Abstract
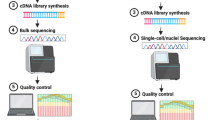
The next-generation sequencing (NGS) of RNA, or RNA-Seq, has significantly changed the way that the transcriptional content of a biological sample is investigated. RNA-Seq is a major advance for the field due to its largely unbiased and digital nature, its ability to empower RNA splice form construction at a genomic level, and its improved dynamic range when compared to a microarray technology. Investigating the healthy or diseased brain presents unique problems from the standpoint of transcriptional analysis as each cell type, and perhaps even each individual cell, is in a unique state of transcription. The organ is a complex mixture of main cell types (neuronal, glial, vascular, etc.), and within each of those types, there are a multitude of subtypes (specific neuronal populations, different classes of glial cells, etc.) that could each be targeted for investigation and could each respond to health and disease in distinct and functionally important ways. Here, we discuss the approach of using laser capture microdissection (LCM) to specifically select cell types of interest for transcriptional dissection. We highlight approaches to couple this with RNA-Seq to generate highly specific transcriptional profiles from the brain. Sample inputs into RNA-Seq protocols continue to be pushed lower, including several reports of single-cell transcriptome profiles; therefore, the combination of cell selection approaches, like LCM, with RNA-Seq is well poised to provide a researcher with the cell-specific digital whole transcriptome information that has been desired since transcriptomic profiling became feasible during the earliest days of the microarray.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Beach TG, Sue LI, Walker DG, et al. The Sun Health Research Institute Brain Donation Program: description and experience, 1987–2007. Cell Tissue Bank. 2008;9:229–45.
Chow N, Cox C, Callahan LM, et al. Expression profiles of multiple genes in single neurons of Alzheimer’s disease. Proc Natl Acad Sci U S A. 1998;95:9620–5.
Colangelo V, Schurr J, Ball MJ, et al. Gene expression profiling of 12633 genes in Alzheimer hippocampal CA1: transcription and neurotrophic factor down-regulation and up-regulation of apoptotic and pro-inflammatory signaling. J Neurosci Res. 2002;70:462–73.
Derisi J, Penland L, Brown PO, et al. Use of a cDNA microarray to analyse gene expression patterns in human cancer. Nat Genet. 1996;14:457–60.
Dickey CA, Loring JF, Montgomery J, et al. Selectively reduced expression of synaptic plasticity-related genes in amyloid precursor protein + presenilin-1 transgenic mice. J Neurosci. 2003;23:5219–26.
Dunckley T, Beach TG, Ramsey KE, et al. Gene expression correlates of neurofibrillary tangles in Alzheimer's disease. Neurobiol Aging. 2006;27:1359–71.
Eberwine J, Yeh H, Miyashiro K, et al. Analysis of gene expression in single live neurons. Proc Natl Acad Sci U S A. 1992;89:3010–4.
Emmert-Buck MR, Bonner RF, Smith PD, et al. Laser capture microdissection. Science. 1996;274:998–1001.
Ginsberg SD, Hemby SE, Lee VM, et al. Expression profile of transcripts in Alzheimer's disease tangle-bearing CA1 neurons. Ann Neurol. 2000;48:77–87.
Islam S, Zeisel A, Joost S, et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat Methods. 2014;11:163–6.
Lao KQ, Tang F, Barbacioru C, et al. mRNA-sequencing whole transcriptome analysis of a single cell on the SOLiD system. J Biomol Tech. 2009;20:266–71.
Liang WS, Dunckley T, Beach TG, et al. Gene expression profiles in anatomically and functionally distinct regions of the normal aged human brain. Physiol Genomics. 2007;28:311–22.
Liang WS, Dunckley T, Beach TG, et al. Altered neuronal gene expression in brain regions differentially affected by Alzheimer's disease: a reference data set. Physiol Genomics. 2008;33:240–56.
Liang WS, Dunckley T, Beach TG, et al. Neuronal gene expression in non-demented individuals with intermediate Alzheimer's disease neuropathology. Neurobiol Aging. 2010;31:549–66.
Lovatt D, Ruble BK, Lee J, et al. Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue. Nat Methods. 2014;11:190–6.
Nolan T, Hands RE, Bustin SA. Quantification of mRNA using real-time RT-PCR. Nat Protoc. 2006;1:1559–82.
Qiu S, Luo S, Evgrafov O, et al. Single-neuron RNA-Seq: technical feasibility and reproducibility. Front Genet. 2012;3:124.
Schena M, Shalon D, Davis RW, et al. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995;270:467–70.
Schena M, Shalon D, Heller R, et al. Parallel human genome analysis: microarray-based expression monitoring of 1000 genes. Proc Natl Acad Sci U S A. 1996;93:10614–9.
Streets AM, Zhang X, Cao C, et al. Microfluidic single-cell whole-transcriptome sequencing. Proc Natl Acad Sci U S A. 2014;111:7048–53.
Tang F, Barbacioru C, Nordman E, et al. RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nat Protoc. 2010;5:516–35.
Acknowledgments
The authors gratefully acknowledge the support of the McKnight Brain Research Foundation, State of Arizona, and NIA-NIH Grant #AG049465.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Siniard, A.L., Corneveaux, J.J., De Both, M., Chawla, M.K., Barnes, C.A., Huentelman, M.J. (2015). RNA Sequencing from Laser Capture Microdissected Brain Tissue to Study Normal Aging and Alzheimer’s Disease. In: Jain, K. (eds) Applied Neurogenomics. Neuromethods, vol 97. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2247-5_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2247-5_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2246-8
Online ISBN: 978-1-4939-2247-5
eBook Packages: Springer Protocols