Abstract
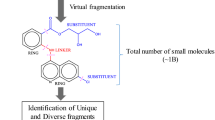
Fragment based screening (FBS) has emerged as a mainstream lead discovery strategy in academia, biotechnology start-ups, and large pharma. As a prerequisite of FBS, a structurally diverse library of fragments is desirable in order to identify chemical matter that will interact with the range of diverse target classes that are prosecuted in contemporary screening campaigns. In addition, it is also desirable to offer synthetically amenable starting points to increase the probability of a successful fragment evolution through medicinal chemistry. Herein we describe a method to identify biologically relevant chemical substructures that are missing from an existing fragment library (chemical gaps), and organize these chemical gaps hierarchically so that medicinal chemists can efficiently navigate the prioritized chemical space and subsequently select purchasable fragments for inclusion in an enhanced fragment library.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dalvit C (2009) NMR methods in fragment screening: theory and a comparison with other biophysical techniques. Drug Discov Today 14:1051–1057
Blaney J, Nienaber V, Burley SK (2006) Fragment-based Lead Discovery and Optimization Using X-Ray Crystallography, Computational Chemistry, and High-throughput Organic Synthesis. In: Jahnke W, Erlanson DA (eds) Fragment-based approaches in drug discovery. Wiley, Weinheim, pp 215–248
Elinder M, Geitmann M, Gossas T et al (2011) Experimental validation of a fragment library for lead discovery using SPR biosensor technology. J Biomol Screen 16:15–25
Lau WF, Withka JM, Hepworth D et al (2011) Design of a multi-purpose fragment screening library using molecular complexity and orthogonal diversity metrics. J Comput Aided Mol Des 25:621–636
Mok NY, Brenk R, Brown N (2014) Increasing the coverage of medicinal chemistry-relevant space in commercial fragments screening. J Chem Inf Model 54:79–85
Kutchukian PS, Vasilyeva NY, Xu J et al (2012) Inside the mind of a medicinal chemist: the role of human bias in compound prioritization during drug discovery. PLoS One 7:e48476
TIBCO Software Inc (2014) TIBCO Spotfire, 5.5.1
Biovia (2013) Pipeline Pilot, 9.1
Overington JP (2009) ChEMBL: large-scale mapping of medicinal chemistry and pharmacology data to genomes. Abstr Pap Am Chem S 238
Bento AP, Gaulton A, Hersey A et al (2014) The ChEMBL bioactivity database: an update. Nucleic Acids Res 42:D1083–D1090
Metabase. http://thomsonreuters.com/metabase/
Annis DA, Nickbarg E, Yang X et al (2007) Affinity selection-mass spectrometry screening techniques for small molecule drug discovery. Curr Opin Chem Biol 11:518–526
Congreve M, Carr R, Murray C et al (2003) A rule of three for fragment-based lead discovery? Drug Discov Today 8:876–877
Baell JB, Holloway GA (2010) New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem 53:2719–2740
Crisman TJ, Bender A, Milik M et al (2008) "Virtual fragment linking": an approach to identify potent binders from low affinity fragment hits. J Med Chem 51:2481–2491
Wassermann AM, Kutchukian PS, Lounkine E et al (2013) Efficient search of chemical space: navigating from fragments to structurally diverse chemotypes. J Med Chem 56:8879–8891
Lewell XQ, Judd DB, Watson SP et al (1998) RECAP – retrosynthetic combinatorial analysis procedure: a powerful new technique for identifying privileged molecular fragments with useful applications in combinatorial chemistry. J Chem Inf Comp Sci 38:511–522
Lounkine E, Kutchukian P, Glick M (2013) Chemometric applications of naïve Bayesian models in drug discovery – beyond compound ranking. In: Bajorath J (ed) Chemoinformatics: case studies and pharmaceutical applications. Wiley, Hoboken, NJ
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Kutchukian, P.S., So, SS., Fischer, C., Waller, C.L. (2015). Fragment Library Design: Using Cheminformatics and Expert Chemists to Fill Gaps in Existing Fragment Libraries. In: Klon, A. (eds) Fragment-Based Methods in Drug Discovery. Methods in Molecular Biology, vol 1289. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2486-8_5
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2486-8_5
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2485-1
Online ISBN: 978-1-4939-2486-8
eBook Packages: Springer Protocols