Abstract
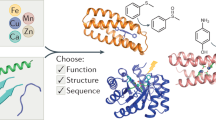
Multinuclear metal ion clusters, coordinated by proteins, catalyze various critical biological redox reactions, including water oxidation in photosynthesis, and nitrogen fixation. Designed metalloproteins featuring synthetic metal clusters would aid in the design of bio-inspired catalysts for various applications in synthetic biology. The design of metal ion-binding sites in a protein chain requires geometrically constrained and accurate placement of several (between three and six) polar and/or charged amino acid side chains for every metal ion, making the design problem very challenging to address. Here, we describe a general computational method to redesign oligomeric interfaces of symmetric proteins for the purpose of creating novel multinuclear metalloproteins with tunable geometries, electrochemical environments, and metal cofactor stability via first and second-shell interactions.
The method requires a target symmetric organometallic cofactor whose coordinating ligands resemble the side chains of a natural or unnatural amino acid and a library of oligomeric protein structures featuring the same symmetry as the target cofactor. Geometric interface matches between target cofactor and scaffold are determined using a program that we call symmetric protein recursive ion-cofactor sampler (SyPRIS). First, the amino acid-bound organometallic cofactor model is built and symmetrically aligned to the axes of symmetry of each scaffold. Depending on the symmetry, rigid body and inverse rotameric degrees of freedom of the cofactor model are then simultaneously sampled to locate scaffold backbone constellations that are geometrically poised to incorporate the cofactor. Optionally, backbone remodeling of loops can be performed if no perfect matches are identified. Finally, the identities of spatially proximal neighbor residues of the cofactor are optimized using Rosetta Design. Selected designs can then be produced in the laboratory using genetically incorporated unnatural amino acid technology and tested experimentally for structure and catalytic activity.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
- Metalloprotein
- Metalloenzyme design
- Multinuclear metal site
- Unnatural amino acid
- 2,2′-Bispyridine
- Computational design
1 Introduction
Much progress has been made in the last two decades toward the de novo design of novel metalloproteins [1–9], where the guiding principle is simultaneous placement of two or more metal coordinating side chain groups from naturally occurring amino acid residues, cysteines, aspartate and glutamate, and histidine residues. However, successful design attempts have been largely dominated by mononuclear (a single metal ion per designed protein) insertions into a single type of scaffold—the geometrically well defined alpha helical bundles [3]. One of the challenges while designing a multinuclear (metal ion site composed of two or more metal ions) metalloproteins is the need to incorporate multiple side chain coordinating groups in close spatial proximity in a single protein—placing exacting constraints on design. Another challenge is the design of the electrostatic environment of the metal ions, which has a large impact on the stability of the highly charged cofactor and the associated catalytic activity.
Computational algorithms could, in principle, aid in addressing both challenges. We previously developed an algorithm that utilized the metal-chelating unnatural amino acid 2,2′-bispyridyl alanine (BPY) [10, 11] for designing mononuclear metal-binding sites [9]. The algorithm uses RosettaMatch [12] to combinatorially search, in a given protein scaffold (typically a single chain), for a constellation of backbone structures that can support the multiple (~3–6) side chain metal-chelating functional groups in the appropriate coordination geometry. The use of BPY simplified the combinatorial design problem as, unlike any natural amino acid side chain, the bipyridyl moiety contributes two metal ligands from the same amino acid side chain. Metalloproteins featuring BPY with His and Asp/Glu residues were designed, and their crystallographic structure demonstrated close agreement with the design model. However, this algorithm is limited by its combinatorial complexity and is not applicable, practically, to construct multinuclear metal-binding sites.
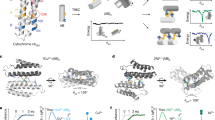
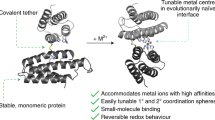
Here, we describe an approach to computationally design incorporation a symmetric multinuclear metallo-cofactor via integration into a similarly symmetric protein scaffold (Fig. 1). For this task, we have developed a matching algorithm, symmetric protein recursive ion-cofactor sampler (SyPRIS), and implemented it in Python. This algorithm allows expanding metalloprotein design to scaffolds other than alpha helical bundles, as well as gaining access to a greater variety of symmetric multinuclear cofactors such as iron-sulfur clusters and cubane complexes. We illustrate the method by describing the incorporation of the D2 symmetric cobalt-oxygen cube-like cofactor (Co-cubane) [13–20]. This cofactor is a mimic of the water oxidation center in photosystem II and features four bipyridyl moieties coordinating four Co-ions, respectively. Though Co-cubane is used as an example, the method is generally applicable to incorporate all types of cofactors of either C or D symmetry within any complementary symmetric scaffold. Theozyme [21] matches generated from SyPRIS can be further designed with the enzyme design modules in the Rosetta macromolecular modeling software [12, 22–25] (Fig. 2).
