Abstract
MicroRNAs (miRNAs) are a class of small, noncoding RNAs which regulate gene expression by directing their target mRNA for degradation or translational repression. Since their discovery in the early 1990s, miRNAs have emerged as key components in the posttranscriptional regulation of gene networks, shaping many biological processes from development, morphogenesis, differentiation, proliferation and apoptosis. Although understanding of the molecular basis of miRNA biology is improving, methods to monitor the dynamic and the spatiotemporal aspects of miRNA expression under physiopathological conditions are required. However, monitoring of miRNAs is difficult due to their small size, low abundance, high degree of sequence similarity, and their dynamic expression pattern which is subjected to tight transcriptional and post-transcriptional controls. Recently, we developed a miRNA monitoring system called RILES, standing for RNAi-inducible expression system, which relies on an engineered regulatable expression system, to switch on the expression of the luciferase gene when the targeted miRNA is expressed in cells. We demonstrated that RILES is a specific, sensitive, and robust method to determine the fine-tuning of miRNA expression during the development of an experimental pathological process in mice. Because RILES offers the possibility for longitudinal studies on individual subjects, sharper insights into miRNA regulation can be generated, with applications in physiology, pathophysiology and development of RNAi-based therapies. This chapter describes methods and protocols to monitor the expression of myomiR-206, -1, and -133 in the tibialis anterior muscle of mice. These protocols can be used and adapted to monitor the expression of other miRNAs in other biological processes.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
- microRNA
- Molecular imaging
- Bioluminescence study
- Spatiotemporal regulation
- RILES (RNAi-inducible luciferase expression system )
1 Introduction
MiRNAs are a class of noncoding RNA molecules that control the expression of target genes at the posttranscriptional level [1]. In the canonical miRNA pathway, miRNA genes are transcribed by polymerase II and/or III from endogenous genes either from individual transcriptional units (intergenic miRNAs ) or in frame with host genes (intronic miRNAs). They are processed in the nucleus as long primary miRNA (pri-miRNA ) transcripts before being shortened as precursor (pre-miRNA ) transcripts of approximately 70 nucleotides by a first endoribonuclease complex consisting of Drosha and DGCR8 . The pre-miRNAs are exported into the cytoplasm by the Exportin-5/Ran-GTP complex and then cleaved by a second endoribonuclease complex, DICER/TRBP , to produce a 20–22-nucleotide-long duplex RNA molecule. This miRNA duplex consists of the mature miRNA, called the guide strand and a less thermostable strand, called the passenger strand which is believed to be degraded. The single-stranded mature miRNA interacts with Argonaute protein s before being loaded in the RLC (RISC loading complex) composed of DICER/TRBP and Argonaute 2 bound to the mature miRNA. In the RLC complex, the miRNA binds to the mRNA target sequence through a perfect (100 % homology) or imperfect (less than 100 % homology) complementary base pairing mechanism, often located on the 3′-UTR part of the target mRNAs. According to the degree of homology, the mRNA is either degraded or the translational machinery is blocked inducing mRNA deadenylation and/or decapping [1].
Research in the field of noncoding RNA has advanced rapidly since the discovery of miRNA in the early 1990s [2]. It is known that the human genome encodes for at least 1200 miRNA genes and that at least 60 % of the transcriptome is under the control of miRNAs. Therefore, it is not surprising that almost all biological processes are directly or indirectly influenced by miRNAs [3] and that deregulated expression of miRNAs has also been found to be associated with human diseases, thus paving the way for novel therapeutic interventions [4].
An important aspect of miRNA function is that their regulation is highly dynamic, subject to tight, spatial, and temporal transcriptional controls which in turn fine-tune the expression of gene networks [5, 6]. The underestimation of these features results in the loss of important information linking miRNA expression to cell function. It is therefore crucial to know the repertoire of miRNAs and how their expression changes in response to cellular environments in order to improve understanding of their biological functions.
Several detection methods have been developed to monitor miRNA expression. These include Northern-blot , real-time PCR, microarray, in situ hybridization, and more recently deep sequencing and biochemical pull-down approaches [7]. At the same time, over the last 10 years, noninvasive molecular imaging methods have been developed to monitor the expression of miRNAs, in real time and noninvasively in small animal models. These methods can be subtyped into two classes according to the nature of the probes used. The first class includes synthetic probes composed mainly of molecular beacons, consisting in fluorescent oligonucleotide sequences, which on binding to a complementary mature miRNA sequence can emit fluorescence. The second class includes fluorescent or bioluminescent reporter genes which once expressed in cells have their light-emission properties modulated by miRNAs. For the latter, the miR-OFF system is based on extinction of reporter gene expression by miRNA binding to miRNA target sequences cloned in the 3′-UTR of the reporter gene [8]. Nowadays, this miR-OFF system is widely used in vitro for functional studies to confirm the presence of potential miRNA binding sequences in the 3′-UTR or 5′-UTR of target mRNA . This system has also been found to be successful in monitoring biogenesis and miRNA activity during neuronal differentiation [9] and tumor development [8, 10] for instance. It has also been employed as a functional screening platform to identify miRNA candidates involved in regulation of tumor-associated mRNA [11].
