Abstract
AlphaScreen® SureFire® assay is a novel technology that combines luminescent oxygen channeling technology, nano-beads, and monocloncal antibodies to detect the level of a selected protein in a volume lower than 5 μl. This method is more sensitive compared with the traditional enzyme-linked immunosorbent assays (ELISA), and can detect an increasing number of new targets. Here, we described a method for AlphaScreen® SureFire® assay that targets ERK1/2 phosphorylation, a primary downstream signaling pathway that conveys activation of GPR55 by l-α-lysophosphatidylinositol (LPI) and certain cannabinoids.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
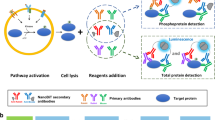
AlphaScreen® SureFire® assay is an alternative technology to the traditional ELISA (Fig. 1).
It is faster and more sensitive than ELISA, requires a micro-quantity of the tested sample, and can be performed in 384 well plates (Fig. 2). This upscale enables the translation to high-throughput screening platforms that employ robotic devices at drug discovery laboratories.
AlphaScreen® SureFire® assay has been applied to study the level of phosphorylated proteins in various fields of research, including GPCR signaling, epigenetics, kinase research, and protein-protein interactions. Using this method, we have recently studied the pharmacological actions of synthetic cannabinoids and phytocannabinoids at the human GPR55 receptor [1]. The latter is activated by the endogenous lipid LPI [1–4], and appears to highly regulate cancer cell function. In selected cancer cells, GPR55 is highly expressed and its activation by LPI increases cell migration, invasion, and proliferation [5–7]. Based on these studies, many resources have been allocated to identify GPR55 inhibitors [1, 8]. Other important clinical targets for identifying GPR55 inhibitors emerged from studies which showed that activation of GPR55 contributes to inflammation and enhances neuropathic pain [9]. We have identified GPR55 enhancers, inhibitors, and modulators of LPI-induced activation of human GPR55 [1]. Our results also suggest that GPR55 has an allosteric binding pocket and point to a cross-talk between the cellular messengers of NFAT-ERK1/2 signaling cascades [1]. Another example is related to the lectin-like oxidized low-density lipoprotein (oxLDL) receptor-1 (LOX-1) [10]. The latter has been shown to directly interact with C-reactive protein (CRP) in addition to binding to oxLDL [10]. These findings suggest that CRP, a risk factor for cardiovascular events, promotes endothelial dysfunction and amplifies vascular inflammation by engaging LOX-1.
2 Materials
-
1.
Drugs. All compounds were dissolved in DMSO, and 10 mM stocks were kept at −20 °C. LPI was stored at −80 °C for up to 3 months.
-
2.
Cells. Untransfected HEK293 cells are grown in Dulbecco’s modified Eagle’s medium (Gibco) containing 2 mM l-glutamine medium (Gibco), and 10 % fetal bovine serum. Cells are slit at a ratio of 1:2 or 1:4, according to the required cell density. For HEK293 cells that stably express the tagged-human GPR55 receptor (hGPR55-HEK293), the GPR55 receptor is tagged with a triple hemagglutinin epitope (HA) at the N terminus (3xHA-GPR55), preceded by the signal sequence from the human growth hormone (HGH; residues 1–33), and is subcloned into pcDNA 3.1 vector. Transfected hGPR55-HEK293 cells are grown in a medium prepared with 500 ml of Dulbecco’s modified Eagle’s medium/F12 (Gibco), 50 ml of 10 % new born calf serum (Gibco), 5 ml of G-418 (50 mg/ml), 5 ml of l-glutamine (Sigma), and 3 ml of penicillin/streptomycin solution (containing 10,000 units of penicillin (base) and 10,000 μg/ml of streptomycin from Gibco).
-
3.
Serum-free Medium. For transfected cells, add to DMEM/F12 medium 5 ml of G418 and 5 ml of l-glutamine. For untransfected cells, add to DMEM/F12 medium 5 ml of l-glutamine without G418.
-
4.
Assay Medium. Prepare serum-free/phenol-free “assay medium” with Dulbecco’s modified Eagle’s medium/F-12 containing 2 nM l-Glutamine (Gibco).
-
5.
AlphaScreen ® Kit. AlphaScreen® SureFire™ Phospho-ERK½ is from PerkinElmer, and its components are listed in Table 1.