Several target cofactors that this method was intended to implement using scaffolds of various symmetries. (a) Co4O4(Ac)2(bipyridine)4 converted from CCDC crystal structure to noncanonical amino acid-bound model featuring D2 symmetry. (b) Cu2(OH)2(bipyridine)2 converted to models featuring C2 symmetry. (c) CuOH(bipyridine)2 converted to models featuring C2 symmetry. (d) Fe4S4(Cys)4 cluster featuring D2 symmetry. (e) Cu(OH)2(His)4 featuring C4 symmetry
2 Methods
2.1 The General Pipeline for the Method (Fig. 3a) Includes the Following Steps (Also See Note 1 )
(a) SyPRIS flow chart starting from generating scaffold library and ultimately ending in designable or discarded match. (b) An example scaffold, part of a library, will be considered by SyPRIS for the incorporation of a target cofactor. (c) A target cofactor, in this case an oxocobalt cubane coordinated by bipyridine ligands, has been modified with the appended magenta atoms creating a noncanonical amino acid. (d) The rotameric degrees of freedom for the atoms comprising the new backbone are sampled recursively with a chi distribution file (or exhaustively if desired) and compared to that of nearby backbone residues of the scaffold. (e) If the matched residue is part of a loop and the match was not geometrically identical, the loop is remodeled. (f) Three residues upstream and downstream of the translated backbone position are remodeled using Generalized KIC in Rosetta. (g) A fully designed oligomeric interface showing incorporated cofactor
-
1.
Generate and standardize a symmetric scaffold library (Fig. 3b).
-
2.
Prepare a target cofactor for symmetric insertion (Fig. 3c).
-
3.
Use SyPRIS to identify inverse rotamer positions suitable for design (Fig. 3d).
-
4.
Perform kinematic loop closure on residue matches that reside within a loop secondary structure (Figs. 3e, f).
-
5.
Design the oligomeric interface with constraints (Fig. 3g).
-
6.
Revert extraneous residue mutations to favor wild-type sequence.
-
7.
Experimental validation through protein expression, purification, and crystallization (not discussed here).
2.2 Generate and Standardize Symmetric Scaffold Library
Potential protein scaffold candidates are selected from the RCSB protein databank to feature a given symmetry in the oligomeric protein, i.e., D2, C2, 3, 4…, etc. Search parameters include symmetry type, chain stoichiometry, expressibility in E. coli, 90 % sequence identity threshold, and <3.0 Å resolution (for structures determined by X-ray crystallography). From these constraints, a raw scaffold library is generated. More than 70 % of the scaffold files generated in this way contain asymmetries in the form of incomplete chains—due to missing electron density in the crystal structures. In order to use the symmetry package of the Rosetta suite, all input files must be composed of chains that are equal in both residue length and residue type. To correct the intrinsic asymmetries, a hybrid Smith-Waterman local alignment is performed on all combinations of chains, removing residues absent from other chains, until a single converging monomeric sequence and all its symmetric partner protomers in the structures are found.
2.3 Target Cofactor
Cofactors of interest include organometallic compounds containing ligands that resemble either canonical amino acids or previously characterized noncanonical amino acids. PDB files are generated for cofactors of interest using their crystal structures and, where needed, the programs Mercury 3.5 and ConQuest 1.17 from the Cambridge Crystallographic Database (CCDC). Small structural changes may be applied to the supplied atom positions to reduce asymmetries within the X-ray crystallographic models. If necessary, backbone atoms are appended to each symmetric ligand, and all dihedrals are set to a default 0.0° prior to matching. To identify dihedral positions acceptable for each cofactor, an ensemble is generated of all dihedral rotations while simultaneously performing internal atomic clash checks. Dihedral rotations that pass the clash check are stored and plotted against each subsequent dihedral rotation within a heat map. Preferred geometries are classified as regions of the heat map with the highest bin density at a determined threshold. These geometric constraints are then converted into a “chi distribution” file necessary for the symmetric protein recursive ion sampler (SyPRIS). A chi distribution file depicts the four atoms participating in a dihedral rotation, a range of values between which to sample, and the degree with which to iterate. A Rosetta parameter file, which stores information about the asymmetric unit of the multinuclear cluster (i.e., one Co-ion and one oxygen atom for the Co-cubane, one Fe and one S atom for an iron-sulfur cluster), is defined for integration within the Rosetta suite during design. Lastly, a Rosetta enzyme design constraints file, which adds an energy term favoring the coordination geometry between ligand and complex, is generated to more accurately determine the energy of the integrated cofactor.
