Abstract
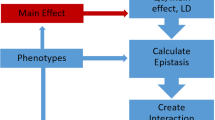
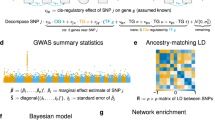
One of the challenges of understanding the genetic basis of complex phenotypes is explaining variability not attributable to individual genes. While most existing methods that investigate variant mutations or differential gene expression focus on individual effects, a complex system of gene interactions (epistasis) and pathways is likely needed to explain phenotypic variation. Herein, we examine methods for treating the interactions in these biological data sets as edges in a network model of the phenotype and review relevant network theory methods for analyzing network structure and identifying important genes. In particular, we review methods for detecting community structure, describing the statistical properties of networks, and computing network centrality of genes that may reveal insights missed by individual genetic effects. We also discuss available tools to facilitate the construction and visualization of epistasis networks of GWAS data.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lin A, Wang RT et al (2010) A genome-wide map of human genetic interactions inferred from radiation hybrid genotypes. Genome Res 20(8):1122–1132
Davis NA, Lareau CA et al (2013) Encore: Genetic Association Interaction Network centrality pipeline and application to SLE exome data. Genet Epidemiol 37(6):614–621
Hu T, Sinnott-Armstrong NA et al (2011) Characterizing genetic interactions in human disease association studies using statistical epistasis networks. BMC Bioinform 12:364
Davis NA, Crowe JE Jr et al (2010) Surfing a genetic association interaction network to identify modulators of antibody response to smallpox vaccine. Genes Immun 11(8):630–636
Pandey A, Davis NA et al (2012) Epistasis network centrality analysis yields pathway replication across two GWAS cohorts for bipolar disorder. Trans Psychiat 2:e154
Proulx SR, Promislow DE et al (2005) Network thinking in ecology and evolution. Trends Ecol Evol 20(6):345–353
Kaleta C, de Figueiredo LF et al (2011) In silico evidence for gluconeogenesis from fatty acids in humans. PLoS Comput Biol 7(7):e1002116
McKinney BA, Reif DM et al (2006) Machine learning for detecting gene-gene interactions: a review. Appl Bioinformatics 5(2):77–88
Ritchie MD, Hahn LW et al (2003) Power of multifactor dimensionality reduction for detecting gene-gene interactions in the presence of genotyping error, missing data, phenocopy, and genetic heterogeneity. Genet Epidemiol 24(2):150–157
Jiang X, Neapolitan RE et al (2011) Learning genetic epistasis using Bayesian network scoring criteria. BMC Bioinform 12:89
McKinney BA, Pajewski NM (2011) Six degrees of epistasis: statistical network models for GWAS. Front Genet 2:109
Clune J, Mouret JB et al (2013) The evolutionary origins of modularity. Proc Roy Soc B 280(1755):20122863
Newman MEJ (2010) Networks: an introduction. Oxford University Press, Oxford
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Gene Mol Biol 4: Article17
Purcell S, Neale B et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575
Schulze TG, Detera-Wadleigh SD et al (2009) Two variants in Ankyrin 3 (ANK3) are independent genetic risk factors for bipolar disorder. Mol Psychiatry 14(5):487–491
Wong AK, Park CY et al (2012) IMP: a multi-species functional genomics portal for integration, visualization and prediction of protein functions and networks. Nucleic Acids Res 40(Web Server issue):W484–W490
Mostafavi S, Ray D et al (2008) GeneMANIA: a real-time multiple association network integration algorithm for predicting gene function. Genome Biol 9(Suppl 1):S4
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Lareau, C.A., McKinney, B.A. (2015). Network Theory for Data-Driven Epistasis Networks. In: Moore, J., Williams, S. (eds) Epistasis. Methods in Molecular Biology, vol 1253. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2155-3_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2155-3_15
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2154-6
Online ISBN: 978-1-4939-2155-3
eBook Packages: Springer Protocols