Table 1 Materials supplied with the kit -
6.
Positive/Negative Control Lysates. Lysates are provided with the AlphaScreen® Kit. Reconstitute them with double-distilled water (DDW) according to the manufacturer’s instructions, and freeze aliquots at −20 °C.
-
7.
Plates. Perform the assay in 384 well white ProxiPlates according to the manufacturer’s instructions.
3 Methods
-
1.
Cell Maintenance. Add cells to 10 ml of medium, centrifuge for 5 min at 200 × g, remove supernatant, add 10 ml of fresh growth medium, seed in a T75 flask, place in the incubator in a humidified atmosphere at 37 °C and 5 % CO2; refresh medium after 24 h.
-
2.
Plating hGPR55-HEK Cells for Assay. The following protocol yields about five 96 well plates. Harvest cells from two T75 flasks by washing once with dissociation buffer (from Sigma), add 10 ml of dissociation buffer, collect cells into a 50 ml tube, add 10 ml of growth medium, centrifuge at 200 × g for 3 min, resuspend in 20 ml of growth medium. Count cells, seed 40,000 cells/well in a 96 well plate. Cells should be 100 % confluent on the next day.
-
3.
Prepare hGPR55-HEK for Assay. Remove the growth medium from the cells, and replace with 100 μl of serum-free medium. Serum-starve the cells for 48 h. Both media should be warmed up to 37 °C (see Note 1 ).
-
4.
ERK1/2 MAP-kinase Phosphorylation Assay
Follow these steps:
-
1.
Design your assay plate (method described below is for a 96 well plate).
-
2.
Warm up the assay medium to 25 °C.
-
3.
Prepare 1× lysis buffer by diluting 1:5 with DDW the lysis buffer ×5, supplied with the kit. For example, mix 1000 ml of lysis buffer with 4000 ml of DDW. Keep on ice (see Note 2 ).
-
4.
Prepare drugs at the required test dilutions. Keep the concentrations of the solvent at a constant level throughout the experiment. Typically, final concentrations of 0.1 or 0.2 % DMSO are used.
-
5.
Label plate and remove medium.
-
6.
Wash once with assay medium.
-
7.
Remove medium from cells.
-
8.
Add the tested drugs at the desire volume.
-
9.
Incubate at 37 °C. Typically, to measure LPI-induced ERK1/2 phosphorylation an incubation of 20 min is recommended. However, the time should be selected based on preliminary experiments.
-
10.
Remove medium from cells.
-
11.
Keep the plate on ice.
-
12.
Add 50 μl/well of 1× lysis buffer.
-
13.
Store at −80 °C for at least 1 h (see Note 3 ).
3.1 ERK1/2 Phosphorylation Assay
To test the level of ERK1/2 phosphorylation, follow these steps:
-
1.
Warm up Activation buffer from the kit to room temperature.
-
2.
Thaw the assay plate (from the above step 13) at room temperature.
-
3.
Transfer 4 μl of the lysed cells from each well into a well of the 384 ProxiPlate.
-
4.
Transfer 4 μl of control lysate (supplied with the kit) into a well of the 384 ProxiPlate.
-
5.
Prepare AlphaScreen® beads-containing ERK1/2 assay mix, as detailed in Table 2.
Table 2 ERK1/2 assay mix -
6.
Add 7 μl per well of ERK1/2 assay mix.
-
7.
Seal the ProxiPlate and protect from light.
-
8.
Incubate the ProxiPlate for 3.5 h at 23–25 °C in the dark.
-
9.
Incubate plate at room temperature and read with the Envision system (PerkinElmer), by using AlphaScreen® settings.
3.2 Data Analysis
Raw data can be presented as “Envision units,” defining basal level (in the presence of vehicle) as zero. Results are presented as means and variability as SEM or 95 % confidence limits (CL) of the percent stimulation of phosphorylated ERK1/2 above the basal level. Data can be analyzed by using nonlinear analysis of log agonist versus-response curves with Prism 5.0 program (GraphPad, San Diego, CA) (Fig. 3).