2.4 Symmetric Protein Recursive Ion Sampler (SyPRIS)
With the scaffold set and cofactor model in place, the following steps are utilized in finding symmetric matches between the cofactor coordinated to an UAA and the protein scaffold.
2.4.1 Align Scaffold and Cofactor Axes of Symmetry
-
1.
The axis of symmetry for the scaffold protein and each cofactor are determined by finding the eigenvector and eigenvalues—multiplying the coordinate matrix by its transpose matrix. Consequently, this creates unit vectors for each set of coordinates and supplies the principal rotational axes defined as the eigen minimum and maximum and their orthogonal cross product. In C-symmetry proteins, the eigen minimum and maximum can each be the target axis of symmetry. To correctly identify the axis of symmetry in a C-system, the midpoint of all symmetric Cα atoms is generated, and the average of all vectors connecting atoms to the origin becomes the symmetric axis.
-
2.
Translate all Cartesian atoms of all files so that the axis of symmetry origin of the scaffold and each model lie on a theoretical (0, 0, 0) origin.
-
3.
Align the axes of symmetry of the complex so that the eigen maximum and eigen minimum are aligned with that of the given scaffold (Fig. 4b). In C-symmetry, the eigen minimum of the cofactor is aligned to the midpoint average vector generated in step 1.
Fig. 4 (a) Residues that satisfy user-specified distance from symmetric axis highlighted in red sticks. (b) Rigid body rotation about symmetric axis to align symmetric axes. (c) Pictorial view of the enumerative exhaustive backbone sampling (left ). Schematic view of the recursive atom placing algorithm for direct matching (right ). (d) Ensemble of backbone positions generated via the recursive method. (e) A matched cofactor output from SyPRIS ready for Rosetta Design
-
4.
If the input features C-symmetry, SyPRIS will locate the midpoint of the Cβ atoms of the cofactor and translate to the midpoint of each protein Cβ combination that is within ± <user input (default = 1.0) > Å of the cofactor Cβ radii (Fig 3a). The cofactor is then rotated about the plane of symmetry until the Cβ atoms of both the cofactor and protein are aligned (Fig 3b). Each rotational/translational position unique to a residue subset will store the lowest atom magnitude difference position as well as two other rotational positions clockwise and counterclockwise to the aligned atoms within a < user input (default = 1.0) > Å direct distance. The four unaligned positions will be stored to further generate an ensemble of positions and dihedrals starting from step 6, below.
-
5.
If the input features d-symmetry, SyPRIS will perform 90° and 180° rotations of the cofactor about the vectors that correspond to each of the defined symmetric axes. Each rotational position will be further sampled in step 6.
2.4.2 Sample Inverse Rotamers
-
1.
A cofactor to scaffold backbone clash check is performed by determining distances between all heavy atoms of the cofactor not included in the chi distribution file and the backbone heavy atoms of nearby residues (not including the residue making the match ± one residue position proximal in sequence). Any distances to heavy atoms < user input (default = 2.8 Å) are considered clashes and discarded.
-
2.
For each unique cofactor rotation, cofactor backbone atoms (branches) are rotated within the range of values about the bonds defined by the atoms in the chi distribution file.
-
3.
To score a given rotation, a vector is produced from the last stationary atom (LASA) to the first atom changing location (FACL). For example, while rotating about a chi1 bond of BPY UAA, the LASA is the alpha carbon, while the FACL would be the backbone nitrogen atom. The vector produced by the LASA and FACL of the cofactor is compared to that of the scaffold. The angle difference is calculated as an AngleLog:
$$ \mathrm{AngleLog}= \log \left(\varSigma \varDelta \left[\left({ \cos}^{-1}\left(<xyz>\bullet <xy{z}^{\prime }>/\left|\right|xyz\left|\right|\times \left|\right|xy{z}^{\prime}\left|\right|\right)n/20\times n\right)\right]\right) $$where n is the number of compared vectors and a value of zero is an average deviation of 20° across all n vectors. To further score a matched position, the magnitude of the cofactor FACL to the compared scaffold atom is calculated. The default threshold for AngleLog and atom magnitude is < user input (default = 0.0) > and < user input (default = 0.8) > Å, respectively.