More recently, we [12] have developed an alternative approach, a miR-ON system , to monitor positively the endogenous expression pattern of miRNAs in cells, tissues, and small animal models. In the strategy developed, the expression of the reporter gene is not suppressed by the miRNA but is instead induced. Therefore miRNA expression in cells is signed by the emission of optical signals which can be detected by standard fluorescence or bioluminescence equipment. The overall strategy relies on engineered regulatable expression systems, also known as genetic switches [13], such as the Tet-Krab , Tet R and the Cumate gene-switch systems [14]. The system is designed so that the transcriptional regulator molecule controlling the expression of a transgene driven by an inducible promoter is placed under control of a miRNA of interest. We engineered the Cumate operon system by subcloning in the 3′-UTR of the CymR transcriptional repressor cDNA , four complementary block sequences of a given miRNA (Fig. 1) [12]. Therefore, when the miRNA of interest is expressed in cells, it binds to the CymR mRNA inducing its degradation through the RLC machinery. In the absence of CymR repressor protein, the second transcription cassette, encoding for the luciferase gene, is transcribed, generating bioluminescence signals in cells. We recently demonstrated that this RNAi-inducing-luciferase expression system, which we named RILES , enabled the monitoring of ectopic and endogenously expressed miRNAs in cell lines and also in tissues of live anesthetized animals. We monitored the expression of miRNA-122 in the liver of mice and the differential expression pattern of myomiRs-133 , -1 and -206 expressed in the skeletal muscle of anterior tibialis of naive mice. Furthermore, we established for the first time in mice, the kinetic of a miRNA during the development of a pathological process over a period of 35 days without repeated administration of RILES probes. Using a model of muscular atrophy we were able to determine the kinetic of miRNA-206 expression during the muscle regeneration phase of muscle atrophy. The bioluminescence data indicated that the expression of miRNA-206 is individual dependent, finely regulated in a time-dependent manner, and characterized by individual heterogeneity during development of the pathology. When compared with data generated from conventional quantitative RT-PCR, miRNA-206 expression was also found to be over expressed but its expression remained constant for 7 days before returning to the basal level. This discrepancy between the two approaches is explained by the invasive nature of the quantitative RT-PCR method that generates a set of information from a heterogeneous population collected at different time points. This invasive method, in contrast to RILES, lacks temporal resolution at the individual level and thus undervalues the crucial significance of the temporal regulation of miRNA expression in response to biological processes [12].
Schematic representation of the RILES method. When present in cells, target miRNA binds to the four complementary-block sequences located in the 3′-untranslated region of the CymR repressor transcript and activates the RNAi silencing complex (RISC) machinery. The CymR mRNA is then cleaved and degraded, resulting in lack of repressor production. The luciferase expression system is thus switched on, generating a positive bioluminescence signal. Reprinted and adapted from [12]
In this chapter we describe in detail methods used to produce the RILES expression plasmid in a context of monitoring the expression of myomiR-206 , -133 , and -1 in the tibialis skeletal muscle of naïve mice and, as a control, the expression of miRNA-122, a liver-specific miRNA not expressed in skeletal muscle tissue. The protocols described include the preparation of the RILES plasmids, from the cloning of the miR targeting sequence in the 3′-UTR of the CymR transcriptional repressor to the functional validation of the RILES plasmids in HEK 293 cells. Finally, we describe the procedure to monitor the expression of myomiRs in the skeletal tibialis muscle of mice following the intramuscular injections of RILES plasmids. These protocols can be adapted to monitor any other miRNA in vitro or in vivo in other tissues as long as the RILES can be transfected to the tissue of interest.
2 Materials
2.1 RILES Plasmid Design and Production
-
1.
