Abstract
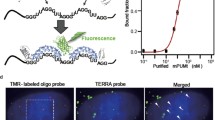
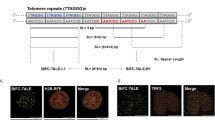
Visualizing RNA in living cells is increasingly important to facilitate accumulation of knowledge about the relation between specific RNA dynamics and physiological events. Single-molecule fluorescence imaging of target RNAs is an excellent approach to analyzing intracellular RNA motion, but it requires special techniques for probe design and microscope setup. Herein, we present a principle and protocol of an RNA visualization probe based on an RNA binding protein of the Pumilio homology domain (PUM-HD). We also describe the setup and operation of a microscope, and introduce an application to visualize telomeric repeats-containing RNA with telomeres and a telomere-related protein: hnRNPA1. This imaging technique is applicable to visualization of different RNAs, especially including repetitive sequences, in living cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Martin KC, Ephrussi A (2009) mRNA localization: gene expression in the spatial dimension. Cell 136(4):719–730. doi:10.1016/j.cell.2009.01.044. S0092-8674(09)00126-3 [pii]
Buxbaum AR, Haimovich G, Singer RH (2015) In the right place at the right time: visualizing and understanding mRNA localization. Nat Rev Mol Cell Biol 16(2):95–109. doi:10.1038/nrm3918. http://www.nature.com/nrm/journal/v16/n2/abs/nrm3918.html#supplementary-information
Geisler S, Coller J (2013) RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 14(11):699–712. doi:10.1038/nrm3679
Heward JA, Lindsay MA (2014) Long non-coding RNAs in the regulation of the immune response. Trends Immunol 35(9):408–419. doi:10.1016/j.it.2014.07.005
Engreitz Jesse M, Sirokman K, McDonel P, Shishkin AA, Surka C, Russell P, Grossman Sharon R, Chow Amy Y, Guttman M, Lander Eric S (2014) RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell 159(1):188–199. doi:10.1016/j.cell.2014.08.018
Oomoto I, Suzuki-Hirano A, Umeshima H, Han Y-W, Yanagisawa H, Carlton P, Harada Y, Kengaku M, Okamoto A, Shimogori T, Wang DO (2015) ECHO-liveFISH: in vivo RNA labeling reveals dynamic regulation of nuclear RNA foci in living tissues. Nucleic Acids Res. doi:10.1093/nar/gkv614
Wang X, McLachlan J, Zamore PD, Hall TMT (2002) Modular recognition of RNA by a human pumilio-homology domain. Cell 110(4):501–512. doi:10.1016/S0092-8674(02)00873-5
Cheong CG, Hall TMT (2006) Engineering RNA sequence specificity of pumilio repeats. Proc Natl Acad Sci U S A 103(37):13635–13639. doi:10.1073/pnas.0606294103
Yamada T, Yoshimura H, Inaguma A, Ozawa T (2011) Visualization of nonengineered single mRNAs in living cells using genetically encoded fluorescent probes. Anal Chem 83(14):5708–5714. doi:10.1021/ac2009405
Yoshimura H, Inaguma A, Yamada T, Ozawa T (2012) Fluorescent probes for imaging endogenous beta-actin mRNA in living cells using fluorescent protein-tagged pumilio. ACS Chem Biol 7(6):999–1005. doi:10.1021/cb200474a
Yoshimura H, Ozawa T (2016) Chapter 3: Monitoring of RNA dynamics in living cells using PUM-HD and fluorescent protein reconstitution technique. In: Grigory SF, Samie RJ (eds) Methods in enzymology, vol 572. Academic Press, New York, pp 65–85. doi:10.1016/bs.mie.2016.03.018
Yamada T, Yoshimura H, Shimada R, Hattori M, Eguchi M, Fujiwara TK, Kusumi A, Ozawa T (2016) Spatiotemporal analysis with a genetically encoded fluorescent RNA probe reveals TERRA function around telomeres. Sci Rep 6:38910. doi:10.1038/srep38910
Green KM, Linsalata AE, Todd PK (2016) RAN translation—what makes it run? Brain Res 1647:30–42. doi:10.1016/j.brainres.2016.04.003
Matsui M, Corey DR (2016) Non-coding RNAs as drug targets. Nat Rev Drug Discov. Advance online publication. doi:10.1038/nrd.2016.117
Faulkner GJ, Kimura Y, Daub CO, Wani S, Plessy C, Irvine KM, Schroder K, Cloonan N, Steptoe AL, Lassmann T, Waki K, Hornig N, Arakawa T, Takahashi H, Kawai J, Forrest ARR, Suzuki H, Hayashizaki Y, Hume DA, Orlando V, Grimmond SM, Carninci P (2009) The regulated retrotransposon transcriptome of mammalian cells. Nat Genet 41(5):563–571. http://www.nature.com/ng/journal/v41/n5/suppinfo/ng.368_s1.html
Vardy L, Orr-Weaver TL (2007) Regulating translation of maternal messages: multiple repression mechanisms. Trends Cell Biol 17(11):547–554. doi:10.1016/j.tcb.2007.09.002
Filipovska A, Razif MF, Nygard KK, Rackham O (2011) A universal code for RNA recognition by PUF proteins. Nat Chem Biol 7(7):425–427. doi:10.1038/nchembio.577
Ozawa T, Yoshimura H, Kim SB (2013) Advances in fluorescence and bioluminescence imaging. Anal Chem 85(2):590–609. doi:10.1021/ac3031724
Yoshimura H, Ozawa T (2014) Methods of split reporter reconstitution for the analysis of biomolecules. Chem Rec 14(3):492–501. doi:10.1002/tcr.201402001
Magliery TJ, Wilson CGM, Pan W, Mishler D, Ghosh I, Hamilton AD, Regan L (2005) Detecting protein−protein interactions with a green fluorescent protein fragment reassembly trap: scope and mechanism. J Am Chem Soc 127(1):146–157. doi:10.1021/ja046699g
Ozawa T, Nogami S, Sato M, Ohya Y, Umezawa Y (2000) A fluorescent indicator for detecting protein−protein interactions in vivo based on protein splicing. Anal Chem 72(21):5151–5157. doi:10.1021/ac000617z
Tokunaga M, Imamoto N, Sakata-Sogawa K (2008) Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat Methods 5(2):159–161. doi:10.1038/nmeth1171
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Yoshimura, H., Ozawa, T. (2018). Real-Time Fluorescence Imaging of Single-Molecule Endogenous Noncoding RNA in Living Cells. In: Gaspar, I. (eds) RNA Detection. Methods in Molecular Biology, vol 1649. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7213-5_22
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7213-5_22
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7212-8
Online ISBN: 978-1-4939-7213-5
eBook Packages: Springer Protocols