Abstract
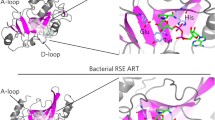
The ARH family of ADP-ribosyl-acceptor hydrolases is composed of three 39-kDa proteins (ARH1, 2, and 3), which hydrolyze specific ADP-ribosylated substrates. ARH1 hydrolyzes mono(ADP-ribosyl)ated arginine, which results from actions of cholera toxin and other nicotinamide adenine dinucleotide (NAD+):arginine ADP-ribosyl-transferases, while ARH3 hydrolyzes poly(ADP-ribose) and O-acetyl-ADP-ribose, resulting from the action of poly(ADP-ribose) polymerases and sirtuins, respectively. ARH2 has not been reported to have enzymatic activity, because of differences in the catalytic domain. Thus, the substrate specificities of ARH1 and ARH3 proteins result in unique cellular functions. In this chapter, we introduce several methods to monitor the activities of the ARH family members.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Verheugd P, Butepage M, Eckei L, Luscher B (2016) Players in ADP-ribosylation: readers and erasers. Curr Protein Pept Sci 17(7):654–667
Palazzo L, Mikoc A, Ahel I (2017) ADP-ribosylation: new facets of an ancient modification. FEBS J 284:2932. https://doi.org/10.1111/febs.14078
Okazaki IJ, Moss J (1996) Mono-ADP-ribosylation: a reversible posttranslational modification of proteins. Adv Pharmacol 35:247–280
Corda D, Di Girolamo M (2003) Functional aspects of protein mono-ADP-ribosylation. EMBO J 22(9):1953–1958. https://doi.org/10.1093/emboj/cdg209
Bütepage M, Eckei L, Verheugd P, Luscher B (2015) Intracellular mono-ADP-ribosylation in signaling and disease. Cell 4(4):569–595. https://doi.org/10.3390/cells4040569
Ogata N, Ueda K, Kagamiyama H, Hayaishi O (1980) ADP-ribosylation of histone H1. Identification of glutamic acid residues 2, 14, and the COOH-terminal lysine residue as modification sites. J Biol Chem 255(16):7616–7620
Altmeyer M, Messner S, Hassa PO, Fey M, Hottiger MO (2009) Molecular mechanism of poly(ADP-ribosyl)ation by PARP1 and identification of lysine residues as ADP-ribose acceptor sites. Nucleic Acids Res 37(11):3723–3738. https://doi.org/10.1093/nar/gkp229
Zhang Y, Wang J, Ding M, Yu Y (2013) Site-specific characterization of the Asp- and Glu-ADP-ribosylated proteome. Nat Methods 10(10):981–984. https://doi.org/10.1038/nmeth.2603
Leidecker O, Bonfiglio JJ, Colby T, Zhang Q, Atanassov I, Zaja R, Palazzo L, Stockum A, Ahel I, Matic I (2016) Serine is a new target residue for endogenous ADP-ribosylation on histones. Nat Chem Biol 12(12):998–1000. https://doi.org/10.1038/nchembio.2180
Iglewski BH, Kabat D (1975) NAD-dependent inhibition of protein synthesis by Pseudomonas aeruginosa toxin. Proc Natl Acad Sci U S A 72(6):2284–2288
Pappenheimer AM Jr (1977) Diphtheria toxin. Annu Rev Biochem 46:69–94. https://doi.org/10.1146/annurev.bi.46.070177.000441
Cassel D, Pfeuffer T (1978) Mechanism of cholera toxin action: covalent modification of the guanyl nucleotide-binding protein of the adenylate cyclase system. Proc Natl Acad Sci U S A 75(6):2669–2673
Tamura M, Nogimori K, Murai S, Yajima M, Ito K, Katada T, Ui M, Ishii S (1982) Subunit structure of islet-activating protein, pertussis toxin, in conformity with the A-B model. Biochemistry 21(22):5516–5522
Vyas S, Chesarone-Cataldo M, Todorova T, Huang YH, Chang P (2013) A systematic analysis of the PARP protein family identifies new functions critical for cell physiology. Nat Commun 4:2240. https://doi.org/10.1038/ncomms3240
Bock FJ, Chang P (2016) New directions in poly(ADP-ribose) polymerase biology. FEBS J 283(22):4017–4031. https://doi.org/10.1111/febs.13737
Gupte R, Liu Z, Kraus WL (2017) PARPs and ADP-ribosylation: recent advances linking molecular functions to biological outcomes. Genes Dev 31(2):101–126. https://doi.org/10.1101/gad.291518.116
