Abstract
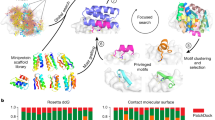
The ability to design novel small-molecule binding sites in proteins is a stringent test of our understanding of the principles of molecular recognition, and would have many practical applications, in synthetic biology and medicine. Here, we describe a computational method in the context of the macromolecular modeling suite Rosetta to designing proteins with sites featuring predetermined interactions to ligands of choice. The required inputs for the method are a model of the small molecule and the desired interactions (e.g., hydrogen bonding, electrostatics, steric packing), and a set of crystallographic structures of proteins containing existing or predicted binding pockets. Constellations of backbones surrounding the putative pocket are searched for compatibility with the desired binding site conception using RosettaMatch and surrounding amino acid side chain identities are optimized using RosettaDesign. Validation of the design is performed using metrics that evaluate the interface energy of the predicted binding pose, the preformation of key binding site features in the apo-state, and the local compatibility of the designed sequence changes with the wild type backbone structure, and top-ranking candidate designs are generated for experimental validation. This approach can allow for the creation of novel binding sites and for the rational tuning of specificity for congeneric ligands by altering the programmed interactions by design, thus offering a general computational protocol for construction and modulation of protein–small molecule interfaces.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
de Wolf FA, Brett GM (2000) Ligand-binding proteins: their potential for application in systems for controlled delivery and uptake of ligands. Pharmacol Rev 52(2):207–236
Hunter MM, Margolies MN, Ju A, Haber E (1982) High-affinity monoclonal antibodies to the cardiac glycoside, digoxin. J Immunol 129(3):1165–1172
Shen XY, Orson FM, Kosten TR (2012) Vaccines against drug abuse. Clin Pharmacol Ther 91:60–70
Bradbury ARM, Sidhu S, Dübel S, McCafferty J (2011) Beyond natural antibodies: the power of in vitro display technologies. Nat Biotechnol 29(3):245–254
Brustad EM, Arnold FH (2011) Optimizing non-natural protein function with directed evolution. Curr Opin Chem Biol 15(2):201–210. doi:10.1016/j.cbpa.2010.11.020
Chen G, Hayhurst A, Thomas JG, Harvey BR, Iverson BL, Georgiou G (2001) Isolation of high-affinity ligand-binding proteins by periplasmic expression with cytometric screening (PECS). Nat Biotechnol 19(6):537–542
Rothlisberger D, Khersonsky O, Wollacott AM, Jiang L, DeChancie J, Betker J, Gallaher JL, Althoff EA, Zanghellini A, Dym O, Albeck S, Houk KN, Tawfik DS, Baker D (2008) Kemp elimination catalysts by computational enzyme design. Nature 453(7192):190–195. doi:10.1038/nature06879
Jiang L, Althoff EA, Clemente FR, Doyle L, Rothlisberger D, Zanghellini A, Gallaher JL, Betker JL, Tanaka F, Barbas CF 3rd, Hilvert D, Houk KN, Stoddard BL, Baker D (2008) De novo computational design of retro-aldol enzymes. Science 319(5868):1387–1391. doi:10.1126/science.1152692
Siegel JB, Zanghellini A, Lovick HM, Kiss G, Lambert AR, St Clair JL, Gallaher JL, Hilvert D, Gelb MH, Stoddard BL, Houk KN, Michael FE, Baker D (2010) Computational design of an enzyme catalyst for a stereoselective bimolecular Diels-Alder reaction. Science 329(5989):309–313. doi:10.1126/science.1190239
Khersonsky O, Rothlisberger D, Dym O, Albeck S, Jackson CJ, Baker D, Tawfik DS (2010) Evolutionary optimization of computationally designed enzymes: Kemp eliminases of the KE07 series. J Mol Biol 396(4):1025–1042. doi:10.1016/j.jmb.2009.12.031
