Abstract
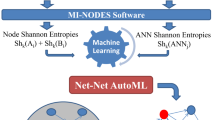
Microbial communities are found in nearly all environments and play a critical role in defining ecosystem service. Understanding the relationship between these microbial communities and their environment is essential for prediction of community structure, robustness, and response to ecosystem changes. Microbial Assemblage Prediction (MAP) describes microbial community structure as an artificial neural network (ANN) that models the microbial community as functions of environmental parameters and community intra-microbial interactions. MAP models can be used to predict community assemblages over a wide range of possible environmental parameters, extrapolate the results of point observations across spatial scales, and make predictions about how microbial communities may fluctuate as the result of changes in their environment.
The submitted manuscript has been created by UChicago Argonne, LLC, operator of Argonne National Laboratory (“Argonne”). Argonne, a US Department of Energy Office of Science laboratory, is operated under contract no. DE-AC02-06CH11357. The US Government retains for itself, and others acting on its behalf, a paid-up nonexclusive, irrevocable worldwide license in the said article to reproduce and prepare derivative works, distribute copies to the public, and perform publicly and display publicly, by or on behalf of the government.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hugenholtz P, Goebel BM, Pace NR (1998) Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J Bacteriol 180:4765–4774
Barns SM, Takala SL, Kuske CR (1999) Wide distribution and diversity of members of the bacterial kingdom Acidobacterium in the environment. Appl Environ Microbiol 65:1731–1737
Shtarkman YM et al (2013) Subglacial Lake Vostok (Antarctica) accretion ice contains a diverse set of sequences from aquatic, marine and sediment-inhabiting bacteria and eukarya. PLoS One 8(7):e67221
Fierer N et al (2008) Short-term temporal variability in airborne bacterial and fungal populations. (Translated from eng). Appl Environ Microbiol 74(1):200–207
Bowers RM et al (2009) Characterization of airborne microbial communities at a high-elevation site and their potential to act as atmospheric ice nuclei. (Translated from eng). Appl Environ Microbiol 75(15):5121–5130
Takai K et al (2001) Archaeal diversity in waters from deep South African gold mines. Appl Environ Microbiol 67(12):5750–5760
Edwards RA et al (2006) Using pyrosequencing to shed light on deep mine microbial ecology. BMC Genomics 7:57
Teske A, Sorensen KB (2008) Uncultured archaea in deep marine subsurface sediments: have we caught them all? ISME J 2(1):3–18
Baune C, Bottcher ME (2010) Experimental investigation of sulphur isotope partitioning during outgassing of hydrogen sulphide from diluted aqueous solutions and seawater. Isotopes Environ Health Stud 46(4):444–453
Bardgett RD, Freeman C, Ostle NJ (2008) Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2(8):805–814
Thomas T, Gilbert J, Meyer F (2012) Metagenomics - a guide from sampling to data analysis. Microb Inform Exp 2(1):3
Larsen PE, Gibbons SM, Gilbert JA (2012) Modeling microbial community structure and functional diversity across time and space. FEMS Microbiol Lett 332(2):91–98
Larsen P, Hamada Y, Gilbert J (2012) Modeling microbial communities: current, developing, and future technologies for predicting microbial community interaction. J Biotechnol 160(1–2):17–24
Larsen PE, Field D, Gilbert JA (2012) Predicting bacterial community assemblages using an artificial neural network approach. Nat Methods 9(6):621–625
Caporaso JG et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Meyer F et al (2008) The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform 9:386
Smith VA et al (2006) Computational inference of neural information flow networks. PLoS Comput Biol 2(11):e161
Sogin ML et al (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci U S A 103(32):12115–12120
Mumby PJ, Clarke KR, Harborne AR (1996) Weighting species abundance estimates for marine resource assessment. Aquat Conserv 6(3):115–120
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Larsen, P., Dai, Y., Collart, F.R. (2015). Predicting Bacterial Community Assemblages Using an Artificial Neural Network Approach. In: Cartwright, H. (eds) Artificial Neural Networks. Methods in Molecular Biology, vol 1260. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-2239-0_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2239-0_3
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-2238-3
Online ISBN: 978-1-4939-2239-0
eBook Packages: Springer Protocols