Abstract
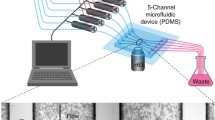
By implementing an external feedback loop one can tightly control the expression of a gene over many cell generations with quantitative accuracy. Controlling precisely the level of a protein of interest will be useful to probe quantitatively the dynamical properties of cellular processes and to drive complex, synthetically-engineered networks. In this chapter we describe a platform for real-time closed-loop control of gene expression in yeast that integrates microscopy for monitoring gene expression at the cell level, microfluidics to manipulate the cells environment, and original software for automated imaging, quantification, and model predictive control. By using an endogenous osmo-stress responsive promoter and playing with the osmolarity of the cells environment, we demonstrate that long-term control can indeed be achieved for both time-constant and time-varying target profiles, at the population level, and even at the single-cell level.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others

References
Bhalla US, Ram PT, Iyengar R (2002) MAP kinase phosphatase as a locus of flexibility in a mitogen-activated protein kinase signaling network. Science 297:1018–23
Hooshangi S, Thiberge S, Weiss R (2005) Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc Natl Acad Sci U S A 102:3581–6
Cai L, Dalal CK, Elowitz MB (2008) Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455:485–90
Celani A, Vergassola M (2010) Bacterial strategies for chemotaxis response. Proc Natl Acad Sci U S A 107:1391–6
Baumgartner BL, Bennett MR, Ferry M et al (2011) Antagonistic gene transcripts regulate adaptation to new growth environments. Proc Natl Acad Sci U S A 108:21087–92
O’Shaughnessy EC, Palani S, Collins JJ et al (2011) Tunable signal processing in synthetic MAP kinase cascades. Cell 144:119–31
de Nadal E, Alepuz PM, Posas F (2002) Dealing with osmostress through MAP kinase activation. EMBO Rep 3:735–40
Hohmann S (2002) Osmotic stress signaling and osmoadaptation in yeasts. Microbiol Mol Biol Rev 66:300–372
Miermont A, Uhlendorf J, McClean M et al (2011) The dynamical systems properties of the HOG signaling cascade. J Signal Transduct 2011:930940
Muzzey D, Gómez-Uribe C, Mettetal JT et al (2009) A systems-level analysis of perfect adaptation in yeast osmoregulation. Cell 138:160–71
Yi TM, Huang Y, Simon MI et al (2000) Robust perfect adaptation in bacterial chemotaxis through integral feedback control. Proc Natl Acad Sci U S A 97:4649–53
Van Voorst F, Neves L, Oliveira R et al (2005) A member of the sugar transporter family, Stl1p is the glycerol/H + symporter in Saccharomyces cerevisiae. Mol Biol Cell 16:2068–2076
O’Rourke SM, Herskowitz I (2004) Unique and redundant roles for HOG MAPK pathway components as revealed by whole-genome expression analysis. Mol Biol Cell 15:532–542
Uhlendorf J, Miermont A, Delaveau T et al (2012) Long-term model predictive control of gene expression at the population and single-cell levels. Proc Natl Acad Sci U S A 35:14271–14276
Klipp E, Nordlander B, Krüger R et al (2005) Integrative model of the response of yeast to osmotic shock. Nat Biotechnol 23:975–82
Hao N, Behar M, Parnell SC et al (2007) A systems-biology analysis of feedback inhibition in the Sho1 osmotic-stress-response pathway. Curr Biol 17:659–67
Mettetal JT, Muzzey D, Gómez-Uribe C et al (2008) The frequency dependence of osmo-adaptation in Saccharomyces cerevisiae. Science 319:482–4
Zi Z, Liebermeister W, Klipp E (2010) A quantitative study of the Hog1 MAPK response to fluctuating osmotic stress in Saccharomyces cerevisiae. PLoS One 5:e9522
Zechner C, Ruess J, Krenn P et al (2012) Moment-based inference predicts bimodality in transient gene expression. Proc Natl Acad Sci U S A 109:8340–8345
Uhlendorf J, Bottani S, Fages F, et al (2011) Towards real-time control of gene expression: controlling the hog signaling cascade. Pac Symp Biocomput 338–349
Menolascina F, di Bernardo M, di Bernardo D (2011) Analysis, design and implementation of a novel scheme for in-vivo control of synthetic gene regulatory networks. Automatica 47:1265–1270
Toettcher JE, Gong D, Lim WA et al (2011) Light-based feedback for controlling intracellular signaling dynamics. Nat Methods 8:837–839
Milias-Argeitis A, Summers S, Stewart-Ornstein J et al (2011) In silico feedback for in vivo regulation of a gene expression circuit. Nat Biotechnol 29:1114–1116
Chen S, Harrigan P, Heineike B et al (2013) Building robust functionality in synthetic circuits using engineered feedback regulation. Curr Opin Biotechnol 24:790–6
Acknowledgments
We acknowledge the support of the Agence Nationale de la Recherche (under the references DiSiP-ANR-07-JCJC-0001 and ICEBERG-ANR-10-BINF-06-01), of the Région Ile de France (C’Nano-ModEnv), of the Action d’Envergure ColAge from INRIA/INSERM (Institut Nationale de la Santé et de la Recherche Médicale), of the MechanoBiology Institute, and of the Laboratoire International Associé CAFS (Cell Adhesion France-Singapour).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Uhlendorf, J. et al. (2015). In Silico Control of Biomolecular Processes. In: Marchisio, M. (eds) Computational Methods in Synthetic Biology. Methods in Molecular Biology, vol 1244. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1878-2_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1878-2_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1877-5
Online ISBN: 978-1-4939-1878-2
eBook Packages: Springer Protocols