The RILES plasmid originates from the Cumate gene-switch , first described by Mullick et al. [14]. We prepared [12] an “all-in-one system” by assembling into a single plasmid unit, the expression cassette encoding for the CymR transcriptional repressor driven by the SV40 promoter and the inducible expression cassette encoding for the luciferase gene reporter driven by the CMV5(Cuo) inducible promoter. A multiple cloning site containing Mlu I and Not I restriction sites was inserted into the 3′-UTR of CymR cDNA , after the stop codon and before the polyadenylation site (Fig. 1) to insert a block of four perfect-match complementary sequences to miRNA (see Note 1 ). This block sequence, consisting of a double-stranded oligonucleotide, is called miR T sequence, standing for miRNA target sequence. Upon cloning, the RILES plasmid is denoted, for example pRILES/206T or pRILES/133T when the miR T sequence contained complementary block sequences to detect miRNA-206 or miRNA-133 respectively (Table 1).
Table 1 Nucleotide sequences of the miR target subcloned in the RILES plasmids -
2.
To prepare the specific miR T sequences to monitor myomiR-206, -133, -1, and miRNA-122 as a control (Table 1), synthetic oligonucleotides are designed to contain, in addition to the miRNA T (see Note 2 ), four different nucleotide sequences, located in between the four block sequences of the miR T cassette, to ensure optimal annealing sequence match between the sense and antisense oligonucleotides. The oligonucleotides are also flanked at the 3′-end by a compatible, overhanging, phosphorylated Not I sequence, and at the 5′-end by a compatible, overhanging, phosphorylated Mlu I sequence. The sense and antisense single-stranded oligonucleotides are obtained commercially (Eurogentec, Seraing, Belgium), PAGE purified (polyacrylamide gel purification method), and coupled with a phosphate group at the 5′-end of each oligonucleotides (see Note 3 ). The purified oligonucleotides are resuspended in nuclease-free water to generate a 100 μM stock solution and conserved at −20 °C.
-
3.
10× annealing buffer: 100 mM Tris–HCl pH7.5, 1 M NaCl, 10 mM EDTA.
-
4.
Water bath at 37 °C.
-
5.
T4 DNA ligase with buffer, Not I and Mlu restriction enzymes (NEB, Hitchin; UK).
-
6.
DH5 calcium-chloride-competent bacteria cells (stored at −80 °C) (NEB, Hitchin; UK).
-
7.
LB broth and LB agar plates containing 100 μg/ml ampicillin (Life Technologies SAS, St. Aubin, France).
-
8.
NucleoBond Xtra Mini and Maxi endotoxin-free extraction kits (Macherey-Nagel, Düren, Germany).
-
9.
Electrophoresis grade agarose (Sigma Aldrich, St. Louis, USA), agarose gel electrophoresis equipment (Biorad, Hercules, CA, USA), ethidium bromide solution 50 μg/ml (Sigma Aldrich, St. Louis, USA).
-
10.
10× gel loading buffer (0.25 % bromophenol bleu, 0.25 % xylene cyanol FF, 50 % glycerol), 1 kb and 100 bp ladders (NEB, Hitchin UK), 50× TAE buffer (Sigma Aldrich, St. Louis, USA) for agarose gel, Gel Doc™ XR+ gel imaging System (Biorad, Hercules, CA, USA).
2.2 Tissue Culture and Transfection
-
1.
HEK 293 cells (human embryonic kidney cell line, ATCC, CRL-1573) (see Note 4 ).
-
2.
Complete growth media: Dulbecco’s Modified Eagle Medium (DMEM, 4 g/l glucose, Life Technologies SAS, St. Aubin, France), supplemented with 10 % (vol/vol) of heat-inactivated fetal bovine serum (Life Technologies SAS, St. Aubin, France) and 100 I.U./ml penicillin and 100 μg/ml streptomycin (Life Technologies SAS, St. Aubin, France).
-
3.
2.5 % (vol/vol) Trypsin-EDTA, phosphate-buffered saline solution, opti-MEM medium and Trypan bleu 0.4 % (Life Technologies SAS, St. Aubin, France).
-
4.
Tissue culture flask (25 cm2), 24-well flat-bottom tissue culture plates, serological sterile polystyrene pipets, conical sterile polypropylene centrifuge tubes, hemocytometer (Starsted, Nümbrecht, Germany).
-
5.
ICAFectin™ 441 (In-cell-Art, Nantes, France, or Eurogentec France SASU, Angers, France).
-
6.
5 nmol of synthetic miRNA precursor (Pre-miR™ miRNA precursor, AM 17100, Life Technologies SAS, St. Aubin, France) specific to miRNA-133a (PM 10412), miRNA-122 (PM 17100) and miRNA-221 (PM 12613). Theses miRNA precursors are resuspended in 1 ml of nuclease-free water to obtain a 5 μM final stock solution (see Note 5 ).
-
7.
pQE30 plasmid (QIAGEN S.A.S, Courtaboeuf, France) (see Note 6 ).
-
8.
Luciferase Assay System with Reporter Lysis Buffer kit (Promega France, Charbonniere, France).
