Abstract
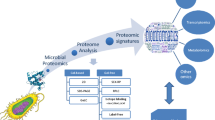
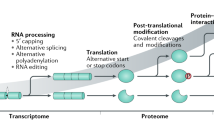
Posttranslational modifications (PTMs) of proteins are an integral part of major cellular regulatory mechanisms dictating protein function, localization, and stability. The capacity to screen PTMs using protein microarrays has advanced our ability to identify their targets and regulatory role. This chapter discusses a unique procedure that combines functional extract-based activity assay with large-scale screening utilities of protein microarrays. This “PTM-profiling” system offers advantages in quantitatively identifying modifications in an unbiased manner in the context of specific cellular conditions. While the possibilities of studying PTMs in different settings are enormous, the immune system presents an attractive model for studying the effects of perturbations in PTMs, and specifically the ubiquitin system, as these were already implicated in both immune function and dysfunction. This chapter discusses the significance of PTM profiling in addressing basic questions in immunology. We describe detailed protocols for the preparation of functional cell extracts from immune cell cultures, following differentiation or induced signals, and screening PTMs on protein arrays, as well as basic guidelines for data analysis and interpretation.
Similar content being viewed by others
References
Merbl Y, Refour P, Patel H et al (2013) Profiling of ubiquitin-like modifications reveals features of mitotic control. Cell 152:1160–1172. doi:10.1016/j.cell.2013.02.007
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479. doi:10.1146/annurev.biochem.67.1.425
Park Y, Jin H, Aki D et al (2014) The ubiquitin system in immune regulation. Adv Immunol 124:17–66. doi:10.1016/B978-0-12-800147-9.00002-9
Wang J, Maldonado MA (2006) The ubiquitin-proteasome system and its role in inflammatory and autoimmune diseases. Cell Mol Immunol 3:255–261
Liu Y-C (2004) Ubiquitin ligases and the immune response. Annu Rev Immunol 22:81–127. doi:10.1146/annurev.immunol.22.012703.104813
Davis ME, Gack MU (2015) Ubiquitination in the antiviral immune response. Virology 479-480:52–65. doi:10.1016/j.virol.2015.02.033
Gómez-Martín D, Díaz-Zamudio M, Alcocer-Varela J (2008) Ubiquitination system and autoimmunity: the bridge towards the modulation of the immune response. Autoimmun Rev 7:284–290. doi:10.1016/j.autrev.2007.11.026
Ciechanover A, Schwartz AL (2004) The ubiquitin system: pathogenesis of human diseases and drug targeting. Biochim Biophys Acta 1695:3–17. doi:10.1016/j.bbamcr.2004.09.018
Komander D, Rape M (2012) The ubiquitin code. Annu Rev Biochem 81:203–229. doi:10.1146/annurev-biochem-060310-170328
Kirkin V, Dikic I (2007) Role of ubiquitin- and Ubl-binding proteins in cell signaling. Curr Opin Cell Biol 19:199–205. doi:10.1016/j.ceb.2007.02.002
Yau R, Rape M (2016) The increasing complexity of the ubiquitin code. Nat Cell Biol 18:579–586. doi:10.1038/ncb3358
Komander D, Clague MJ, Urbé S (2009) Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol 10:550–563. doi:10.1038/nrm2731
Popovic D, Vucic D, Dikic I (2014) Ubiquitination in disease pathogenesis and treatment. Nat Med 20:1242–1253. doi:10.1038/nm.3739
Schartner JM, Fathman CG, Seroogy CM (2007) Preservation of self: an overview of E3 ubiquitin ligases and T cell tolerance. Semin Immunol 19:188–196. doi:10.1016/j.smim.2007.02.010
Liu Y-C, Penninger J, Karin M (2005) Immunity by ubiquitylation: a reversible process of modification. Nat Rev Immunol 5:941–952. doi:10.1038/nri1731
Chen ZJ (2005) Ubiquitin signalling in the NF-κB pathway. Nat Cell Biol 7:758–765. doi:10.1038/ncb0805-758
Smahi A, Courtois G, Vabres P et al (2000) Genomic rearrangement in NEMO impairs NF-kappaB activation and is a cause of incontinentia pigmenti. The international Incontinentia Pigmenti (IP) consortium. Nature 405:466–472. doi:10.1038/35013114
Döffinger R, Smahi A, Bessia C et al (2001) X-linked anhidrotic ectodermal dysplasia with immunodeficiency is caused by impaired NF-kappaB signaling. Nat Genet 27:277–285. doi:10.1038/85837
Orange JS, Brodeur SR, Jain A et al (2002) Deficient natural killer cell cytotoxicity in patients with IKK-gamma/NEMO mutations. J Clin Invest 109:1501–1509. doi:10.1172/JCI14858