Borra MT, O'Neill FJ, Jackson MD, Marshall B, Verdin E, Foltz KR, Denu JM (2002) Conserved enzymatic production and biological effect of O-acetyl-ADP-ribose by silent information regulator 2-like NAD+-dependent deacetylases. J Biol Chem 277(15):12632–12641. https://doi.org/10.1074/jbc.M111830200
Liou GG, Tanny JC, Kruger RG, Walz T, Moazed D (2005) Assembly of the SIR complex and its regulation by O-acetyl-ADP-ribose, a product of NAD-dependent histone deacetylation. Cell 121(4):515–527. https://doi.org/10.1016/j.cell.2005.03.035
Grubisha O, Rafty LA, Takanishi CL, Xu X, Tong L, Perraud AL, Scharenberg AM, Denu JM (2006) Metabolite of SIR2 reaction modulates TRPM2 ion channel. J Biol Chem 281(20):14057–14065. https://doi.org/10.1074/jbc.M513741200
Moss J, Jacobson MK, Stanley SJ (1985) Reversibility of arginine-specific mono(ADP-ribosyl)ation: identification in erythrocytes of an ADP-ribose-L-arginine cleavage enzyme. Proc Natl Acad Sci U S A 82(17):5603–5607
Oka S, Kato J, Moss J (2006) Identification and characterization of a mammalian 39-kDa poly(ADP-ribose) glycohydrolase. J Biol Chem 281(2):705–713. https://doi.org/10.1074/jbc.M510290200
Mashimo M, Kato J, Moss J (2014) Structure and function of the ARH family of ADP-ribosyl-acceptor hydrolases. DNA Repair (Amst) 23:88–94. https://doi.org/10.1016/j.dnarep.2014.03.005
Konczalik P, Moss J (1999) Identification of critical, conserved vicinal aspartate residues in mammalian and bacterial ADP-ribosylarginine hydrolases. J Biol Chem 274(24):16736–16740
Ono T, Kasamatsu A, Oka S, Moss J (2006) The 39-kDa poly(ADP-ribose) glycohydrolase ARH3 hydrolyzes O-acetyl-ADP-ribose, a product of the Sir2 family of acetyl-histone deacetylases. Proc Natl Acad Sci U S A 103(45):16687–16691. https://doi.org/10.1073/pnas.0607911103
Moss J, Tsai SC, Adamik R, Chen HC, Stanley SJ (1988) Purification and characterization of ADP-ribosylarginine hydrolase from Turkey erythrocytes. Biochemistry 27(15):5819–5823
Moss J, Balducci E, Cavanaugh E, Kim HJ, Konczalik P, Lesma EA, Okazaki IJ, Park M, Shoemaker M, Stevens LA, Zolkiewska A (1999) Characterization of NAD:arginine ADP-ribosyltransferases. Mol Cell Biochem 193(1–2):109–113
Freissmuth M, Gilman AG (1989) Mutations of GS alpha designed to alter the reactivity of the protein with bacterial toxins. Substitutions at ARG187 result in loss of GTPase activity. J Biol Chem 264(36):21907–21914
Gill DM, Meren R (1978) ADP-ribosylation of membrane proteins catalyzed by cholera toxin: basis of the activation of adenylate cyclase. Proc Natl Acad Sci U S A 75(7):3050–3054
Kahn RA, Gilman AG (1984) Purification of a protein cofactor required for ADP-ribosylation of the stimulatory regulatory component of adenylate cyclase by cholera toxin. J Biol Chem 259(10):6228–6234
Kahn RA, Gilman AG (1986) The protein cofactor necessary for ADP-ribosylation of Gs by cholera toxin is itself a GTP binding protein. J Biol Chem 261(17):7906–7911
Kato J, Zhu J, Liu C, Moss J (2007) Enhanced sensitivity to cholera toxin in ADP-ribosylarginine hydrolase-deficient mice. Mol Cell Biol 27(15):5534–5543. https://doi.org/10.1128/MCB.00302-07
Kato J, Zhu J, Liu C, Stylianou M, Hoffmann V, Lizak MJ, Glasgow CG, Moss J (2011) ADP-ribosylarginine hydrolase regulates cell proliferation and tumorigenesis. Cancer Res 71(15):5327–5335. https://doi.org/10.1158/0008-5472.CAN-10-0733
Kato J, Vekhter D, Heath J, Zhu J, Barbieri JT, Moss J (2015) Mutations of the functional ARH1 allele in tumors from ARH1 heterozygous mice and cells affect ARH1 catalytic activity, cell proliferation and tumorigenesis. Oncogene 4:e151. https://doi.org/10.1038/oncsis.2015.5
