Abstract
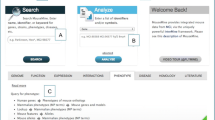
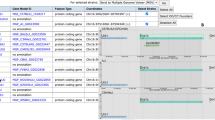
The Mouse Genome Informatics (MGI), resource (www.informatics.jax.org) has existed for over 25 years, and over this time its data content, informatics infrastructure, and user interfaces and tools have undergone dramatic changes (Eppig et al., Mamm Genome 26:272–284, 2015). Change has been driven by scientific methodological advances, rapid improvements in computational software, growth in computer hardware capacity, and the ongoing collaborative nature of the mouse genomics community in building resources and sharing data. Here we present an overview of the current data content of MGI, describe its general organization, and provide examples using simple and complex searches, and tools for mining and retrieving sets of data.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Eppig JT, Richardson JE, Kadin JA, Ringwald M, Blake JA, Bult CJ (2015) Mouse Genome Informatics (MGI): reflecting on 25 years. Mamm Genome 26:272–284
Eppig JT, Richardson JE, Kadin JA, Smith CL, Blake JA, Bult CJ, MGD Team (2015) Mouse Genome Database: from sequence to phenotypes and disease models. Genesis 53:458–473
Finger JH, Smith CM, Hayamizu TF, McCright IJ, Xu J, Eppig JT, Kadin JA, Richardson JE, Ringwald M (2015) The mouse Gene Expression Database (GXD): new features and how to get the most out of them. Genesis 53:510–522
Murray SA, Eppig JT, Smedley D, Simpson EM, Rosenthal N (2012) Beyond knockouts: cre resources for conditional mutagenesis. Mamm Genome 23:587–599
Bult CJ, Krupke DM, Begley DA, Richardson JE, Neuhauser SB, Sundberg JP, Eppig JT (2015) Mouse Tumor Biology (MTB): a database of mouse models for human cancer. Nucleic Acids Res 43:D818–D824
Shultz LD, Lyons BL, Burzenski LM, Gott B, Chen X, Chaleff S, Kotb M, Gillies SD, King M, Mangada J, Greiner DL, Handgretinger R (2005) Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2R gamma null mice engrafted with mobilized human hemopoietic stem cells. J Immunol 174:6477–6489
Eppig JT, Motenko H, Richardson JE, Richards-Smith B, Smith CL (2015) The International Mouse Strain Resource (IMSR): cataloging worldwide mouse and ES cell line resources. Mamm Genome 26:448–455
Eppig JT, Blake JA, Bult CJ, Kadin JA, Richardson JE, Mouse Genome Database Group (2015) The Mouse Genome Database (MGD): facilitating mouse as a model for human biology and disease. Nucleic Acids Res 43:D726–D736
Motenko H, Neuhauser SB, O'Keefe M, Richardson JE (2015) MouseMine: a new data warehouse for MGI. Mamm Genome 26:325–330
Smith RN, Aleksic J, Butano D, Carr A, Contrino S, Hu F, Lyne M, Lyne R, Kalderimis A, Rutherford K, Stepan R, Sullivan J, Wakeling M, Watkins X, Micklem G (2012) InterMine: a flexible data warehouse system for the integration and analysis of heterogeneous biological data. Bioinformatics 28:3163–3165
Richardson JE, Bult CJ (2015) Visual annotation display (VLAD): a tool for finding functional themes in lists of genes. Mamm Genome 26:567–573
Köhler S, Doelken SC, Mungall CJ, Bauer S, Firth HV, Bailleul-Forestier I, Black GC, Brown DL, Brudno M, Campbell J, FitzPatrick DR, Eppig JT, Jackson AP, Freson K, Girdea M, Helbig I, Hurst JA, Jähn J, Jackson LG, Kelly AM, Ledbetter DH, Mansour S, Martin CL, Moss C, Mumford A, Ouwehand WH, Park SM, Riggs ER, Scott RH, Sisodiya S, Van Vooren S, Wapner RJ, Wilkie AO, Wright CF, Vulto-van Silfhout AT, de Leeuw N, de Vries BB, Washingthon NL, Smith CL, Westerfield M, Schofield P, Ruef BJ, Gkoutos GV, Haendel M, Smedley D, Lewis SE, Robinson PN (2014) The Human Phenotype Ontology project: linking molecular biology and disease through phenotype data. Nucleic Acids Res 42:D966–D974
Richardson JE (2006) fjoin: simple and efficient computation of feature overlaps. J Comput Biol 13:1457–1464
Zhu Y, Richardson JE, Hale P, Baldarelli RM, Reed DJ, Recla JM, Sinclair R, Reddy TBK, Bult CJ (2015) A unified gene catalog for the laboratory mouse reference genome. Mamm Genome 26:295–304
Blake JA, Bult CJ, Eppig JT, Kadin JA, Richardson JE, Mouse Genome Database Group (2014) The Mouse Genome Database: integration of and access to knowledge about the laboratory mouse. Nucleic Acids Res 42:D810–D817
Smith CL, Eppig JT (2012) The Mammalian Phenotype Ontology as a unifying standard for experimental and high-throughput phenotyping data. Mamm Genome 23:653–668
Smith CL, Eppig JT (2015) Expanding the mammalian phenotype ontology to support automated exchange of high throughput mouse phenotyping data generated by large-scale mouse knockout screens. J Biomed Semantics 6:11
Bello SM, Eppig JT, MGI Software Group (2016) Inferring gene-to-phenotype and gene-to-disease relationships at mouse genome informatics: challenges and solutions. J Biomed Semantics 7:14
Acknowledgements
MGI is supported by the following NIH grants: HG000330 and HG002273 from the National Human Genome Research Institute; HD062499 from the Eunice Kennedy Shriver National Institute of Child Health and Human Development; CA089713 from the National Cancer Institute; OD011190 from the Office of the Director, Division of Comparative Medicine; and NS082666 from the National Institute of Neurological Disorder and Stroke.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Eppig, J.T. et al. (2017). Mouse Genome Informatics (MGI): Resources for Mining Mouse Genetic, Genomic, and Biological Data in Support of Primary and Translational Research. In: Schughart, K., Williams, R. (eds) Systems Genetics. Methods in Molecular Biology, vol 1488. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6427-7_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6427-7_3
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6425-3
Online ISBN: 978-1-4939-6427-7
eBook Packages: Springer Protocols