-
4.
Enumerative sampling. A predefined ensemble of inverse rotameric states is stored within one cofactor file. Each state is sampled exhaustively (Fig. 4c, left).
-
5.
Recursive sampling. For any range of values tested in the chi distribution file, the best scoring rotation (as long as it meets the thresholds) is stored along with the best adjacent rotation. Recursive ½ angles are sampled within this range to minimize to the best solution. The algorithm to locate new half dihedrals:
$$ \mathrm{A})\kern1em {\left({\varphi}_o+{\varphi}_n/2\right)}^n\kern1em \mathrm{or}\kern1em \mathrm{B})\kern1em {\left({\varphi}_{n-1}+{\varphi}_n/2\right)}^n $$where n is the number of half angles calculated as set by the user, φo is first dihedral (best scored), and n = 1 is the best scoring adjacent dihedral. SyPRIS starts with the algorithm in A. If two of the newly calculated half angles score better than the original dihedral, the B algorithm takes over for subsequent tests. Only the φ o, φ 1, and φ n (n = max) FACL rotated branches will be stored to further sample a wider ensemble of positions (Fig. 4c, right). This algorithm occurs for each subsequent torsion angle at all stored positions (3^# of chis). Therefore, a cofactor with three chis featuring D2 symmetry will store 27 positions (with tunable tolerance) at a given rotation. A C2 cofactor with the same number of chis will store up to five times this many positions due to the rigid body rotational degrees of freedom (Fig. 4d).
-
6.
For both the recursive and enumerative methods, final matches are determined by scoring the average AngleLog and RMSD over all FACL atom positions as defined in step 8 (Fig. 4e).
-
7.
A table for each protein is generated containing all the intrinsic properties of the ion cluster at a given match—model number and rotation about an axis. The table also includes the residue matched within the scaffold, the average AngleLog score, each individual AngleLog for all chains, the RMSD for all compared atoms, and the scaffold name. If an exact match is found (priority 1 designs), the scaffold will be mutated at the given residue position and passed to Rosetta Design. All other matches are subjects for the KIC procedure (priority 2 designs).
2.5 Kinematic Loop Closure (KIC)
This predesign method takes the tables generated by SyPRIS and locates the preferred residues for replacement with the ligand-like amino acid within the protein scaffold. The secondary structure of that residue with ± <user input (default = 3) > residues is determined based on Ramachandran preferred angles of phi and psi using a standard DSSP check. If the query within the scaffold is a loop region, the scaffold is accepted as designable; otherwise, if the region is helical or forms beta sheets, the scaffold is rejected. The scaffolds containing loops at match locations are then subjects of programs that:
-
1.
Take the scaffold and corresponding model as arguments.
-
2.
Translate the backbone coordinates of the matched residue on the scaffold to the location of the model to ensure exact match (generally changing atom positions by 0.5 Å across the entire residue).
-
3.
Generate a coordinate constraint file (see Note 2 ) of the heavy atoms comprising the multinuclear cluster in the model corresponding to chain A for use during design. A coordinate constraint (CST) file contains coordinates that ensures that the metal cluster atoms do not change positions during design.
-
4.
Generate two “loops” files (upstream and downstream of the matched residue) specific to each scaffold and matching residues necessary for performing KIC. The loop file contains information for which residue backbones will be sampled to make connection to another end point residue (i.e., remodeling the upstream or downstream loop about the ligand-like residue).
-
5.
Utilizing a Rosetta-generalized KIC [26, 27], the four residues upstream and downstream are remodeled to accommodate the new position of the matched residue (step II). The remodeling includes sampling of backbone phi and psi angles while progressively closing the chain break. More details can be found in Kortemme et al.
-
6.
A deterministic de novo loop is generated for each use of generalized KIC.
-
7.
Generated loops are evaluated based on void formation, electrostatic repulsion, etc.