-
9.
AutoLumat LB 953 Multi-tube luminometer (Berthol France, Thoiry, France).
-
10.
Pierce™ BCA Protein Assay Kit (Life Technologies SAS, St. Aubin, France).
2.3 Animal Experiments and Bioluminescence Imaging
-
1.
8-week-old female Swiss or BALB/C mice (Harlan laboratories, Paris, France).
-
2.
Specific pathogen free (SPF) animal facility, litter, neutral pH water, individual ventilated cages, self-contained isoflurane-based anesthesia unit (FUJIFILM VisualSonics, Ontario, Canada), animal identification via ear notch (FUJIFILM VisualSonics, Ontario, Canada).
-
3.
Amphiphilic block copolymer 704 as in vivo transfection reagent [15].
-
4.
Tyrode salt solution (endotoxin free, Sigma Aldrich, St. Louis, USA).
-
5.
In vivo -grade luciferin substrate (VivoGLo, Promega France, Charbonniere, France).
-
6.
NightOWL LB in vivo imaging system (Berthol France, Thoiry, France) coupled with isoflurane-based anesthesia induction.
3 Methods
3.1 Molecular Cloning of myomiR T Sequences and Production of Recombinant RILES Plasmids
-
1.
To anneal the miR T sequence, in a 1.5 ml Eppendorf tube, mix 0.5 nmol of sense oligonucleotides with 0.5 nmol of antisense oligonucleotides in a 20 μl final reaction volume of 1× annealing buffer. Heat the sample for 5 min at 95 °C and allow the reaction mixture to cool down to 37 °C for 30 min. The annealed miR T oligonucleotide products can be visualized on agarose gel (see Note 7 ) before being stored at −20 °C.
-
2.
Digest 5 μg of RILES plasmids with 10 U of Not I and Mlu I restriction digest enzymes in a 50 μl reaction volume for 2 h at 37 °C. Gel purified the digested 6.2 kb RILES plasmid of 6.2 kb and measure absorbance at 260 nm using a spectrophotometer. A DNA band of 120 base pairs corresponding to the miR T sequence should also be detectable on the agarose gel.
-
3.
Ligate 50 ng of purified, digested RILES plasmids with the 2 μl of a 1/250 dilution of miR T oligonucleotides from step 1 in a 20 μl reaction volume containing 2.5 U of T4 DNA ligase. Incubate the ligation mixture for 30 min at 25 °C (see Note 8 ). Stop the reaction by inactivating the ligase at 65 °C for 10 min.
-
4.
Transform 50 μl of competent DH5a cells (2.5 × 106 pfu cells) with 5 μl of the ligation product from step 3 according to the manufacturer’s instructions (NEB, Hitchin UK). Incubate transformed bacteria on an LB agar ampicillin plate for 16 h at 37 °C.
-
5.
The following day, pick ten individual recombinant colonies and grow each in 3 ml of LB containing 50 μg/ml of ampicillin for 16 h at 37 °C with agitation using an orbital shaker. Extract and purify the pRILES plasmids using the standard alkaline lysis method (NucleoBond Xtra Mini kit, Macherey-Nagel).
-
6.
To identify the recombinant pRILES plasmids, containing the miR T sequence, from the parental, non-recombinant plasmid, digest 1 μg of plasmid from step 5 with 5 U of Not I and Mlu I restriction enzymes for 1 h at 37 °C in 20 μl reaction volume. Analyse the digestion product on a 1 % agarose electrophoresis gel. A DNA band of 120 base pairs corresponding to the miR T cassette should be visible among the digested RILES plasmid DNA of 6.2 kb.
-
7.
Amplify the corresponding positive colonies in 250 ml of LB containing ampicillin at 50 μg/ml for 16 h at 37 °C with agitation.
-
8.
Extract the pRILES using endotoxin NucleoBond Xtra Maxi kit (Macherey-Nagel) and measure absorbance at 260 and 280 nm to estimate the concentration of the pRILES and the purity of the preparation. The RILES plasmids can be stored at −20 °C for at least 1 year without loss of activity. An aliquot is used for sequencing the plasmid to ensure specificity of oligonucleotide miR T sequences cloned in the RILES.
3.2 In Vitro Functional Validation Study Using Synthetic miRNA Mimics
-
1.
HEK 293 cell are grown in complete medium at 37 °C, 5 % CO2, in a humidified atmosphere (tissue culture incubator). It is important that the cells are split at least twice at constant cell density to ensure optimal transfection efficiency.
-
2.
One day before transfection, plate 1.5 × 105 cells into wells of a 24-well plate and incubate overnight in the tissue culture incubator.
-
3.
