Abstract
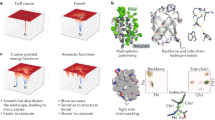
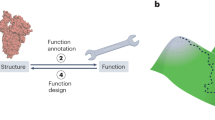
Computational protein design (CPD) has established itself as a leading field in basic and applied science with a strong coupling between the two. Proteins are computationally designed from the level of amino acids to the level of a functional protein complex. Design targets range from increased thermo- (or other) stability to specific requested reactions such as protein–protein binding, enzymatic reactions, or nanotechnology applications. The design scheme may encompass small regions of the proteins or the entire protein. In either case, the design may aim at the side-chains or at the full backbone conformation. Herein, the main framework for the process is outlined highlighting key elements in the CPD iterative cycle. These include the very definition of CPD, the diverse goals of CPD, components of the CPD protocol, methods for searching sequence and structure space, scoring functions, and augmenting the CPD with other optimization tools. Taken together, this chapter aims to introduce the framework of CPD.
“Most people make the mistake of thinking design is what it looks like. People think it’s this veneer—that the designers are handed this box and told, ‘Make it look good!’ That’s not what we think design is. It’s not just what it looks like and feels like. Design is how it works.” Steve Jobs, Apple’s C.E.O in an interview to the New-York Times. Nov. 30th 2003, The Guts of a New Machine
http://www.nytimes.com/2003/11/30/magazine/the-guts-of-a-new-machine.html
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Samish I, MacDermaid CM, Perez-Aguilar JM, Saven JG (2011) Theoretical and computational protein design. Annu Rev Phys Chem 62:129–149. doi:10.1146/annurev-physchem-032210-103509
Pantazes RJ, Grisewood MJ, Maranas CD (2011) Recent advances in computational protein design. Curr Opin Struct Biol 21(4):467–472. doi:10.1016/j.sbi.2011.04.005
Saven JG (2011) Computational protein design: engineering molecular diversity, nonnatural enzymes, nonbiological cofactor complexes, and membrane proteins. Curr Opin Chem Biol 15(3):452–457. doi:10.1016/j.cbpa.2011.03.014
Tian P (2010) Computational protein design, from single domain soluble proteins to membrane proteins. Chem Soc Rev 39(6):2071–2082. doi:10.1039/b810924a
Suarez M, Jaramillo A (2009) Challenges in the computational design of proteins. J R Soc Interface 6 Suppl 4:S477–S491. doi:10.1098/rsif.2008.0508.focus
Lippow SM, Tidor B (2007) Progress in computational protein design. Curr Opin Biotechnol 18(4):305–311. doi:10.1016/j.copbio.2007.04.009
Rosenberg M, Goldblum A (2006) Computational protein design: a novel path to future protein drugs. Curr Pharm Des 12(31):3973–3997
Butterfoss GL, Kuhlman B (2006) Computer-based design of novel protein structures. Annu Rev Biophys Biomol Struct 35:49–65. doi:10.1146/annurev.biophys.35.040405.102046
Johnson LB, Huber TR, Snow CD (2014) Methods for library-scale computational protein design. Methods Mol Biol 1216:129–159. doi:10.1007/978-1-4939-1486-9_7
Davey JA, Chica RA (2012) Multistate approaches in computational protein design. Protein Sci 21(9):1241–1252. doi:10.1002/pro.2128
Lassila JK (2010) Conformational diversity and computational enzyme design. Curr Opin Chem Biol 14(5):676–682. doi:10.1016/j.cbpa.2010.08.010
