Abstract
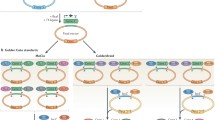
Biopart Assembly Standard for Idempotent Cloning (BASIC) is a simple, accurate, and robust DNA assembly method. The method is based on linker-mediated DNA assembly and provides highly accurate DNA assembly with 99 % correct assemblies for four parts and 90 % correct assemblies for seven parts [1]. The BASIC standard defines a single entry vector for all parts flanked by the same prefix and suffix sequences and its idempotent nature means that the assembled construct is returned in the same format. Once a part has been adapted into the BASIC format it can be placed at any position within a BASIC assembly without the need for reformatting. This allows laboratories to grow comprehensive and universal part libraries and to share them efficiently. The modularity within the BASIC framework is further extended by the possibility of encoding ribosomal binding sites (RBS) and peptide linker sequences directly on the linkers used for assembly. This makes BASIC a highly versatile library construction method for combinatorial part assembly including the construction of promoter, RBS, gene variant, and protein-tag libraries. In comparison with other DNA assembly standards and methods, BASIC offers a simple robust protocol; it relies on a single entry vector, provides for easy hierarchical assembly, and is highly accurate for up to seven parts per assembly round [2].
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Storch M, Casini A, Mackrow B et al (2015) BASIC: a new biopart assembly standard for idempotent cloning provides accurate, single-tier DNA assembly for synthetic biology. ACS Synth Biol 4(7):781–787. doi:10.1021/sb500356
Casini A, Storch M, Baldwin G, Ellis T (2015) Bricks and blueprints: methods and standards for DNA assembly. Nat Rev Mol Cell Biol 16:568–576. doi:10.1038/nrm4014
Casini A, Macdonald JT, Jonghe JD et al (2013) One-pot DNA construction for synthetic biology: the Modular Overlap-Directed Assembly with Linkers (MODAL) strategy. Nucleic Acids Res 42:e7. doi:10.1093/nar/gkt915
Casini A, Christodoulou G, Freemont P et al (2014) R2oDNA designer: computational design of biologically neutral synthetic DNA sequences. ACS Synth Biol 3:525–528. doi:10.1021/sb4001323
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28. doi:10.1016/0378-1119(90)90336-P
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Storch, M., Casini, A., Mackrow, B., Ellis, T., Baldwin, G.S. (2017). BASIC: A Simple and Accurate Modular DNA Assembly Method. In: Hughes, R. (eds) Synthetic DNA. Methods in Molecular Biology, vol 1472. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6343-0_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6343-0_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6341-6
Online ISBN: 978-1-4939-6343-0
eBook Packages: Springer Protocols