2.6 Rosetta Design
All redesigned loop scaffolds that pass are subject to four rounds of rotamer sampling followed by gradient-based minimization of side chain and backbone atoms. Design and repack shells are defined as residues with Cα atoms within 12 and 16 Å radii, respectively, about the matched residue. The design shell specifies that all residues within the shell excluding the metal cofactor and UAA will be allowed to mutate to other more favorably scoring residues. Residues within the repack shell sample their rotameric preferred side chain conformations while keeping their identity fixed. The talaris2013 symmetric score function with constraints is used to evaluate the states of the protein during design. The coordinate constraint file generated in step 3 of Subheading 2.5 is used to force the ligand-like residue into a conformation conducive for coordinating the ions of the cofactor. The symmetry definition file generated in stage 2 was used to copy any change made on the master unit to all slave units as defined by Rosetta symmetry. Backbone minimization is allowed for residues that are part of the UAA-containing loop and nearby residues. Heavy coordinate constraints are placed on the scaffold to only allow movement of backbone atoms if necessary due to redesigned loop clashes. Final designs are chosen by low backbone RMSD of the design shell, smallest change to void volume, and favorable energies of interaction of the design shell residues with the cofactor (see Notes 3 and 4 ). Lastly, reversions are made on extraneous residues (see Note 5 ) to favor the wild-type sequence, and the protein is ready for expression (Fig. 5).
Two designs incorporating a catalytic D2 symmetric organometallic cofactor (Co4O4(Ac)2(bipyridine)4). The noncanonical amino acid bipyridine is incorporated on one chain, forming the cofactor upon oligomerization. The design protein (green and white) is compared to the wild-type scaffold (wheat). Mutation positions are represented by sticks
3 Notes
-
1.
All Python scripts and skeleton RosettaScripts XML files are attached.
-
2.
The Rosetta force field, as other molecular mechanics force fields, does not accurately model interactions of protein functional groups with metal ions. Therefore, it is necessary to treat these interactions with restraints. The weights used in the restraints will be system dependent, but in the final models, one should end up with a metal site geometry similar to the one from the starting crystal structure with some small deviation. If the metal site is completely distorted, the weights of the restraints should be increased to keep the geometry fixed.
-
3.
Another metric that is currently evaluated by human intuition in our protocol is that access of small ions/substrates to the metal site has not been blocked by new mutations introduced in the design protocol. Conformational changes upon substrate binding are not modeled, and system-dependent knowledge of the dynamics of the closure and opening of the active site should be kept in mind when either picking out scaffolds for design and evaluating designs by inspection.
-
4.
Many substitutions can be introduced, but as a designer, one should also make sure that the initial protein scaffold can accommodate these changes in the absence of any substrate; otherwise, the enzyme will either not express or be unfolded. In particular, we paid special attention to the maintenance of the symmetric interface of the oligomer in question.
-
5.
Chemical intuition is almost always required to evaluate the goodness of designs.
References
Ghosh D, Pecoraro VL (2004) Understanding metalloprotein folding using a de novo design strategy. Inorg Chem 43:7902–7915. doi:10.1021/ic048939z
Hellinga HW (1996) Metalloprotein design. Curr Opin Biotechnol 7:437–441. doi:10.1016/S0958-1669(96)80121-2
Peacock AFA (2013) Incorporating metals into de novo proteins. Curr Opin Chem Biol 17:934–939. doi:10.1016/j.cbpa.2013.10.015
Zastrow ML, Pecoraro VL (2013) Designing functional metalloproteins: from structural to catalytic metal sites. Coord Chem Rev 257:2565–2588. doi:10.1016/j.ccr.2013.02.007
Lu Y, Yeung N, Sieracki N, Marshall NM (2009) Design of functional metalloproteins. Nature 460:855–862. doi:10.1038/nature08304
Grzyb J, Xu F, Weiner L et al (2010) De novo design of a non-natural fold for an iron-sulfur protein: Alpha-helical coiled-coil with a four-iron four-sulfur cluster binding site in its central core. Biochim Biophys Acta Bioenerg 1797:406–413. doi:10.1016/j.bbabio.2009.12.012
