Abstract
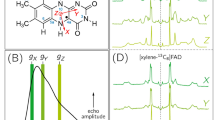
Flavin is a general name given to molecules having the heteroaromatic ring system of 7,8-dimethylisoalloxazine but practically means riboflavin (Rfl), flavin adenine dinucleotide (FAD), and flavin mononucleotide (FMN) in biological systems, whose structures are illustrated in Fig. 1, together with the atomic numbering scheme and ring numbering of the isoalloxazine moiety. As the isoalloxazine skeleton cannot be synthesized in human cells, it is obtained from diet as Rfl (vitamin B2). FAD and FMN can act as cofactors in flavoenzymes but Rfl does not. Most flavoenzymes catalyze redox reactions of substrates (Miura, Chem Rec 1:183–194, 2001). When O2 serves as the oxidant in the oxidation half cycle of an enzymic reaction, the enzyme is called “flavo-oxidase” but when others do, the enzyme is called “flavo-dehydrogenase.” The difference between the two types of oxidative catalysis arises from delicate differences in the π-electron distributions in the isoalloxazine ring, which can be revealed by Raman spectroscopy (Miura, Chem Rec 1:183–194, 2001). Since a flavin is an extremely versatile molecule, the scientific field including chemistry, biochemistry, and enzymology is collectively called “flavonology.” It was found recently, however, that the flavin also acts as a chromophore to initiate light-induced DNA repair and signal transductions (Sancar, Chem Rev 103:2203–2237, 2003).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Miura R (2001) Versatility and specificity in flavoenzymes: control mechanisms of flavin reactivity. Chem Rec 1:183–194
Sancar A (2003) Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem Rev 103:2203–2237
Li J, Uchida T, Todo T, Kitagawa T (2006) Similarities and differences between cyclobutane pyrimidine dimer photolyase and (6-4) photolyase as revealed by resonance Raman spectroscopy: electron transfer from the FAD cofactor to ultraviolet-damaged DNA. J Biol Chem 281:25551–25559
Li J, Uchida T, Ohta T, Todo T, Kitagawa T (2006) Characteristic structure and environment in FAD cofactor of (6-4) photolyase along function revealed by resonance Raman spectroscopy. J Phys Chem B 110: 16724–16732
Mataga N, Chosrowjan H, Shibata Y, Tanaka F (1998) Ultrafast fluorescence quenching dynamics of flavin chromophores in protein nanospace. J Phys Chem B 102:7081–7084
Li J, Liu Z, Tan C, Guo X, Wang L, Sancar A, Zhong D (2010) Dynamics and mechanism of repair of ultraviolet-induced (6-4) photoproduct by photolyase. Nature 466:887–890
Dutta PK, Nestor JR, Spiro TG (1977) Resonance coherent anti-stokes Raman-scattering spectra of fluorescent biological chromophores. Vibrational evidence for hydrogen-bonding of flavin to glucose oxidase and for rapid solvent exchange. Proc Natl Acad Sci U S A 74:4146–4149
Nishina Y, Kitagawa T, Shiga K, Horiike K, Matsumura Y, Watari H, Yamano T (1978) Resonance Raman spectra of riboflavin and its derivatives in the bound state with egg riboflavin binding proteins. J Biochem 84:925–932
Kitagawa T, Nishina Y, Kyogoku Y, Yamano T, Ohishi N, Takai-Suzuki A, Yagi K (1979) Resonance Raman spectra of carbon-13- and nitrogen-15-labeled riboflavin bound to egg-white flavoprotein. Biochemistry 18:1804–1808
Nishina Y, Shiga K, Horiike K, Tojo H, Kasai S, Yanase K, Matsui K, Watari H, Yamano T (1980) Vibrational-modes of flavin bound to riboflavin binding-protein from egg-white. Resonance Raman-spectra of lumiflavin and 8-substituted riboflavin. J Biochem 88:403–409
Abe M, Kyogoku Y (1987) Vibrational analysis of flavin derivatives: normal coordinate treatments of lumiflavin. Spectrochim Acta A Mol Spectrosc 43:1027–1037
Bowman WD, Spiro TG (1981) Normal mode analysis of lumiflavin and interpretation of resonance Raman-spectra of flavoproteins. Biochemistry 20:3313–3318
