Abstract
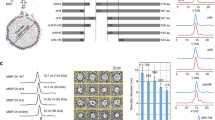
Structural studies of membrane proteins (MP) in a native or native-like environment remain a challenge. X-ray crystallography of three-dimensional crystals of MP in lipids and cryo-electron microscopy of two-dimensional crystals also in lipids have given atomic structures of several MP. Recent developments of solid-state NMR (ssNMR) provided structural data of MP in lipids and should give access to the dynamic behavior of MP’s in a native-like environment. Preparation of samples for ssNMR is not trivial with overexpressed proteins since purified recombinant MP have to be reincorporated in proteoliposomes and concentrated in the small volume of the rotor used for ssNMR studies. We present here the protocol that we have used to study the recombinant mouse TSPO1, an integral membrane protein of 20 kDa mostly found in the outer membrane of mitochondria and overexpressed in E. coli bacteria.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lacapere J-J et al (2007) Determining membrane protein structures: still a challenge! Trends Biochem Sci 32(6):259–270
Ubarretxena-Belandia I, Stokes DL (2010) Present and future of membrane protein structure determination by electron crystallography. Adv Protein Chem Struct Biol 81:33–60
Moraes I et al (2014) Membrane protein structure determination – the next generation. Biochim Biophys Acta 1838:78–87
Barends TRM et al (2014) De novo protein crystal structure determination from X-ray free-electron laser data. Nature 505(7482):244–247
Binshteim E, Ohi MD (2015) Cryo-electron microscopy and the amazing race to atomic resolution. Biochemistry 54(20):3133–3141
De Zorzi R et al (2016) Single-particle electron microscopy in the study of membrane protein structure. Microscopy (Oxf) 65(1):81–96
McDermott A (2009) Structure and dynamics of membrane proteins by magic angle spinning solid-state NMR. Annu Rev Biophys 38:385–403
Kunert B et al (2014) Efficient and stable reconstitution of the ABC transporterBmrA for solid-state NMR studies. Front Mol Biosci 1(5):1–11
Rigaud J-L, Levy D (2003) Reconstitution of membrane proteins into liposomes. Methods Enzymol 372:65–86
Lacapere J-J et al (2001) Structural and functional study of reconstituted peripheral benzodiazepine receptor. Biochem Biophys Res Commun 284:536–541
Ostuni MA et al (2010) Characterization of membrane protein preparations: measurement of detergent content and ligand binding after proteoliposomes reconstitution. Methods Mol Biol 654:3–18
Teboul D et al (2012) Mouse TSPO in a lipid environment interacting with a functionalized monolayer. BBA-Biomembranes 1818(11):2791–2800
Lacapere J-J et al (2014) Structural studies of TSPO, a mitochondrial protein. In: Mus-Veteau I (ed) Membrane proteins production for structural studies. Springer, New York, NY, pp 393–422
Böckmann A et al (2009) Characterization of different water pools in solid-state NMR protein samples. J Biomol NMR 45:319–327
Abdine A, Park KH, Warschawski DE (2012) Cell-free membrane protein expression for solid-state NMR. Methods Mol Biol 831:85–109
Das BB et al (2013) Lipid bilayer preparations of membrane proteins for oriented and magic-angle spinning solid-state NMR samples. Nat Protoc 8(11):2256–2270
Dvinskikh SV et al (2005) Efficient solid-state NMR methods for measuring heteronuclear dipolar couplings in unoriented lipid membrane systems. Phys Chem Chem Phys 7:607–613
Bennett AE et al (1995) Heteronuclear decoupling in rotating solids. J Chem Phys 103:6951–6958
Morcombe CR, Zilm KW (2003) Chemical shift referencing in MAS solid state NMR. J Magn Reson 162:479–486
Purusottam R et al (2015) Probing the gel to liquid-crystalline phase transition and relevant conformation changes in liposomes by (13)C magic-angle spinning NMR spectroscopy. BBA-Biomembranes 1848(12):3134–3139
Das BB et al (2014) Dipolar assisted assignment protocol (DAAP) for MAS solid-state NMR of rotationally aligned membrane proteins in phospholipid bilayers. J Magn Reson 242:224–232
Das BB et al (2015) Membrane protein structure from rotational diffusion. Biochim Biophys Acta 1848:229–245
Eddy MT et al (2015) Lipid bilayer-bound conformation of an integral membrane beta barrel protein by multidimensional MAS NMR. J Biomol NMR 61:299–310
Bertani P et al (2014) 15N chemical shift referencing in solid state NMR. Solid State Nucl Magn Reson 61-62:15–18
Detken A et al (2002) Simple and efficient decoupling in magic-angle spinning solid-state NMR: the XiX scheme. Chem Phys Lett 356:298–304
Loening et al. (2012) A comparison of NCO and NCA transfer methods for biological solid-state NMR spectroscopy. J Magn Reson 214:81–90
Fritzsching KJ et al (2013) Practical use of chemical shift databases for protein solid-state NMR: 2D chemical shift maps and amino-acid assignment with secondary-structure information. J Biomol NMR 56:155–167
Wiegand T et al (2015) Solid-state NMR sequential assignments the N-terminal domain of HpDnaB heilcase. Biomol NMR Assign 9:1–11
Ong et al. (2013) Detecting substrates bound to the secondary multidrug efflux pump EmrE by DNP-enhanced solid-state NMR. J Am Chem Soc 135:15754–15762
Hellstrand E et al (2013) Membrane lipid co-aggregation with α-synuclein fibrils. PLoS ONE 8(10):e77235
Acknowledgments
We would like to thank Stefanie Finet and Jean-Michel Guignier (UPMC, Paris) for their help and advice with DLS and cryo-EM experiments, Dominique Cailleu (Plateforme Analytique, Amiens) for help with NMR experiments on the 11.7 T magnet and Olivier Lequin (UPMC, Paris) for fruitful discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Senicourt, L., Duma, L., Papadopoulos, V., Lacapere, JJ. (2017). Solid-State NMR of Membrane Protein Reconstituted in Proteoliposomes, the Case of TSPO. In: Lacapere, JJ. (eds) Membrane Protein Structure and Function Characterization. Methods in Molecular Biology, vol 1635. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7151-0_18
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7151-0_18
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7149-7
Online ISBN: 978-1-4939-7151-0
eBook Packages: Springer Protocols