Abstract
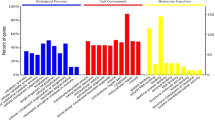
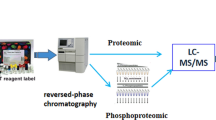
Proteomics is a technology that allows to decipher the molecular networks involved in the regulation of biological processes such as spermatogenesis. Sertoli cells (SCs) are key players in the paracrine control of this process. Envisioning to increase the knowledge on the molecular networks harbored in SCs, we propose a methodology based on GeLC-MS/MS for the characterization of these cells’ proteome. Proteins are separated by SDS-PAGE hyphenated to HPLC and identified by mass spectrometry. The integration of data with bioinformatics tools such as ClueGO + CluePedia from Cytoscape allows the identification of the biological pathways more prevalent in SCs, and that might be modulated by pathophysiological conditions. Moreover, the proteome analysis with tools as SignalP/SecretomeP highlights the proteins more prone to be secreted and involved in the paracrine control of germ cells, which might also be deregulated by diseases.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Li X, Wang W, Chen J (2015) From pathways to networks: connecting dots by establishing protein-protein interaction networks in signaling pathways using affinity purification and mass spectrometry. Proteomics 15(2–3):188–202. https://doi.org/10.1002/pmic.201400147
Zheng B et al (2015) Quantitative proteomics reveals the essential roles of stromal interaction molecule 1 (STIM1) in the testicular cord formation in mouse testis. Mol Cell Proteomics 14(10):2682–2691. https://doi.org/10.1074/mcp.M115.049569
Jegou B (1993) The Sertoli-germ cell communication network in mammals. Int Rev Cytol 147:25–96
Stanton PG, Sluka P, Foo CF, Stephens AN, Smith AI, McLachlan RI, O'Donnell L (2012) Proteomic changes in rat spermatogenesis in response to in vivo androgen manipulation; impact on meiotic cells. PLoS One 7(7):e41718. https://doi.org/10.1371/journal.pone.0041718
Chalmel F, Com E, Lavigne R, Hernio N, Teixeira-Gomes AP, Dacheux JL, Pineau C (2014) An integrative omics strategy to assess the germ cell secretome and to decipher sertoli-germ cell crosstalk in the Mammalian testis. PLoS One 9(8):e104418. https://doi.org/10.1371/journal.pone.0104418
Com E, Melaine N, Chalmel F, Pineau C (2014) Proteomics and integrative genomics for unraveling the mysteries of spermatogenesis: the strategies of a team. J Proteome 107:128–143. https://doi.org/10.1016/j.jprot.2014.04.013
Rato L, Alves MG, Socorro S, Duarte AI, Cavaco JE, Oliveira PF (2012) Metabolic regulation is important for spermatogenesis. Nat Rev Urol 9(6):330–338. https://doi.org/10.1038/nrurol.2012.77
Yates JR, Ruse CI, Nakorchevsky A (2009) Proteomics by mass spectrometry: approaches, advances, and applications. Annu Rev Biomed Eng 11:49–79. https://doi.org/10.1146/annurev-bioeng-061008-124934
Padrao AI, Vitorino R, Duarte JA, Ferreira R, Amado F (2013) Unraveling the phosphoproteome dynamics in mammal mitochondria from a network perspective. J Proteome Res 12(10):4257–4267. https://doi.org/10.1021/pr4003917
Aslam B, Basit M, Nisar MA, Khurshid M, Rasool MH (2017) Proteomics: technologies and their applications. J Chromatogr Sci 55(2):182–196. https://doi.org/10.1093/chromsci/bmw167
Paulo JA (2016) Sample preparation for proteomic analysis using a GeLC-MS/MS strategy. J Biol Methods 3(3). https://doi.org/10.14440/jbm.2016.106
Chen C, Huang H, Wu CH (2011) Protein bioinformatics databases and resources. Methods Mol Biol 694:3–24. https://doi.org/10.1007/978-1-60761-977-2_1
Bindea G, Galon J, Mlecnik B (2013) CluePedia Cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics. https://doi.org/10.1093/bioinformatics/btt019
Bindea G et al (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25(8):1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Bendtsen JD, Jensen LJ, Blom N, Von Heijne G, Brunak S (2004) Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng Des Sel 17(4):349–356. https://doi.org/10.1093/protein/gzh037
Bindea G, Galon J, Mlecnik B (2013) CluePedia Cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 29(5):661–663. https://doi.org/10.1093/bioinformatics/btt019
Palmero S, Bardi G, Coniglio L, Falugi C (1999) Presence and localization of molecules related to the cholinergic system in developing rat testis. Eur J Histochem 43(4):277–283
Matzuk MM, Finegold MJ, Su JG, Hsueh AJ, Bradley A (1992) Alpha-inhibin is a tumour-suppressor gene with gonadal specificity in mice. Nature 360(6402):313–319. https://doi.org/10.1038/360313a0
Tripathi UK et al (2014) Differential proteomic profile of spermatogenic and Sertoli cells from peri-pubertal testes of three different bovine breeds. Front Cell Dev Biol 2:24. https://doi.org/10.3389/fcell.2014.00024
Carvalho PC et al (2016) Integrated analysis of shotgun proteomic data with PatternLab for proteomics 4.0. Nat Protoc 11(1):102–117. https://doi.org/10.1038/nprot.2015.133
Killcoyne S, Carter GW, Smith J, Boyle J (2009) Cytoscape: a community-based framework for network modeling. Methods Mol Biol 563:219–239. https://doi.org/10.1007/978-1-60761-175-2_12
Acknowledgments
The authors thank Portuguese Foundation for Science and Technology (FCT), European Union, QREN, FEDER, and COMPETE for funding the iBiMED (UID/BIM/04501/2013), QOPNA (UID/QUI/00062/2013), UnIC (UID/IC/00051/2013), research project (POCI-01-0145-FEDER-016728; PTDC/DTP-DES/6077/2014), and RV’s (IF/00286/2015) and FT’s (SFRH/BD/111633/2015) fellowship grants.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Materials
Supplementary Table S1
List of proteins previously identified in mouse SCs [5] (XLSX 29 kb)
Supplementary Table S2
List of proteins retrieved from SecretomeP and SignalP analysis (XLSX 635 kb)
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Ferreira, R., Trindade, F., Vitorino, R. (2018). Proteome Profiling of Sertoli Cells Using a GeLC-MS/MS Strategy. In: Alves, M., Oliveira, P. (eds) Sertoli Cells. Methods in Molecular Biology, vol 1748. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7698-0_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7698-0_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7697-3
Online ISBN: 978-1-4939-7698-0
eBook Packages: Springer Protocols