Eisenberg AS, Schelvis JP (2008) Contributions of the 8-methyl group to the vibrational normal modes of flavin mononucleotide and its 5-methyl semiquinone radical. J Phys Chem A 112:6179–6189
Unno M, Sano R, Masuda S, Ono TA, Yamauchi S (2005) Light-induced structural changes in the active site of the BLUF domain in AppA by Raman spectroscopy. J Phys Chem B 109:12620–12626
Zheng YG, Dong J, Palfey BA, Carey PR (1999) Using Raman spectroscopy to monitor the solvent-exposed and “buried” forms of flavin in p-hydroxybenzoate hydroxylase. Biochemistry 38:16727–16732
Lively CR, McFarland JT (1990) Assignment and the effect of hydrogen bonding on the vibrational normal modes of flavins and flavoproteins. J Phys Chem 94:3980–3994
Tegoni M, Gervais M, Desbois A (1997) Resonance Raman study on the oxidized and anionic semiquinone forms of flavocytochrome b 2 and L-lactate monooxygenase. Influence of the structure and environment of the isoalloxazine ring on the flavin function. Biochemistry 36:8932–8946
Yang K-Y, Swenson RP (2007) Modulation of the redox properties of the flavin cofactor through hydrogen-bonding interactions with the N(5) atom: role of αSer254 in the electron-transfer flavoprotein from the methylotrophic bacterium W3A1. Biochemistry 46:2289–2297
Yang K-Y, Swenson RP (2007) Nonresonance Raman study of the flavin cofactor and its interactions in the methylotrophic bacterium W3A1 electron-transfer flavoprotein. Biochemistry 46:2298–2305
Hazekawa I, Nishina Y, Sato K, Shichiri M, Miura R, Shiga K (1997) A Raman study on the C(4)=O stretching mode of flavins in flavoenzymes: hydrogen bonding at the C(4)=O moiety. J Biochem 121:1147–1154
Su Y, Tripathi GNR (1994) Time-resolved resonance Raman observation of protein-free riboflavin semiquinone radicals. J Am Chem Soc 116:4405–4407
Schelvis JPM, Pun D, Goyal N, Sokolova O (2006) Resonance Raman spectra of the neutral and anionic radical semiquinones of flavin adenine dinucleotide in glucose oxidase revisited. J Raman Spectrosc 37:822–829
Dutta PK, Spiro TG (1980) Resonance coherent anti-Stokes Raman scattering spectra of oxidized and semiquinone forms of Clostridium MP flavodoxin. Biochemistry 19:1590–1593
Nishina Y, Shiga K, Horiike K, Tojo H, Kasai S, Matsui K, Watari H, Yamano T (1980) Resonance Raman spectra of semiquinone forms of flavins bound to riboflavin binding protein. J Biochem 88:411–416
Kitagawa T, Sakamoto H, Sugiyama T, Yamano T (1982) Formation of the semiquinone form in the anaerobic reduction of adrenodoxin reductase by NADPH. Resonance Raman, EPR, and optical spectroscopic evidence. J Biol Chem 257:12075–12080
Schelvis JPM, Ramsey M, Sokolova O, Tavares C, Cecala C, Connell K, Wagner S, Gindt YM (2003) Resonance Raman and UV-Vis spectroscopic characterization of FADH• in the complex of photolyase with UV-damaged DNA. J Phys Chem B 107:12352–12362
Sugiyama T, Nisimoto Y, Mason HS, Loehr TM (1985) Flavins of NADPH-cytochrome-P-450 reductase: evidence for structural alteration of flavins in their one-electron-reduced semiquinone states from resonance Raman spectroscopy. Biochemistry 24:3012–3019
Park HW, Kim ST, Sancar A, Deisenhofer J (1995) Crystal structure of DNA photolyase from Escherichia coli. Science 268:1866–1872
Maul MJ, Barends TR, Glas AF, Cryle MJ, Domratcheva T, Schneider S, Schlichting I, Carell T (2008) Crystal structure and mechanism of a DNA (6-4) photolyase. Angew Chem Int Ed 47:10076–10080
Nishina Y, Tojo H, Shiga K (1988) Resonance Raman-spectra of anionic semiquinoid form of a flavoenzyme, D-amino-acid oxidase. J Biochem 104:227–231
Zheng YG, Carey PR, Palfey BA (2004) Raman spectrum of fully reduced flavin. J Raman Spectrosc 35:521–524
Zheng YJ, Ornstein RL (1996) A theoretical study of the structures of flavin in different oxidation and protonation states. J Am Chem Soc 118:9402–9408