Majumdar I, Paul J (2014) The deubiquitinase A20 in immunopathology of autoimmune diseases. Autoimmunity 47(5):307–319. doi:10.3109/08916934.2014.900756
Ma A, Malynn BA (2012) A20: linking a complex regulator of ubiquitylation to immunity and human disease. Nat Rev Immunol 12:774–785. doi:10.1038/nri3313
Compagno M, Lim WK, Grunn A et al (2009) Mutations of multiple genes cause deregulation of NF-kappaB in diffuse large B-cell lymphoma. Nature 459:717–721. doi:10.1038/nature07968
Kato M, Sanada M, Kato I et al (2009) Frequent inactivation of A20 in B-cell lymphomas. Nature 459:712–716. doi:10.1038/nature07969
Novak U, Rinaldi A, Kwee I et al (2009) The NF-{kappa}B negative regulator TNFAIP3 (A20) is inactivated by somatic mutations and genomic deletions in marginal zone lymphomas. Blood 113:4918–4921. doi:10.1182/blood-2008-08-174110
Schmitz R, Stanelle J, Hansmann M-L, Küppers R (2009) Pathogenesis of classical and lymphocyte-predominant Hodgkin lymphoma. Annu Rev Pathol 4:151–174. doi:10.1146/annurev.pathol.4.110807.092209
Schwartz RH (2003) T cell anergy. Annu Rev Immunol 21:305–334. doi:10.1146/annurev.immunol.21.120601.141110
Bachmaier K, Krawczyk C, Kozieradzki I et al (2000) Negative regulation of lymphocyte activation and autoimmunity by the molecular adaptor Cbl-b. Nature 403:211–216. doi:10.1038/35003228
Paolino M, Thien CBF, Gruber T et al (2011) Essential role of E3 ubiquitin ligase activity in Cbl-b-regulated T cell functions. J Immunol 186:2138–2147. doi:10.4049/jimmunol.1003390
Jeon M-S, Atfield A, Venuprasad K et al (2004) Essential role of the E3 ubiquitin ligase Cbl-b in T cell Anergy induction. Immunity 21:167–177. doi:10.1016/j.immuni.2004.07.013
Gómez-Martín D, Ibarra-Sánchez M, Romo-Tena J et al (2013) Casitas B lineage lymphoma b is a key regulator of peripheral tolerance in systemic lupus erythematosus. Arthritis Rheum 65:1032–1042. doi:10.1002/art.37833
Doníz-Padilla L, Martínez-Jiménez V, Niño-Moreno P et al (2011) Expression and function of Cbl-b in T cells from patients with systemic lupus erythematosus, and detection of the 2126 a/G Cblb gene polymorphism in the Mexican mestizo population. Lupus 20:628–635. doi:10.1177/0961203310394896
Stürner KH, Borgmeyer U, Schulze C et al (2014) A multiple sclerosis-associated variant of CBLB links genetic risk with type I IFN function. J Immunol 193:4439–4447. doi:10.4049/jimmunol.1303077
Anandasabapathy N, Ford GS, Bloom D et al (2003) GRAIL: an E3 ubiquitin ligase that inhibits cytokine Gene transcription is expressed in Anergic CD4+ T cells. Immunity 18:535–547. doi:10.1016/S1074-7613(03)00084-0
Heissmeyer V, Macián F, Im S-H et al (2004) Calcineurin imposes T cell unresponsiveness through targeted proteolysis of signaling proteins. Nat Immunol 5:255–265. doi:10.1038/ni1047
Sande OJ, Karim AF, Li Q et al (2016) Mannose-capped Lipoarabinomannan from mycobacterium tuberculosis induces CD4+ T cell Anergy via GRAIL. J Immunol 196:691–702. doi:10.4049/jimmunol.1500710
Gu H, Chiang YJ, Kole HK et al (2000) Cbl-b regulates the CD28 dependence of T-cell activation. Nature 403:216–220. doi:10.1038/35003235
Lohr NJ, Molleston JP, Strauss KA et al (2010) Human ITCH E3 ubiquitin ligase deficiency causes syndromic multisystem autoimmune disease. Am J Hum Genet 86:447–453. doi:10.1016/j.ajhg.2010.01.028
Williamson A, Jin L, Rape M (2009) Preparation of synchronized human cell extracts to study ubiquitination and degradation. Methods Mol Biol 545:301–312. doi:10.1007/978-1-60327-993-2_19
Acknowledgments
Yifat Merbl is supported by the I-CORE Program of the Planning and Budgeting Committee and The Israel Science Foundation (grant No. 1775/12), and by the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation programme.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Eisenberg-Lerner, A., Regev, I., Merbl, Y. (2017). Post-Translational Modification Profiling-Functional Proteomics for the Analysis of Immune Regulation. In: Lazar, I., Kontoyianni, M., Lazar, A. (eds) Proteomics for Drug Discovery. Methods in Molecular Biology, vol 1647. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7201-2_9
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7201-2_9
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7200-5
Online ISBN: 978-1-4939-7201-2
eBook Packages: Springer Protocols