Mandell DJ, Kortemme T (2009) Backbone flexibility in computational protein design. Curr Opin Biotechnol 20(4):420–428. doi:10.1016/j.copbio.2009.07.006
Ollikainen N, Smith CA, Fraser JS, Kortemme T (2013) Flexible backbone sampling methods to model and design protein alternative conformations. Methods Enzymol 523:61–85. doi:10.1016/B978-0-12-394292-0.00004-7
Vizcarra CL, Mayo SL (2005) Electrostatics in computational protein design. Curr Opin Chem Biol 9(6):622–626. doi:10.1016/j.cbpa.2005.10.014
Verschueren E, Vanhee P, van der Sloot AM, Serrano L, Rousseau F, Schymkowitz J (2011) Protein design with fragment databases. Curr Opin Struct Biol 21(4):452–459. doi:10.1016/j.sbi.2011.05.002
Saven JG (2001) Designing protein energy landscapes. Chem Rev 101(10):3113–3130
Kuhlman B, Baker D (2004) Exploring folding free energy landscapes using computational protein design. Curr Opin Struct Biol 14(1):89–95. doi:10.1016/j.sbi.2004.01.002
Hwang I, Park S (2008) Computational design of protein therapeutics. Drug Discov Today Technol 5(2-3):e43–e48. doi:10.1016/j.ddtec.2008.11.004
Feldmeier K, Hocker B (2013) Computational protein design of ligand binding and catalysis. Curr Opin Chem Biol 17(6):929–933
Wijma HJ, Janssen DB (2013) Computational design gains momentum in enzyme catalysis engineering. FEBS J 280(13):2948–2960. doi:10.1111/febs.12324
Khare SD, Fleishman SJ (2013) Emerging themes in the computational design of novel enzymes and protein-protein interfaces. FEBS Lett 587(8):1147–1154. doi:10.1016/j.febslet.2012.12.009
Nanda V, Koder RL (2010) Designing artificial enzymes by intuition and computation. Nat Chem 2(1):15–24. doi:10.1038/nchem.473
Havranek JJ (2010) Specificity in computational protein design. J Biol Chem 285(41):31095–31099. doi:10.1074/jbc.R110.157685
Sharabi O, Erijman A, Shifman JM (2013) Computational methods for controlling binding specificity. Methods Enzymol 523:41–59. doi:10.1016/B978-0-12-394292-0.00003-5
Senes A (2011) Computational design of membrane proteins. Curr Opin Struct Biol 21(4):460–466. doi:10.1016/j.sbi.2011.06.004
Perez-Aguilar JM, Saven JG (2012) Computational design of membrane proteins. Structure 20(1):5–14. doi:10.1016/j.str.2011.12.003
Parmar AS, Pike D, Nanda V (2014) Computational design of metalloproteins. Methods Mol Biol 1216:233–249. doi:10.1007/978-1-4939-1486-9_12
Nanda V, Zahid S, Xu F, Levine D (2011) Computational design of intermolecular stability and specificity in protein self-assembly. Methods Enzymol 487:575–593. doi:10.1016/B978-0-12-381270-4.00020-2
Ambroggio XI, Kuhlman B (2006) Design of protein conformational switches. Curr Opin Struct Biol 16(4):525–530. doi:10.1016/j.sbi.2006.05.014
Kortemme T, Baker D (2004) Computational design of protein-protein interactions. Curr Opin Chem Biol 8(1):91–97. doi:10.1016/j.cbpa.2003.12.008
Joyce GF (2007) Forty years of in vitro evolution. Angew Chem Int Ed Engl 46(34):6420–6436. doi:10.1002/anie.200701369
Fleishman SJ, Whitehead TA, Ekiert DC, Dreyfus C, Corn JE, Strauch EM, Wilson IA, Baker D (2011) Computational design of proteins targeting the conserved stem region of influenza hemagglutinin. Science 332(6031):816–821. doi:10.1126/science.1202617
Shlyk-Kerner O, Samish I, Kaftan D, Holland N, Sai PS, Kless H, Scherz A (2006) Protein flexibility acclimatizes photosynthetic energy conversion to the ambient temperature. Nature 442(7104):827–830. doi:10.1038/nature04947
Lane MD, Seelig B (2014) Advances in the directed evolution of proteins. Curr Opin Chem Biol 22:129–136. doi:10.1016/j.cbpa.2014.09.013