Jorns MS, Baldwin ET, Sancar GB, Sancar A (1987) Action mechanism of Escherichia coli DNA photolyase. II. Role of the chromophores in catalysis. J Biol Chem 262:486–491
Lin C, Robertson DE, Ahmad M, Raibekas AA, Jorns MS, Dutton PL, Cashmore AR (1995) Association of flavin adenine dinucleotide with the Arabidopsis blue light receptor CRY1. Science 269:968–970
Fogle KJ, Parson KG, Dahm NA, Holmes TC (2011) Cryptochrome is a blue-light sensor that regulates neuronal firing rate. Science 331:1409–1413
Huala E, Oeller PW, Liscum E, Han IS, Larsen E, Briggs WR (1997) Arabidopsis NPH1: a protein kinase with a putative redox-sensing domain. Science 278:2120–2123
Iseki M, Matsunaga S, Murakami A, Ohno K, Shiga K, Yoshida K, Sugai M, Takahashi T, Hori T, Watanabe M (2002) A blue-light-activated adenylyl cyclase mediates photoavoidance in Euglena gracilis. Nature 415:1047–1051
Li J, Kitagawa T (2011) Applications of vibrational spectroscopy in the study of flavin-based photoactive proteins. Spectroscopy 25:261–269
Murphy AK, Tammaro M, Cortazar F, Gindt YM, Schelvis JPM (2008) Effect of the cyclobutane cytidine dimer on the properties of Escherichia coli DNA photolyase. J Phys Chem B 112:15217–15226
Mees A, Klar T, Gnau P, Hennecke U, Eker AP, Carell T, Essen L-O (2004) Crystal structure of a photolyase bound to a CPD-like DNA lesion after in situ repair. Science 306:1789–1793
Liu Z, Tan C, Guo X, Kao YT, Li J, Wang L, Sancar A, Zhong D (2011) Dynamics and mechanism of cyclobutane pyrimidine dimer repair by DNA photolyase. Proc Natl Acad Sci U S A 108:14831–14836
Benecky M, Li TY, Schmidt J, Frerman F, Watters KL, McFarland J (1979) Resonance Raman study of flavins and the flavoprotein fatty acyl coenzyme A dehydrogenase. Biochemistry 18:3471–3476
Sokolowsky K, Newton M, Lucero C, Wertheim B, Freedman J, Cortazar F, Czochor J, Schelvis JPM, Gindt YM (2010) Spectroscopic and thermodynamic comparisons of Escherichia coli DNA photolyase and Vibrio cholerae cryptochrome 1. J Phys Chem B 114:7121–7130
Salomon M, Christie JM, Knieb E, Lempert U, Briggs WR (2000) Photochemical and mutational analysis of the FMN-binding domains of the plant blue light receptor, phototropin. Biochemistry 39:9401–9410
Schopfer LM, Haushalter JP, Smith M, Milad M, Morris MD (1981) Resonance Raman spectra for flavin derivatives modified in the 8 position. Biochemistry 20:6734–6739
Alexandre MT, van Grondelle R, Hellingwerf KJ, Robert B, Kennis JT (2008) Perturbation of the ground-state electronic structure of FMN by the conserved cysteine in phototropin LOV2 domains. Phys Chem Chem Phys 10:6693–6702
Kikuchi S, Unno M, Zikihara K, Tokutomi S, Yamauchi S (2009) Vibrational assignment of the flavin-cysteinyl adduct in a signaling state of the LOV domain in FKF1. J Phys Chem B 113:2913–2921
Unno M, Masuda S, Ono TA, Yamauchi S (2006) Orientation of a key glutamine residue in the BLUF domain from AppA revealed by mutagenesis, spectroscopy, and quantum chemical calculations. J Am Chem Soc 128:5638–5639
Unno M, Kikuchi S, Masuda S (2010) Structural refinement of a key tryptophan residue in the BLUF photoreceptor AppA by ultraviolet resonance Raman spectroscopy. Biophys J 98:1949–1956
Nishina Y, Kitagawa T, Shiga K, Watari H, Yamano T (1980) Resonance Raman study of flavoenzyme-inhibitor charge-transfer interactions. Old yellow enzyme-phenol complexes. J Biochem 87:831–839
Kitagawa T, Nishina Y, Shiga K, Watari H, Matsumura Y, Yamano T (1979) Resonance Raman evidence for charge-transfer interactions of phenols with the flavin mono-nucleotide of old yellow enzyme. J Am Chem Soc 101:3376–3378
Nishina Y, Sato K, Shi R, Setoyama C, Miura R, Shiga K (2001) On the ligands in charge-transfer complexes of porcine kidney flavoenzyme D-amino acid oxidase in three redox states: a resonance Raman study. J Biochem 130:637–647
