Abstract
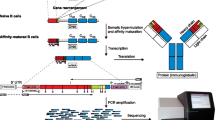
The B cell receptor (BCR) repertoire is highly diverse. Repertoire diversity is achieved centrally by somatic recombination of immunoglobulin (Ig) genes and peripherally by somatic hypermutation and Ig heavy chain class-switching. Throughout these processes, there is selection for functional gene rearrangements, selection against gene combinations resulting in self-reactive BCRs, and selection for BCRs with high affinity for exogenous antigens after challenge. Hence, investigation of BCR repertoires from different groups of B cells can provide information on stages of B cell development and shed light on the etiology of B cell pathologies. In most instances, the third complementarity determining region of the Ig heavy chain (CDR-H3) contributes the majority of amino acids to the antibody/antigen binding interface. Although CDR-H3 spectratype analysis provides information on the overall diversity of BCR repertoires, this fairly simple technique analyzes the relative quantities of CDR-H3 regions of each size, within a range of approximately 10–80 bp, without sequence detail and thus is limited in scope. High-throughput sequencing (HTS) techniques on the Roche 454 GS FLX Titanium system, however, can generate a wide coverage of Ig sequences to provide more qualitative data such as V, D, and J usage as well as detailed CDR3 sequence information. Here we present protocols in detail for CDR-H3 spectratype analysis and HTS of human BCR repertoires.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sanz I (1991) Multiple mechanisms participate in the generation of diversity of human H chain CDR3 regions. J Immunol 147:1720–1729
Tonegawa S (1983) Somatic generation of antibody diversity. Nature 302:575–581
Raaphorst FM, Raman CS, Tami J, Fischbach M, Sanz I (1997) Human Ig heavy chain CDR3 regions in adult bone marrow pre-B cells display an adult phenotype of diversity: evidence for structural selection of DH amino acid sequences. Int Immunol 9:1503–1515
Volpe JM, Kepler TB (2008) Large-scale analysis of human heavy chain V(D)J recombination patterns. Immunome Res 4:3
Larimore K, McCormick MW, Robins HS, Greenberg PD (2012) Shaping of human germline IgH repertoires revealed by deep sequencing. J Immunol 189:3221–3230
Ademokun A, Wu YC, Martin V, Mitra R, Sack U, Baxendale H, Kipling D, Dunn-Walters DK (2011) Vaccination-induced changes in human B-cell repertoire and pneumococcal IgM and IgA antibody at different ages. Aging Cell 10:922–930
Wu YC, Kipling D, Dunn-Walters DK (2012) Age-related changes in human peripheral blood IGH repertoire following vaccination. Front Immunol 3:193
Cuisinier AM, Guigou V, Boubli L, Fougereau M, Tonnelle C (1989) Preferential expression of VH5 and VH6 immunoglobulin genes in early human B-cell ontogeny. Scand J Immunol 30:493–497
Gibson KL, Wu YC, Barnett Y, Duggan O, Vaughan R, Kondeatis E, Nilsson BO, Wikby A, Kipling D, Dunn-Walters DK (2009) B-cell diversity decreases in old age and is correlated with poor health status. Aging Cell 8:18–25
Tschumper RC, Asmann YW, Hossain A, Huddleston PM, Wu X, Dispenzieri A, Eckloff BW, Jelinek DF (2012) Comprehensive assessment of potential multiple myeloma immunoglobulin heavy chain V-D-J intraclonal variation using massively parallel pyrosequencing. Oncotarget 3:502–513
Warsame AA, Aasheim HC, Nustad K, Troen G, Tierens A, Wang V, Randen U, Dong HP, Heim S, Brech A, Delabie J (2011) Splenic marginal zone lymphoma with VH1-02 gene rearrangement expresses poly- and self-reactive antibodies with similar reactivity. Blood 118:3331–3339
Luo C, Tsementzi D, Kyrpides N, Read T, Konstantinidis KT (2012) Direct comparisons of Illumina vs. Roche 454 sequencing technologies on the same microbial community DNA sample. PLoS One 7:e30087
Meyer M, Stenzel U, Myles S, Prufer K, Hofreiter M (2007) Targeted high-throughput sequencing of tagged nucleic acid samples. Nucleic Acids Res 35, e97
Wu YC, Kipling D, Dunn-Walters DK (2011) The relationship between CD27 negative and positive B cell populations in human peripheral blood. Front Immunol 2:81
Wu YC, Kipling D, Leong HS, Martin V, Ademokun AA, Dunn-Walters DK (2010) High-throughput immunoglobulin repertoire analysis distinguishes between human IgM memory and switched memory B-cell populations. Blood 116:1070–1078
Alamyar E, Duroux P, Lefranc MP, Giudicelli V (2012) IMGT® tools for the nucleotide analysis of immunoglobulin (IG) and T cell receptor (TR) V-(D)-J repertoires, polymorphisms, and IG mutations: IMGT/V-QUEST and IMGT/HighV-QUEST for NGS. Methods Mol Biol 882:569–604
Alamyar E, Giudicelli V, Li S, Duroux P, Lefranc MP (2012) IMGT/HIGHV-QUEST: the IMGT® web portal for immunoglobulin (Ig) or antibody and T cell receptor (TR) analysis from NGS high throughput and deep sequencing. Immunome Res 8:26
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein Identification and Analysis Tools on the ExPASy Server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, Clifton, UK, pp 571–607
R Development Core Team, R Foundation for Statistical Computing, Vienna, Austria (2010) ISBN 3-900051-07-0
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Wu, YC., Kipling, D., Dunn-Walters, D. (2015). Assessment of B Cell Repertoire in Humans. In: Shaw, A. (eds) Immunosenescence. Methods in Molecular Biology, vol 1343. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2963-4_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2963-4_16
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2962-7
Online ISBN: 978-1-4939-2963-4
eBook Packages: Springer Protocols