Packer MS, Liu DR (2015) Methods for the directed evolution of proteins. Nat Rev Genet 16(7):379–394. doi:10.1038/nrg3927
Moult J, Fidelis K, Kryshtafovych A, Schwede T, Tramontano A (2014) Critical assessment of methods of protein structure prediction (CASP)—round x. Proteins 82(Suppl 2):1–6. doi:10.1002/prot.24452
Yue K, Dill KA (1992) Inverse protein folding problem: designing polymer sequences. Proc Natl Acad Sci U S A 89(9):4163–4167
Whitehead TA, Baker D, Fleishman SJ (2013) Computational design of novel protein binders and experimental affinity maturation. Methods Enzymol 523:1–19. doi:10.1016/B978-0-12-394292-0.00001-1
Rothlisberger D, Khersonsky O, Wollacott AM, Jiang L, DeChancie J, Betker J, Gallaher JL, Althoff EA, Zanghellini A, Dym O, Albeck S, Houk KN, Tawfik DS, Baker D (2008) Kemp elimination catalysts by computational enzyme design. Nature 453(7192):190–195. doi:10.1038/nature06879
Tanford C (1978) The hydrophobic effect and the organization of living matter. Science 200(4345):1012–1018
Bowie JU, Luthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253(5016):164–170
Godzik A, Kolinski A, Skolnick J (1992) Topology fingerprint approach to the inverse protein folding problem. J Mol Biol 227(1):227–238
Carbonell P, Trosset JY (2015) Computational protein design methods for synthetic biology. Methods Mol Biol 1244:3–21. doi:10.1007/978-1-4939-1878-2_1
Richter F, Baker D (2013) Computational protein design for synthetic biology. In: Zhao H (ed) Synthetic biology tools and applications. Elsevier Inc., San Diego, CA
Quinn TP, Tweedy NB, Williams RW, Richardson JS, Richardson DC (1994) Betadoublet: de novo design, synthesis, and characterization of a beta-sandwich protein. Proc Natl Acad Sci U S A 91(19):8747–8751
Kuhlman B, Dantas G, Ireton GC, Varani G, Stoddard BL, Baker D (2003) Design of a novel globular protein fold with atomic-level accuracy. Science 302(5649):1364–1368. doi:10.1126/science.1089427
Kaplan J, DeGrado WF (2004) De novo design of catalytic proteins. Proc Natl Acad Sci U S A 101(32):11566–11570. doi:10.1073/pnas.0404387101
Joh NH, Wang T, Bhate MP, Acharya R, Wu Y, Grabe M, Hong M, Grigoryan G, DeGrado WF (2014) De novo design of a transmembrane Zn(2)(+)-transporting four-helix bundle. Science 346(6216):1520–1524. doi:10.1126/science.1261172
Huang PS, Love JJ, Mayo SL (2007) A de novo designed protein protein interface. Protein Sci 16(12):2770–2774. doi:10.1110/ps.073125207
Desjarlais JR, Handel TM (1995) De novo design of the hydrophobic cores of proteins. Protein Sci 4(10):2006–2018. doi:10.1002/pro.5560041006
Ventura S, Vega MC, Lacroix E, Angrand I, Spagnolo L, Serrano L (2002) Conformational strain in the hydrophobic core and its implications for protein folding and design. Nat Struct Biol 9(6):485–493. doi:10.1038/nsb799
Keating AE, Malashkevich VN, Tidor B, Kim PS (2001) Side-chain repacking calculations for predicting structures and stabilities of heterodimeric coiled coils. Proc Natl Acad Sci U S A 98(26):14825–14830. doi:10.1073/pnas.261563398
Slovic AM, Kono H, Lear JD, Saven JG, DeGrado WF (2004) Computational design of water-soluble analogues of the potassium channel KcsA. Proc Natl Acad Sci U S A 101(7):1828–1833. doi:10.1073/pnas.0306417101
Slovic AM, Summa CM, Lear JD, DeGrado WF (2003) Computational design of a water-soluble analog of phospholamban. Protein Sci 12(2):337–348. doi:10.1110/ps.0226603