DeGrado WF, Summa CM, Pavone V et al (1999) De novo design and structural characterization of proteins and metalloproteins. Annu Rev Biochem 68:779–819. doi:10.1146/annurev.biochem.68.1.779
Degrado WF, Summa CM, Pavone V et al (1999) De novo design and structural characterization of proteins. Biochemistry 68:779–819
Mills JH, Khare SD, Bolduc JM et al (2013) Computational design of an unnatural amino acid dependent metalloprotein with atomic level accuracy. J Am Chem Soc 135:13393–13399. doi:10.1021/ja403503m
Liu CC, Schultz PG (2010) Adding new chemistries to the genetic code. Annu Rev Biochem 79:413–444. doi:10.1146/annurev.biochem.052308.105824
Imperiali B, Fisher SL (1991) (S)-u-amino-2,2′-bipyridine-6-propanoic acid: a versatile amino acid for de novo metalloprotein design. J Am Chem Soc 113:8527–8528. doi:10.1021/ja00022a053
Richter F, Leaver-Fay A, Khare SD et al (2011) De novo enzyme design using Rosetta3. PLoS One 6:1–12. doi:10.1371/journal.pone.0019230
Smith PF, Kaplan C, Sheats JE et al (2014) What determines catalyst functionality in molecular water oxidation? Dependence on ligands and metal nuclearity in cobalt clusters. Inorg Chem 53:2113–2121. doi:10.1021/ic402720p
Li X, Clatworthy EB, Masters AF, Maschmeyer T (2015) Molecular cobalt clusters as precursors of distinct active species in electrochemical, photochemical, and photoelectrochemical water oxidation reactions in phosphate electrolytes. Chemistry 21(46):16578–16584. doi:10.1002/chem.201502428
Dimitrou K, Brown AD, Christou G et al (2001) Mixed-valence, tetranuclear cobalt(iii, iv) complexes: preparation and properties of [Co4O4(O2CR)2(bpy)4]3+ salts. Chem Commun 4:1284–1285. doi:10.1039/b102008k
Evangelisti F, Guettinger R, More R et al (2013) Closer to photosystem II: A Co4O4 cubane catalyst with flexible ligand architecture. J Am Chem Soc 135(50):18734–18737. doi:10.1021/ja4098302
McCool NS, Robinson DM, Sheats JE, Dismukes GC (2011) A Co4O4 cubane water oxidation catalyst inspired by photosynthesis. J Am Chem Soc 133:11446–11449. doi:10.1021/ja203877y
Berardi S, La Ganga G, Natali M et al (2012) Photocatalytic water oxidation: tuning light-induced electron transfer by molecular Co4O4 cores. J Am Chem Soc 134:11104–11107. doi:10.1021/ja303951z
Chakrabarty R, Bora SJ, Das BK (2007) Synthesis, structure, spectral and electrochemical properties, and catalytic use of cobalt (III)−oxo cubane clusters. Polyhedron 46:9450–9462
Najafpour MM, Rahimi F, Aro E-M et al (2012) Nano-sized manganese oxides as biomimetic catalysts for water oxidation in artificial photosynthesis: a review. J R Soc Interface 9:2383–2395. doi:10.1098/rsif.2012.0412
Tantillo DJ, Chen J, Houk KN (1998) Theozymes and compuzymes: theoretical models for biological catalysis. Curr Opin Chem Biol 2:743–750. doi:10.1016/S1367-5931(98)80112-9
Siegel JB, Zanghellini A, Lovick HM et al (2010) Computational design of an enzyme catalyst for a stereoselective bimolecular diels-alder reaction. Science 105:1–6
Röthlisberger D, Khersonsky O, Wollacott AM et al (2008) Kemp elimination catalysts by computational enzyme design. Nature 453:190–195. doi:10.1038/nature06879
Jiang L, Althoff EA, Clemente FR et al (2008) De novo computational design of retro-aldol enzymes. Science 319:1387–1391. doi:10.1126/science.1152692
Bradley P, Misura KMS, Baker D (2005) Toward high-resolution de novo structure prediction for small proteins. Science 309:1868–1871. doi:10.1126/science.1113801
Mandell DJ, Kortemme T (2009) Backbone flexibility in computational protein design. Curr Opin Biotechnol 20:420–428. doi:10.1016/j.copbio.2009.07.006
Mandell DJ, Coutsias EA, Kortemme T (2009) Sub-angstrom accuracy in protein loop reconstruction by robotics-inspired conformational sampling. Nat Methods 6:551–552. doi:10.1038/nmeth0809-551
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Hansen, W.A., Mills, J.H., Khare, S.D. (2016). Computational Design of Multinuclear Metalloproteins Using Unnatural Amino Acids. In: Stoddard, B. (eds) Computational Design of Ligand Binding Proteins. Methods in Molecular Biology, vol 1414. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3569-7_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3569-7_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3567-3
Online ISBN: 978-1-4939-3569-7
eBook Packages: Springer Protocols