(a) Mean log concentration-response curves of LPI effect on ERK1/2 phosphorylation in hGPR55-HEK293 cells after 20 min (n = 4 each in triplicate). No significant differences are observed in basal levels of phosphorylated ERK1/2 in untransfected HEK293 cells and hGPR55-HEK293 cells incubated for 20 min in 0.1 % DMSO (n = 2 each in duplicate). (b) LPI-induced stimulation of ERK1/2 phosphorylation is attenuated by 10 μM PD98059, a MEK1 inhibitor, that significantly reduces basal pERK levels. *p < 0.05, **p < 0.01, ***p < 0.001 by means of one-sample t-test. (c) Mean log concentration-response curves of ERK1/2 phosphorylation after 20 min stimulation at 37 °C with Δ9-tetrahydrocannabinol (Δ9-THC) (n = 3). Each symbol represents the mean percentage change in bound phosphorylated ERK1/2 ± SEM over the basal level
When curves cannot be fitted by nonlinear analysis of log agonist versus-response curves, statistical significance of the stimulation can be determined with an unpaired Student’s t-test at each specific concentration. Results are considered significant only when F-test comparing the variance is not significantly different.
4 Notes
-
1.
All media should be warmed up to 37 °C, unless specified otherwise.
-
2.
Portions of the diluted lysis buffer can be stored frozen (−20 °C) for subsequent experiments.
-
3.
Lysates can be frozen (−80 °C) at this stage, to be assayed later.
References
Anavi-Goffer S, Baillie G, Irving AJ et al (2012) Modulation of L-alpha-lysophosphatidylinositol/GPR55 mitogen-activated protein kinase (MAPK) signaling by cannabinoids. J Biol Chem 287:91–104
Kotsikorou E, Sharir H, Shore DM et al (2013) Identification of the GPR55 antagonist binding site using a novel set of high-potency GPR55 selective ligands. Biochemistry 52:9456–9469
Heynen-Genel S, Dahl R, Shi S, et al (2010) Screening for selective ligands for GPR55 agonists in Probe Reports from the NIH Molecular Libraries Program (Internet), National Center for Biotechnology Information, Bethesda, MD, NBK66152 [bookaccession]
Henstridge CM, Balenga NA, Ford LA et al (2009) The GPR55 ligand L-alpha-lysophosphatidylinositol promotes RhoA-dependent Ca2+ signaling and NFAT activation. FASEB J 23:183–193
Andradas C, Caffarel MM, Perez-Gomez E et al (2011) The orphan G protein-coupled receptor GPR55 promotes cancer cell proliferation via ERK. Oncogene 30:245–252
Ford LA, Roelofs AJ, Anavi-Goffer S et al (2010) A role for L-alpha-lysophosphatidylinositol and GPR55 in the modulation of migration, orientation and polarization of human breast cancer cells. Br J Pharmacol 160:762–771
Pineiro R, Maffucci T, Falasca MT (2011) The putative cannabinoid receptor GPR55 defines a novel autocrine loop in cancer cell proliferation. Oncogene 30:142–152
Heynen-Genel S, Dahl R, Shi S, et al (2010) Screening for selective ligands for GPR55 antagonists in Probe Reports from the NIH Molecular Libraries Program (Internet), National Center for Biotechnology Information, Bethesda, MD, NBK66153 [bookaccession]
Staton PC, Hatcher JP, Walker DJ et al (2008) The putative cannabinoid receptor GPR55 plays a role in mechanical hyperalgesia associated with inflammatory and neuropathic pain. Pain 139:225–236
Shih HH, Zhang S, Cao W et al (2009) CRP is a novel ligand for the oxidized LDL receptor LOX-1. Am J Physiol Heart Circ Physiol 296:H1643–H1650
Acknowledgments
This work was supported by the US National Institutes of Health grant numbers DA-03672 and DA-09789 (RAR). We thank the University of Aberdeen for a Knowledge Transfer Grant Award.
Conflicts of interest: RAR received funding from GW Pharmaceuticals.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Anavi-Goffer, S., Ross, R.A. (2016). A Functional Assay for GPR55: Envision Protocol. In: Maccarrone, M. (eds) Endocannabinoid Signaling. Methods in Molecular Biology, vol 1412. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3539-0_8
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3539-0_8
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3537-6
Online ISBN: 978-1-4939-3539-0
eBook Packages: Springer Protocols