Khare SD, Fleishman SJ (2013) Emerging themes in the computational design of novel enzymes and protein-protein interfaces. FEBS Lett 587(8):1147–1154. doi:10.1016/j.febslet.2012.12.009
Fleishman SJ, Khare SD, Koga N, Baker D (2011) Restricted sidechain plasticity in the structures of native proteins and complexes. Protein Sci 20(4):753–757. doi:10.1002/pro.604
Korndörfer IP, Schlehuber S, Skerra A (2003) Structural mechanism of specific ligand recognition by a lipocalin tailored for the complexation of digoxigenin. J Mol Biol 330(2):385–396
Röthlisberger D, Khersonsky O, Wollacott AM, Jiang L, DeChancie J, Betker J, Gallaher JL, Althoff EA, Zanghellini A, Dym O, Albeck S, Houk KN, Tawfik DS, Baker D (2008) Kemp elimination catalysts by computational enzyme design. Nature 453(7192):190–195
Holm L, Rosenström P (2010) Dali server: conservation mapping in 3D. Nucleic Acids Res 38(suppl 2):W545–W549
Bloom JD, Labthavikul ST, Otey CR, Arnold FH (2006) Protein stability promotes evolvability. Proc Natl Acad Sci U S A 103(15):5869–5874
Tokuriki N, Tawfik DS (2009) Chaperonin overexpression promotes genetic variation and enzyme evolution. Nature 459(7247):668–673
Jeffrey PD, Strong RK, Sieker LC, Chang CY, Campbell RL, Petsko GA, Haber E, Margolies MN, Sheriff S (1993) 26-10 Fab–digoxin complex: affinity and specificity due to surface complementarity. Proc Natl Acad Sci U S A 90(21):10310–10314
Richter F, Leaver-Fay A, Khare SD, Bjelic S, Baker D (2011) De novo enzyme design using Rosetta3. PLoS One 6(5), e19230
Fleishman SJ, Leaver-Fay A, Corn JE, Strauch E-M, Khare SD, Koga N, Ashworth J, Murphy P, Richter F, Lemmon G, Meiler J, Baker D (2011) RosettaScripts: a scripting language interface to the Rosetta macromolecular modeling suite. PLoS One 6(6), e20161
Davis IW, Baker D (2009) RosettaLigand docking with full ligand and receptor flexibility. J Mol Biol 385(2):381–392
Kellogg EH, Leaver-Fay A, Baker D (2010) Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins 79(3):830–838. doi:10.1002/prot.22921
Collaborative Computational Project N (1994) The CCP4 suite: programs for protein crystallography. Acta Crystallogr Sect D 50(5), 760–763. doi: 10.1107/S0907444994003112
Lawrence MC, Colman PM (1993) Shape complementarity at protein/protein interfaces. J Mol Biol 234(4):946–950
Cooper S, Khatib F, Treuille A, Barbero J, Lee J, Beenen M, Leaver-Fay A, Baker D, Popovic Z, Players F (2010) Predicting protein structures with a multiplayer online game. Nature 466(7307):756–760
Fleishman SJ, Khare SD, Koga N, Baker D (2011) Restricted sidechain plasticity in the structures of native proteins and complexes. Protein Sci 20(4):753–757
Raman S, Vernon R, Thompson J, Tyka M, Sadreyev R, Pei J, Kim D, Kellogg E, DiMaio F, Lange O, Kinch L, Sheffler W, Kim B-H, Das R, Grishin NV, Baker D (2009) Structure prediction for CASP8 with all-atom refinement using Rosetta. Proteins 77(S9):89–99. doi:10.1002/prot.22540
Tinberg CE, Khare SD, Dou J, Doyle L, Nelson JW, Schena A, Jankowski W, Kalodimos CG, Johnsson K, Stoddard BL, Baker D (2013) Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501(7466):212–216. doi:10.1038/nature12443
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Tinberg, C.E., Khare, S.D. (2017). Computational Design of Ligand Binding Proteins. In: Samish, I. (eds) Computational Protein Design. Methods in Molecular Biology, vol 1529. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6637-0_19
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6637-0_19
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6635-6
Online ISBN: 978-1-4939-6637-0
eBook Packages: Springer Protocols