The following day, ensure that the adherent cells have reached at least 80 % confluence. Renew the cell culture media in wells with 400 μl of serum-free media and return the plate to the tissue culture incubator.
-
4.
Prepare the pRILES/ICAFectin complexes, according to a transfection ratio of 1/2 (μg pDNA /μl ICAfectin) (see Note 9 ) as recommended by the manufacturer (In Cell Art). The following protocol is for the transfection of pRILES into 1 well of the 24-well plate. In a first Eppendorf tube (A) add 0.5 μg RILES plasmids plus the desired amount of miRNA precursor (see Note 10 ) and complete the total amount of nucleic acids to 2 μg using the pQE30 carrier plasmid (see Note 6 ). Adjust the final volume to 50 μl with OptiMEN medium. In a second Eppendorf tube (B) add 2 μl of ICAfectin 441 and adjust the final volume to 50 μl with OptiMEN medium. Equilibrate the tubes for 5 min at room temperature. Then add the content of tube B to tube A and homogenate the solution thoroughly by pipetting up and down at least five times. Incubate for 30 min at room temperature to allow the formation of the pRILES/ICAfectin 441 complexes.
-
5.
Add the 100 μl complexes from step 4 drop-by-drop into the well of the 24-well plate. Transfect the cells for 4 h in the tissue culture incubator. Thereafter, the transfection media is replaced with 500 μl of complete medium.
-
6.
Incubate the 24-well plate for an additional 48 h in the tissue culture incubator.
-
7.
Determine expression of the luciferase gene using the Luciferase Assay System with Reporter Lysis Buffer kit following the manufacturer’s recommendations (Promega). Briefly, the cell monolayer is washed once with PBS, and dried completely before adding 100 μl of cell culture lysis reagent (CCLR). The cell monolayer is then scraped carefully using a cell scraper or alternatively using a 200 μl TIPS incurved manually to produce a right angle. The resulting cell lysate is incubated on ice for 10 min and pre-cleared by a centrifugation step at 6000 × g for 5 min. An aliquot of 20 μl of cell supernatant is transferred to a luminometer tube and 100 μl of Luciferase Assay Reagent containing the luciferin substrate is added to the tube using an automatic luminometer. The light emitted from the catalyzed luciferin product is recorded for a 2-s integration time. Each sample is read in duplicate to ensure accurate and reproducible results.
-
8.
A 10 μl aliquot of pre-cleared lysate from step 7 is used to determine the protein content in sample using the standardized BCA method (Pierce™ BCA Protein Assay Kit).
-
9.
Relative luciferase activities (RLU) are normalized to protein content and expressed as RLU/mg of protein. The final data are represented as fold of luciferase induction relative to control cells, transfected with the RILES /miR T plasmid alone or transfected in the presence of irrelevant synthetic miRNAs, set to the arbitral value of 1. A typical example of results that can be generated is shown in Fig. 2.
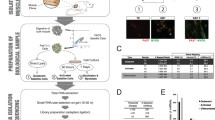
Fig. 2 Dose–response study of luciferase induction in HEK 293 cells transfected with pRILES/122 T. (a) The HEK 293 cells are transfected with pRILES/122 T in presence of increasing concentrations of synthetic miRNA-122 precursor. (b) Selective luciferase expression in HEK 293 cells transfected either with pRILES/122T in the presence of two concentrations of synthetic miRNA-122 precursor and as a control of specificity with the irrelevant synthetic miRNA-133 and miRNA-221. The luciferase expression in cells was determined 48 h post-transfection, and expressed as fold of luciferase induction relative to control cells transfected with the control pRILES plasmids, not regulated by miRNA and set to the arbitral value of 1. Data shown are the mean +/− SD of one representative experiment performed in triplicate and reproduced at least three times. Statistics using the two-tailed t-test, *p < 0.05; **p < 0.01, n.s. (no statistically significant difference) compared to control cells. Reprinted and adapted from [12]
3.3 In Vivo Bioluminescence Imaging of myomiRS Expression in the Tibialis Anterior Skeletal Muscles of Mice
-
1.
Adult, 8-week-old female Swiss mice (Harlam) are housed in a specific pathogen-free (SPF) animal facility in accordance with the European Ethical Conduct and Animal Recommendation guidelines (CARE, directive 2010/63/EU).
-
2.
A total of 15 mice are randomly separated into five groups as follows: RILES (control, non-regulated miRNA plasmid), RILES/1T (RILES plasmid containing the miRNA-1 targeting cassette), RILES/206T (RILES plasmid containing the miRNA-206 targeting cassette), RILES/133T (RILES plasmid containing the miRNA-133 targeting cassette), and the RILES/1T (RILES plasmid containing the miRNA-1 targeting cassette).
