eBook Packages: Springer Protocols