Tamaoki H, Setoyama C, Miura R, Hazekawa I, Nishina Y, Shiga K (1997) Spectroscopic studies of rat liver acyl-CoA oxidase with reference to recognition and activation of substrate. J Biochem 121:1139–1146
Nishina Y, Sato K, Miura R, Shiga K (1995) Structures of charge-transfer complexes of flavoenzyme D-amino-acid oxidase. A study by resonance Raman-spectroscopy and extended Hückel molecular-orbital method. J Biochem 118:614–620
Suzuki H, Koyama H, Nishina Y, Sato K, Shiga K (1991) A resonance Raman study on a reaction intermediate of Pseudomonas L-phenylalanine oxidase (deaminating and decarboxylating). J Biochem 110:169–172
Nishina Y, Sato K, Shiga K (1991) Isomerization of Δ1-piperideine-2-carboxylate to Δ2-piperideine-2-carboxylate on complexation with flavoprotein D-amino-acid oxidase. J Biochem 109:705–710
Nishina Y, Miura R, Tojo H, Miyake Y, Watari H, Shiga K (1986) A resonance Raman-study on the structures of complexes of flavoprotein D-amino-acid oxidase. J Biochem 99:329–337
Nishina Y, Shiga K, Miura R, Tojo H, Ohta M, Miyake Y, Yamano T, Watari H (1983) On the structures of flavoprotein D-amino-acid oxidase purple intermediates. A resonance Raman-study. J Biochem 94:1979–1990
Miura R, Nishina Y, Ohta M, Tojo H, Shiga K, Watari H, Yamano T, Miyake Y (1983) Resonance Raman study on the flavin in the purple intermediates of D-amino acid oxidase. Biochem Biophys Res Commun 111:588–594
Nishina Y, Shiga K, Watari H, Miura R, Miyake Y, Tojo H, Yamano T (1982) Resonance Raman on the purple intermediates of the flavoenzyme D-amino acid oxidase. Biochem Biophys Res Commun 106:818–822
Miura R, Nishina Y, Shiga K, Tojo H, Watari H, Miyake Y, Yamano T (1982) A resonance Raman study on the reaction intermediates of D-amino acid oxidase. J Biochem 91:837–843
Nishina Y, Shiga K, Tojo H, Miura R, Watari H, Yamano T (1981) Resonance Raman study of D-amino acid oxidase-inhibitor complexes. J Biochem 90:1515–1520
Tamaoki H, Nishina Y, Shiga K, Miura R (1999) Mechanism for the recognition and activation of substrate in medium-chain acyl-CoA dehydrogenase. J Biochem 125:285–296
Nishina Y, Sato K, Hazekawa I, Shiga K (1995) Structural modulation of 2-enoyl-CoA bound to reduced acyl-CoA dehydrogenases: a resonance Raman study of a catalytic intermediate. J Biochem 117:800–808
Hazekawa I, Nishina Y, Sato K, Shichiri M, Shiga K (1995) Substrate activating mechanism of short-chain acyl-CoA, medium-chain acyl-CoA, long-chain acyl-CoA, and isovaleryl-CoA dehydrogenases from bovine liver: a resonance Raman study on the 3-ketoacyl-CoA complexes. J Biochem 118:900–910
Nishina Y, Sato K, Shiga K, Fujii S, Kuroda K, Miura R (1992) Resonance Raman study on complexes of medium-chain acyl-CoA dehydrogenase. J Biochem 111:699–706
Williamson G, Engel PC, Nishina Y, Shiga K (1982) A resonance Raman study on the nature of charge-transfer interactions in butyryl CoA dehydrogenase. FEBS Lett 138:29–32
Schmidt J, Reinsch J, McFarland JT (1981) Mechanistic studies on fatty acyl-CoA dehydrogenase. J Biol Chem 256:11667–11670
Nishina Y, Sato K, Miura R, Matsui K, Shiga K (1998) Resonance Raman study on reduced flavin in purple intermediate of flavoenzyme: use of [4-carbonyl-18O]-enriched flavin. J Biochem 124:200–208
Miura R, Nishina Y, Sato K, Fujii S, Kuroda K, Shiga K (1993) 13C- and 15N-NMR studies on medium-chain acyl-CoA dehydrogenase reconstituted with 13C- and 15N-enriched flavin adenine dinucleotide. J Biochem 113:106–113
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Li, J., Kitagawa, T. (2014). Resonance Raman Spectroscopy. In: Weber, S., Schleicher, E. (eds) Flavins and Flavoproteins. Methods in Molecular Biology, vol 1146. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-0452-5_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-0452-5_15
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-0451-8
Online ISBN: 978-1-4939-0452-5
eBook Packages: Springer Protocols