-
3.
Two days before the intramuscular injection of RILES plasmids, the mice are anesthetized with 2.5 % isoflurane/02 (oxygen) at a flow rate of 1 l/min. Then the fur on the tibialis anterior muscles of each mouse is shaved using either a razor or an epilator cream to guide latter the intramuscular injection.
-
4.
The day of the intramuscular injection, the pRILES/amphiphilic block copolymer 704 complexes are prepared before the mice are anesthetized. The general principle involves injecting 50 μl of solution containing 10 μg of pRILES formulated with 3 % (w/v) of amphiphilic block copolymer 704. Alternatively, the Lutrol block copolymer F-66 transfection reagent (Sigma Aldrich, St. Louis, USA) can also be used with same results. The following protocol is for eight intramuscular injections (1 group of mice plus 2 extra volumes). In a first Eppendorf tube (A) add 200 μl of 6 % (w/v) amphiphilic block copolymer 704 (see Note 11 ) prepared in tyrode solution at pH 7.6. In a second Eppendorf tube (B) add 80 μg of pRILES and complete the volume to 200 μl using the tyrode solution at pH 7.6. Equilibrate the tubes for 5 min at room temperature. Then add the content of tube B to tube A and homogenate the solution thoroughly by pipetting up and down at least five times. Incubate for 30 min at room temperature to allow the formation of pRILES/amphiphilic block copolymer 704 complexes. The complexes are stable for at least 2 h at room temperature.
-
5.
Before the intramuscular injection of the pRILES/amphiphilic block copolymer 704 complexes, the mice are first anesthetized as described in step 3. The intramuscular injection involves a rapid injection 50 μl of pRILES/amphiphilic block copolymer 704 complexes (see Note 12 ) at one site in the proximal region of the tendon tibialis (see Note 12 ). The injection is performed using a microfine insulin syringe (U100). Thereafter, the mice are returned to the cage and placed under surveillance until their behavior returns to normal.
-
6.
Six days (see Note 13 ) later the mice are injected intraperitoneally with the luciferin substrate (1.5 mg/mouse prepared in PBS). Five minutes after the injection (see Note 14 ), the mice are anesthetized with isoflurane as described in step 3 and placed in the bioluminescence scanner, connected to an isoflurane ventilation system to maintain constant anaesthesia. The first bioluminescence scan starts exactly 10 min after the injection of luciferin. The acquisition parameters are as follows: 2 min integration time, F/Stop, binin 8. At the end of the scan, a picture of the mouse is taken to enable bioluminescence images to be superposed. Final data are expressed as photons/second/pixel from reconstructed superposed images. A typical example of results generated is shown in Fig. 3.
Fig. 3 Noninvasive bioluminescence imaging of the muscle-specific myomiRs-206, -133, and -1 in mice. 8 μg pRILES/133T, pRILES/1T, and pRILES/206T were formulated with the amphiphilic block polymer 704 transfection reagent and intramuscularly injected in the left and right tibialis anterior to transfect the skeletal muscles of Swiss mice. Negative control included the pRILES, not regulated by miRNA and the pRILES/122T, regulated by the miRNA-122, a liver specific miRNA not expressed in the mouse tibialis skeletal muscle. Bioluminescence imaging was performed 6 days later and light emission quantified using regions of interest covering the lower legs of the mice. (a) Representative bioluminescence images collected in the lower left legs of mice. (b) Quantitative bioluminescence values detected in the mice described in (a) and expressed as luciferase induction relative to the control pRILES group of animals set arbitrarily to the value of 1. Error bars in (b), mean +/− SEM (n = 6) of one representative experiment repeated at least three times. Statistics using the two-tailed t-test, *p < 0.05; **p < 0.01 n.s. (no statistically significant difference) compared with the pRILES control group. Reprinted and adapted from [12]
4 Notes
-
1.
The nucleotide sequence of mature miRNAs can be found from the miRNA data base as miR base (http://www.mirbase.org/). It is important to specify the species of miRNA. MiRNA nucleotide sequences from the same annotated miRNA can vary slightly from species to species, for instance from human to mice.
-
2.
The number of the miR T block sequence can be adjusted from 2 to 8. Previous reports have demonstrated that a minimum of two block sequences are sufficient to repress expression of the target gene [16, 17]. In our study, we found that four complementary block sequences of a miRNA were optimal to switch on the RILES in cells.
-
3.
Sense and antisense oligonucleotides are chemically synthesized with a 5′-end phosphate group to allow the direct ligation of the annealed miR T oligonucleotide to the digested RILES plasmid.
-
4.