Voet AR, Noguchi H, Addy C, Simoncini D, Terada D, Unzai S, Park SY, Zhang KY, Tame JR (2014) Computational design of a self-assembling symmetrical beta-propeller protein. Proc Natl Acad Sci U S A 111(42):15102–15107. doi:10.1073/pnas.1412768111
Woolfson DN, Bartlett GJ, Bruning M, Thomson AR (2012) New currency for old rope: from coiled-coil assemblies to alpha-helical barrels. Curr Opin Struct Biol 22(4):432–441. doi:10.1016/j.sbi.2012.03.002
Lanci CJ, MacDermaid CM, Kang SG, Acharya R, North B, Yang X, Qiu XJ, DeGrado WF, Saven JG (2012) Computational design of a protein crystal. Proc Natl Acad Sci U S A 109(19):7304–7309. doi:10.1073/pnas.1112595109
Swift J, Wehbi WA, Kelly BD, Stowell XF, Saven JG, Dmochowski IJ (2006) Design of functional ferritin-like proteins with hydrophobic cavities. J Am Chem Soc 128(20):6611–6619. doi:10.1021/ja057069x
Summa CM, Rosenblatt MM, Hong JK, Lear JD, DeGrado WF (2002) Computational de novo design, and characterization of an A(2)B(2) diiron protein. J Mol Biol 321(5):923–938
Cochran FV, Wu SP, Wang W, Nanda V, Saven JG, Therien MJ, DeGrado WF (2005) Computational de novo design and characterization of a four-helix bundle protein that selectively binds a nonbiological cofactor. J Am Chem Soc 127(5):1346–1347. doi:10.1021/ja044129a
Shifman JM, Mayo SL (2003) Exploring the origins of binding specificity through the computational redesign of calmodulin. Proc Natl Acad Sci U S A 100(23):13274–13279. doi:10.1073/pnas.2234277100
Kortemme T, Joachimiak LA, Bullock AN, Schuler AD, Stoddard BL, Baker D (2004) Computational redesign of protein-protein interaction specificity. Nat Struct Mol Biol 11(4):371–379. doi:10.1038/nsmb749
Potapov V, Reichmann D, Abramovich R, Filchtinski D, Zohar N, Ben Halevy D, Edelman M, Sobolev V, Schreiber G (2008) Computational redesign of a protein-protein interface for high affinity and binding specificity using modular architecture and naturally occurring template fragments. J Mol Biol 384(1):109–119. doi:10.1016/j.jmb.2008.08.078
Lippow SM, Wittrup KD, Tidor B (2007) Computational design of antibody-affinity improvement beyond in vivo maturation. Nat Biotechnol 25(10):1171–1176. doi:10.1038/nbt1336
Dagliyan O, Shirvanyants D, Karginov AV, Ding F, Fee L, Chandrasekaran SN, Freisinger CM, Smolen GA, Huttenlocher A, Hahn KM, Dokholyan NV (2013) Rational design of a ligand-controlled protein conformational switch. Proc Natl Acad Sci U S A 110(17):6800–6804. doi:10.1073/pnas.1218319110
Korendovych IV, Kulp DW, Wu Y, Cheng H, Roder H, DeGrado WF (2011) Design of a switchable eliminase. Proc Natl Acad Sci U S A 108(17):6823–6827. doi:10.1073/pnas.1018191108
Yin H, Slusky JS, Berger BW, Walters RS, Vilaire G, Litvinov RI, Lear JD, Caputo GA, Bennett JS, DeGrado WF (2007) Computational design of peptides that target transmembrane helices. Science 315(5820):1817–1822. doi:10.1126/science.1136782
Samish I (2009) Search and sampling in structural bioinformatics. In: Bourne P, Gu J (eds) Structural bioinformatics. Wiley, New York, pp 207–236
Dunbrack RL Jr, Karplus M (1993) Backbone-dependent rotamer library for proteins. Application to side-chain prediction. J Mol Biol 230(2):543–574. doi:10.1006/jmbi.1993.1170
Dunbrack RL Jr (2002) Rotamer libraries in the 21st century. Curr Opin Struct Biol 12(4):431–440
