Abstract
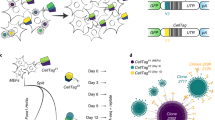
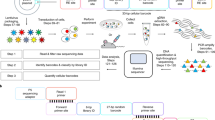
We describe here a method to identify the position of retroviral insertion sites and simultaneously to quantify the absolute abundance of each clone, i.e., the number of cells having the provirus inserted at a given place in the host genome. The method is based on random shearing of the host cell DNA, followed by a linker-mediated PCR to amplify the genomic regions flanking the proviruses, and high-throughput sequencing of the amplicons. The quantification of the abundance of each infected clone allowed us to develop two new metrics: i. the oligoclonality index, which quantifies the nonuniformity of the distribution of clone abundance, and ii. an estimator of the total number of clones in the body of the host. These new tools are valuable for the study of retroviral infections and can also be adapted for the tracking of gene-edited cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lewinski MK, Yamashita M, Emerman M, Ciuffi A, Marshall H et al (2006) Retroviral DNA integration: viral and cellular determinants of target-site selection. PLoS Pathog 2:e60
Craigie R, Bushman FD (2012) HIV DNA integration. Cold Spring Harb Perspect Med 2:a006890
Gillet NA, Malani N, Melamed A, Gormley N, Carter R et al (2011) The host genomic environment of the provirus determines the abundance of HTLV-1-infected T-cell clones. Blood 117:3113–3122
Cook LB, Rowan AG, Melamed A, Taylor GP, Bangham CR (2012) HTLV-1-infected T cells contain a single integrated provirus in natural infection. Blood 120:3488–3490
Melamed A, Laydon D, Gillet N, Tanaka Y, Taylor G et al (2013) Genome-wide determinants of proviral targeting, clonal abundance and expression in natural HTLV-1 Infection. PLoS Pathog 9
Gillet NA, Gutierrez G, Rodriguez SM, de Brogniez A, Renotte N et al (2013) Massive depletion of bovine leukemia virus proviral clones located in genomic transcriptionally active sites during primary infection. PLoS Pathog 9:e1003687
Melamed A, Witkover AD, Laydon DJ, Brown R, Ladell K et al (2014) Clonality of HTLV-2 in natural infection. PLoS Pathog 10:e1004006
Cook LB, Melamed A, Niederer H, Valganon M, Laydon D et al (2014) The role of HTLV-1 clonality, proviral structure, and genomic integration site in adult T-cell leukemia/lymphoma. Blood 123:3925–3931
Bangham CR, Cook LB, Melamed A (2014) HTLV-1 clonality in adult T-cell leukaemia and non-malignant HTLV-1 infection. Semin Cancer Biol 26C:89–98
Berry CC, Gillet NA, Melamed A, Gormley N, Bangham CR et al (2012) Estimating abundances of retroviral insertion sites from DNA fragment length data. Bioinformatics 28:755–762
Laydon DJ, Melamed A, Sim A, Gillet NA, Sim K et al (2014) Quantification of HTLV-1 clonality and TCR diversity. PLoS Comput Biol 10:e1003646
Maldarelli F, Wu X, Su L, Simonetti FR, Shao W et al (2014) Specific HIV integration sites are linked to clonal expansion and persistence of infected cells. Science 345(6193):179–183
Trobridge GD (2011) Genotoxicity of retroviral hematopoietic stem cell gene therapy. Expert Opin Biol Ther 11:581–593
Gaj T, Gersbach CA, Barbas CF 3rd (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405
Bahassi EM, Stambrook PJ (2014) Next-generation sequencing technologies: breaking the sound barrier of human genetics. Mutagenesis 29:303–310
Krueger F, Andrews SR, Osborne CS (2011) Large scale loss of data in low-diversity illumina sequencing libraries can be recovered by deferred cluster calling. PLoS One 6:e16607
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Firouzi S, Lopez Y, Suzuki Y, Nakai K, Sugano S et al (2014) Development and validation of a new high-throughput method to investigate the clonality of HTLV-1-infected cells based on provirus integration sites. Genome Med 6:46
Gini (1914) Sulla misura della concentrazione e della variabilita dei caratteri: transactions of the real istituto veneto di scienze
Handcock MS, Morris M (1999) Relative distribution methods in the social sciences. Springer, New York, NY
Acknowledgments
Nicolas A. Gillet and Anat Melamed contributed equally to this paper.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Gillet, N.A., Melamed, A., Bangham, C.R.M. (2017). High-Throughput Mapping and Clonal Quantification of Retroviral Integration Sites. In: Casoli, C. (eds) Human T-Lymphotropic Viruses. Methods in Molecular Biology, vol 1582. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6872-5_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6872-5_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6870-1
Online ISBN: 978-1-4939-6872-5
eBook Packages: Springer Protocols