HEK 293 cells are easy to transfect enabling both efficient expression of the RILES plasmids and delivery of miRNA mimics in a large proportion of the same cells. We recommend determining the endogenous expression level of the studied miRNAs in these cells by quantitative RT-PCR. Elevated endogenous expression of miRNAs in these cells will in turn elevate the basal expression level of luciferase gene in the cells transfected with the pRILES/miR T plasmid alone. This could compromise the dose response study carried out with increasing doses of synthetic miRNA precursors.
-
5.
There are two types of commercially available synthetic miRNA agonists developed by Ambion and distributed by Life technology. The first generation consists in the miR precursor (Pre-mir™ miRNA Precursor) and the second (latest generation) consists in the miR mimics (mirVana® miRNA mimic). Recently (unpublished data), we found that the miR mimics are up to ten times more potent in switching on the RILES than the miR precursors at the same final concentration.
-
6.
We usually normalize our transfection condition to a final amount of 2 μg of nucleic acids (RILES plus synthetic miRNA precursors) to ensure that all formulated nucleic acids are the same from well to well. The amount and nature of nucleic acids formulated with the transfection reagent impact directly on the size and charge of nanoparticles formed after complexation and consequently on the transfection efficiency. This normalization procedure could also be carried out with the widely used “empty plasmid” or “mock plasmid”.
-
7.
One aliquot of the annealed double-stranded miR T oligonucleotides can be analyzed on 3 % TAE/agarose gel to differentiate the double-stranded oligonucleotides from the single-stranded, non-annealed oligonucleotides. Upon electrophoretic migration, the double-stranded oligonucleotides migrate the same distance to the 200 bp DNA band of a 1 kb ladder, while the single-stranded oligonucleotides migrate the same distance to the 100 bp DNA of the 1 kb ladder.
-
8.
A negative control of this experiment includes the commonly used “self-ligation product” which is obtained by carrying out the same procedure, but without adding the miR T oligonucleotides in the reaction volume. This step is not strictly necessary but allows the efficiency of the ligation reaction to be predicted, enabling an estimation of the total number of bacterial colonies which need to be screened to identify colonies containing the recombinant pRILES plasmid.
-
9.
Other transfection reagent such as Lipofectamine 2000 (Life Technologies SAS, St. Aubin, France) could also be used.
-
10.
We recommend that each experiment includes the following samples in triplicate: (1) HEK 293 cells transfected with pRILES/122T alone, (2) HEK 293 cells transfected with pRILES/122T plus 4 μl (20 μmol) of stock miRNA 122 precursor solution to generate a final concentration of miRNA-122 precursor of 40 nM during the transfection procedure (see step 5 from Subheading 3), (3) HEK 293 cells transfected with pRILES/122T plus 2 μl (10 μmol) of stock miRNA 122 precursor solution to generate a final concentration of miRNA-122 precursor of 20 nM during the transfection procedure, (4) HEK 293 cells transfected with pRILES/122T plus 1 μl (5 μmol) of stock miRNA 122 precursor solution to generate a final concentration of miRNA-122 precursor of 10 nM during the transfection procedure, (5) HEK 293 cells transfected with pRILES/122T plus 0.5 μl (0.25 μmol) of stock miRNA 122 precursor solution to generate a final concentration of miRNA-122 precursor of 0.5 nM during the transfection procedure, (6) HEK 293 cells transfected with pRILES/122T plus 0.25 μl (0.125 μmol) of stock miRNA 122 mimic solution to a final concentration of miRNA-122 mimics of 0.25 nM during the transfection procedure, and (7) HEK 293 cells transfected with pRILES/122T plus 0.125 μl (0.625 μmol) of stock miRNA 122 mimic solution to generate a final concentration of miRNA-122 mimics of 0.125 nM during the transfection procedure. As a control, the HEK 293 cells are transfected with pRILES/122T plus 1 or 10 nM final concentration of irrelevant miRNA such as miRNA-133 precursor or miRNA-206 precursor (see Fig. 2).
-
11.
We recommend preparing the amphiphilic block copolymer 704 solution the day before the intramuscular injection and leaving the solution at 4 °C overnight. We noticed that the transfection efficacy of the tibialis anterior skeletal muscle is optimal under this condition.
-
12.
A rapid intramuscular injection (approximately 3 s) of the formulated RILES plasmids in the tibialis anterior muscle increases the efficacy of gene transfer in the skeletal muscle tissue. This procedure aims to mimic the principle of hydrodynamic injection of pDNA as described in detail in [18].
-
13.
Optimal gene expression of pRILES is detected 6 days after the intramuscular injection although a minimum of 2 days is sufficient to detect significant bioluminescent signals in the tibialis skeletal muscle of the mice.