Kasamatsu A, Nakao M, Smith BC, Comstock LR, Ono T, Kato J, Denu JM, Moss J (2011) Hydrolysis of O-acetyl-ADP-ribose isomers by ADP-ribosylhydrolase 3. J Biol Chem 286(24):21110–21117. https://doi.org/10.1074/jbc.M111.237636
Mashimo M, Kato J, Moss J (2013) ADP-ribosyl-acceptor hydrolase 3 regulates poly (ADP-ribose) degradation and cell death during oxidative stress. Proc Natl Acad Sci U S A 110(47):18964–18969. https://doi.org/10.1073/pnas.1312783110
Fontana P, Bonfiglio JJ, Palazzo L, Bartlett E, Matic I, Ahel I (2017) Serine ADP-ribosylation reversal by the hydrolase ARH3. elife 6. https://doi.org/10.7554/eLife.28533
Mashimo M, Moss J (2016) Functional role of ADP-Ribosyl-acceptor hydrolase 3 in poly(ADP-ribose) polymerase-1 response to oxidative stress. Curr Protein Pept Sci 17(7):633–640
Yu SW, Wang H, Poitras MF, Coombs C, Bowers WJ, Federoff HJ, Poirier GG, Dawson TM, Dawson VL (2002) Mediation of poly(ADP-ribose) polymerase-1-dependent cell death by apoptosis-inducing factor. Science 297(5579):259–263. https://doi.org/10.1126/science.1072221
Andrabi SA, Kang HC, Haince JF, Lee YI, Zhang J, Chi Z, West AB, Koehler RC, Poirier GG, Dawson TM, Dawson VL (2011) Iduna protects the brain from glutamate excitotoxicity and stroke by interfering with poly(ADP-ribose) polymer-induced cell death. Nat Med 17(6):692–699. https://doi.org/10.1038/nm.2387
Lee Y, Karuppagounder SS, Shin JH, Lee YI, Ko HS, Swing D, Jiang H, Kang SU, Lee BD, Kang HC, Kim D, Tessarollo L, Dawson VL, Dawson TM (2013) Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss. Nat Neurosci 16(10):1392–1400. https://doi.org/10.1038/nn.3500
Wang Y, An R, Umanah GK, Park H, Nambiar K, Eacker SM, Kim B, Bao L, Harraz MM, Chang C, Chen R, Wang JE, Kam TI, Jeong JS, Xie Z, Neifert S, Qian J, Andrabi SA, Blackshaw S, Zhu H, Song H, Ming GL, Dawson VL, Dawson TM (2016) A nuclease that mediates cell death induced by DNA damage and poly(ADP-ribose) polymerase-1. Science 354(6308):aad6872. https://doi.org/10.1126/science.aad6872
Andrabi SA, Umanah GK, Chang C, Stevens DA, Karuppagounder SS, Gagne JP, Poirier GG, Dawson VL, Dawson TM (2014) Poly(ADP-ribose) polymerase-dependent energy depletion occurs through inhibition of glycolysis. Proc Natl Acad Sci U S A 111(28):10209–10214. https://doi.org/10.1073/pnas.1405158111
Niere M, Mashimo M, Agledal L, Dolle C, Kasamatsu A, Kato J, Moss J, Ziegler M (2012) ADP-ribosylhydrolase 3 (ARH3), not poly(ADP-ribose) glycohydrolase (PARG) isoforms, is responsible for degradation of mitochondrial matrix-associated poly(ADP-ribose). J Biol Chem 287(20):16088–16102. https://doi.org/10.1074/jbc.M112.349183
Parihar P, Solanki I, Mansuri ML, Parihar MS (2015) Mitochondrial sirtuins: emerging roles in metabolic regulations, energy homeostasis and diseases. Exp Gerontol 61:130–141. https://doi.org/10.1016/j.exger.2014.12.004
Osborne B, Bentley NL, Montgomery MK, Turner N (2016) The role of mitochondrial sirtuins in health and disease. Free Radic Biol Med 100:164–174. https://doi.org/10.1016/j.freeradbiomed.2016.04.197
Alvarez-Gonzalez R, Juarez-Salinas H, Jacobson EL, Jacobson MK (1983) Evaluation of immobilized boronates for studies of adenine and pyridine nucleotide metabolism. Anal Biochem 135(1):69–77
Acknowledgment
Funding: The study was funded by the Intramural Research Program, NIH, NHLBI.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Mashimo, M., Moss, J. (2018). ADP-Ribosyl-Acceptor Hydrolase Activities Catalyzed by the ARH Family of Proteins. In: Chang, P. (eds) ADP-ribosylation and NAD+ Utilizing Enzymes. Methods in Molecular Biology, vol 1813. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-8588-3_12
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8588-3_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-8587-6
Online ISBN: 978-1-4939-8588-3
eBook Packages: Springer Protocols