Abstract
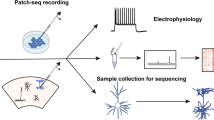
Cells exhibit diverse morphologic phenotypes, biophysical and functional properties, and gene expression patterns. Understanding how these features are interrelated at the level of single cells has been challenging due to the lack of techniques for multimodal profiling of individual cells. We recently developed Patch-seq, a technique that combines whole-cell patch clamp recording, immunohistochemistry, and single-cell RNA-sequencing (scRNA-seq) to comprehensively profile single cells. Here we present a detailed step-by-step protocol for obtaining high-quality morphological, electrophysiological, and transcriptomic data from single cells. Patch-seq enables researchers to explore the rich, multidimensional phenotypic variability among cells and to directly correlate gene expression with phenotype at the level of single cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Picelli S, Bjorklund AK, Faridani OR, Sagasser S, Winberg G, Sandberg R (2013) Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods 10(11):1096–1098. https://doi.org/10.1038/nmeth.2639
Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, Wang X, Bodeau J, Tuch BB, Siddiqui A, Lao K, Surani MA (2009) mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6(5):377–382. https://doi.org/10.1038/nmeth.1315
Eberwine J, Yeh H, Miyashiro K, Cao Y, Nair S, Finnell R, Zettel M, Coleman P (1992) Analysis of gene expression in single live neurons. Proc Natl Acad Sci U S A 89(7):3010–3014. https://doi.org/10.1073/pnas.89.7.3010
Sucher NJ, Deitcher DL (1995) PCR and patch-clamp analysis of single neurons. Neuron 14(6):1095–1100. https://doi.org/10.1016/0896-6273(95)90257-0
Sucher NJ, Deitcher DL, Baro DJ, Warrick RM, Guenther E (2000) Genes and channels: patch/voltage-clamp analysis and single-cell RT-PCR. Cell Tissue Res 302(3):295–307. https://doi.org/10.1007/s004410000289
Subkhankulova T, Yano K, Robinson HP, Livesey FJ (2010) Grouping and classifying electrophysiologically-defined classes of neocortical neurons by single cell, whole-genome expression profiling. Front Mol Neurosci 3:10. https://doi.org/10.3389/fnmol.2010.00010
Qiu S, Luo S, Evgrafov O, Li R, Schroth GP, Levitt P, Knowles JA, Wang K (2012) Single-neuron RNA-Seq: technical feasibility and reproducibility. Front Genet 3:124. https://doi.org/10.3389/fgene.2012.00124
Cadwell CR, Palasantza A, Jiang X, Berens P, Deng Q, Yilmaz M, Reimer J, Shen S, Bethge M, Tolias KF, Sandberg R, Tolias AS (2016) Electrophysiological, transcriptomic and morphologic profiling of single neurons using patch-seq. Nat Biotechnol 34(2):199–203. https://doi.org/10.1038/nbt.3445
Fuzik J, Zeisel A, Mate Z, Calvigioni D, Yanagawa Y, Szabo G, Linnarsson S, Harkany T (2016) Integration of electrophysiological recordings with single-cell RNA-seq data identifies neuronal subtypes. Nat Biotechnol 34(2):175–183. https://doi.org/10.1038/nbt.3443
Cadwell CR, Scala F, Li S, Livrizzi G, Shen S, Sandberg R, Jiang X, Tolias AS (2017) Multimodal profiling of single-cell morphology, electrophysiology, and gene expression using Patch-seq. Nat Protoc 12(12):2531–2553. https://doi.org/10.1038/nprot.2017.120
Scala F, Kobak D, Bernabucci M, Bernaerts Y, Cadwell CR, Castro JR, Hartmanis L, Jiang X, Laturnus S, Miranda E, Mulherkar S, Tan ZH, Yao Z, Zeng H, Sandberg R, Berens P, Tolias AS (2021) Phenotypic variation of transcriptomic cell types in mouse motor cortex. Nature 598(7879):144–150. https://doi.org/10.1038/s41586-020-2907-3
Scala F, Kobak D, Shan S, Bernaerts Y, Laturnus S, Cadwell CR, Hartmanis L, Froudarakis E, Castro JR, Tan ZH, Papadopoulos S, Patel SS, Sandberg R, Berens P, Jiang X, Tolias AS (2019) Layer 4 of mouse neocortex differs in cell types and circuit organization between sensory areas. Nat Commun 10(1):4174. https://doi.org/10.1038/s41467-019-12058-z
Picelli S, Bjorklund AK, Reinius B, Sagasser S, Winberg G, Sandberg R (2014) Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res 24(12):2033–2040. https://doi.org/10.1101/gr.177881.114
Jiang X, Shen S, Cadwell CR, Berens P, Sinz F, Ecker AS, Patel S, Tolias AS (2015) Principles of connectivity among morphologically defined cell types in adult neocortex. Science 350(6264):aac9462. https://doi.org/10.1126/science.aac9462
Kobak D, Berens P (2019) The art of using t-SNE for single-cell transcriptomics. Nat Commun 10(1):5416. https://doi.org/10.1038/s41467-019-13056-x
Kobak D, Bernaerts Y, Weis MA, Scala F, Tolias AS, Berens P (2021) Sparse reduced-rank regression for exploratory visualisation of paired multivariate data. J R Stat Soc Ser C Appl Stat 70(4):980–1000. https://doi.org/10.1111/rssc.12494
Tripathy SJ, Toker L, Bomkamp C, Mancarci BO, Belmadani M, Pavlidis P (2018) Assessing transcriptome quality in patch-seq datasets. Front Mol Neurosci 11:363. https://doi.org/10.3389/fnmol.2018.00363
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Cadwell, C.R., Tolias, A.S. (2024). Patch-seq: Multimodal Profiling of Single-Cell Morphology, Electrophysiology, and Gene Expression. In: Gužvić, M. (eds) Single Cell Analysis. Methods in Molecular Biology, vol 2752. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3621-3_15
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3621-3_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3620-6
Online ISBN: 978-1-0716-3621-3
eBook Packages: Springer Protocols