-
14.
It is recommended to administrate the luciferin substrate to non-anesthetized mice to allow optimal diffusion of the substrate in the whole-body animal tissue. Moreover, to ensure equal and homogenate diffusion of the luciferin in all the group of mice, the scanning of the mice should start exactly 10 min after the intraperitonial injection of the luciferin substrate.
References
Winter J, Jung S, Keller S, Gregory RI, Diederichs S (2009) Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 11:228–234
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Berezikov E (2011) Evolution of microRNA diversity and regulation in animals. Nat Rev Genet 12:846–860
Kole R, Krainer AR, Altman S (2012) RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat Rev Drug Discov 11:125–140
Kloosterman WP, Plasterk RH (2006) The diverse functions of microRNAs in animal development and disease. Dev Cell 11:441–450
Sayed D, Abdellatif M (2011) MicroRNAs in development and disease. Physiol Rev 91:827–887
Dong H, Lei J, Ding L, Wen Y, Ju H, Zhang X (2013) MicroRNA: function, detection, and bioanalysis. Chem Rev 113:6207–6233
Ko HY, Lee YS, Kim S (2014) Bioluminescence reporter gene-based detection of microRNAs. Methods Mol Biol 1098:85–95
Ko MH, Kim S, Hwang do W, Ko HY, Kim YH, Lee DS (2008) Bioimaging of the unbalanced expression of microRNA9 and microRNA9* during the neuronal differentiation of P19 cells. FEBS J 275:2605–2616
Kim HJ, Kim YH, Lee DS, Chung JK, Kim S (2008) In vivo imaging of functional targeting of miR-221 in papillary thyroid carcinoma. J Nucl Med 49:1686–1693
Maurel M, Jalvy S, Ladeiro Y, Combe C, Vachet L, Sagliocco F, Bioulac-Sage P, Pitard V, Jacquemin-Sablon H, Zucman-Rossi J, Laloo B, Grosset CF (2013) A functional screening identifies five microRNAs controlling glypican-3: role of miR-1271 down-regulation in hepatocellular carcinoma. Hepatology 57:195–204
Ezzine S, Vassaux G, Pitard B, Barteau B, Malinge JM, Midoux P, Pichon C, Baril P (2013) RILES, a novel method for temporal analysis of the in vivo regulation of miRNA expression. Nucleic Acids Res 41, e192
Stieger K, Belbellaa B, Le Guiner C, Moullier P, Rolling F (2009) In vivo gene regulation using tetracycline-regulatable systems. Adv Drug Deliv Rev 61:527–541
Mullick A, Xu Y, Warren R, Koutroumanis M, Guilbault C, Broussau S, Malenfant F, Bourget L, Lamoureux L, Lo R, Caron AW, Pilotte A, Massie B (2006) The cumate gene-switch: a system for regulated expression in mammalian cells. BMC Biotechnol 6:43
Chevre R, Le Bihan O, Beilvert F, Chatin B, Barteau B, Mevel M, Lambert O, Pitard B (2011) Amphiphilic block copolymers enhance the cellular uptake of DNA molecules through a facilitated plasma membrane transport. Nucleic Acids Res 39:1610–1622
Brown BD, Venneri MA, Zingale A, Sergi Sergi L, Naldini L (2006) Endogenous microRNA regulation suppresses transgene expression in hematopoietic lineages and enables stable gene transfer. Nat Med 12:585–591
Brown BD, Gentner B, Cantore A, Colleoni S, Amendola M, Zingale A, Baccarini A, Lazzari G, Galli C, Naldini L (2007) Endogenous microRNA can be broadly exploited to regulate transgene expression according to tissue, lineage and differentiation state. Nat Biotechnol 25:1457–1467
Suda T, Liu D (2007) Hydrodynamic gene delivery: its principles and applications. Mol Ther 15:2063–2069
Acknowledgments
This work was supported by La ligue Contre le Cancer du Loiret, Région Centre. We would like thank the staff of the CIPA animal facility (TAAM/CIPA, CNRS UPS 44, Orléans) for their technical assistance and access to the Lumina Bioluminescence scanner. In addition, we would like to thank Dr. Lily Mijouin for helping in the preparation of the manuscript for this chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Baril, P., Pichon, C. (2016). Positive Bioluminescence Imaging of MicroRNA Expression in Small Animal Models Using an Engineered Genetic-Switch Expression System, RILES. In: Medarova, Z. (eds) RNA Imaging. Methods in Molecular Biology, vol 1372. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3148-4_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3148-4_15
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3147-7
Online ISBN: 978-1-4939-3148-4
eBook Packages: Springer Protocols