Shapovalov MV, Dunbrack RL Jr (2011) A smoothed backbone-dependent rotamer library for proteins derived from adaptive kernel density estimates and regressions. Structure 19(6):844–858. doi:10.1016/j.str.2011.03.019
Subramaniam S, Senes A (2014) Backbone dependency further improves side chain prediction efficiency in the Energy-based Conformer Library (bEBL). Proteins 82(11):3177–3187. doi:10.1002/prot.24685
Kuhlman B, Baker D (2000) Native protein sequences are close to optimal for their structures. Proc Natl Acad Sci U S A 97(19):10383–10388
Grigoryan G, Degrado WF (2011) Probing designability via a generalized model of helical bundle geometry. J Mol Biol 405(4):1079–1100. doi:10.1016/j.jmb.2010.08.058
Schramm CA, Hannigan BT, Donald JE, Keasar C, Saven JG, Degrado WF, Samish I (2012) Knowledge-based potential for positioning membrane-associated structures and assessing residue-specific energetic contributions. Structure 20(5):924–935. doi:10.1016/j.str.2012.03.016
Xu F, Zahid S, Silva T, Nanda V (2011) Computational design of a collagen A:B:C-type heterotrimer. J Am Chem Soc 133(39):15260–15263. doi:10.1021/ja205597g
Shifman JM, Mayo SL (2002) Modulating calmodulin binding specificity through computational protein design. J Mol Biol 323(3):417–423
Havranek JJ, Harbury PB (2003) Automated design of specificity in molecular recognition. Nat Struct Biol 10(1):45–52. doi:10.1038/nsb877
Bolon DN, Grant RA, Baker TA, Sauer RT (2005) Specificity versus stability in computational protein design. Proc Natl Acad Sci U S A 102(36):12724–12729. doi:10.1073/pnas.0506124102
Grigoryan G, Reinke AW, Keating AE (2009) Design of protein-interaction specificity gives selective bZIP-binding peptides. Nature 458(7240):859–864. doi:10.1038/nature07885
Fry HC, Lehmann A, Saven JG, DeGrado WF, Therien MJ (2010) Computational design and elaboration of a de novo heterotetrameric alpha-helical protein that selectively binds an emissive abiological (porphinato)zinc chromophore. J Am Chem Soc 132(11):3997–4005. doi:10.1021/ja907407m
Koga N, Tatsumi-Koga R, Liu G, Xiao R, Acton TB, Montelione GT, Baker D (2012) Principles for designing ideal protein structures. Nature 491(7423):222–227. doi:10.1038/nature11600
Fry HC, Lehmann A, Sinks LE, Asselberghs I, Tronin A, Krishnan V, Blasie JK, Clays K, DeGrado WF, Saven JG, Therien MJ (2013) Computational de novo design and characterization of a protein that selectively binds a highly hyperpolarizable abiological chromophore. J Am Chem Soc 135(37):13914–13926. doi:10.1021/ja4067404
Jamroz M, Kolinski A (2010) Modeling of loops in proteins: a multi-method approach. BMC Struct Biol 10:5. doi:10.1186/1472-6807-10-5
Hildebrand PW, Goede A, Bauer RA, Gruening B, Ismer J, Michalsky E, Preissner R (2009) SuperLooper—a prediction server for the modeling of loops in globular and membrane proteins. Nucleic Acids Res 37:W571–W574. doi:10.1093/nar/gkp338
Soto CS, Fasnacht M, Zhu J, Forrest L, Honig B (2008) Loop modeling: sampling, filtering, and scoring. Proteins 70(3):834–843. doi:10.1002/prot.21612
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Samish, I. (2017). The Framework of Computational Protein Design. In: Samish, I. (eds) Computational Protein Design. Methods in Molecular Biology, vol 1529. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6637-0_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6637-0_1
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6635-6
Online ISBN: 978-1-4939-6637-0
eBook Packages: Springer Protocols