Abstract
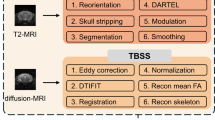
In this chapter, we provide a comprehensive historical overview of algorithms for feature extraction of structural magnetic resonance imaging data and the corresponding analytical frameworks. We then focus on the use of T1-weighted images, which still represent the working horse of surface- and voxel-based morphometry to elaborate on the complex relationships between magnetic resonance imaging contrast and underlying brain tissue properties. This critical point is the motivation for embarking on novel structural imaging protocols based on biophysical models, which are sensitive to tissue myelin, iron, and water content. We expand on the concept of voxel-based quantification to demonstrate the added value to existing methods using T1-weighted data—both in terms of robust brain tissue classification and of straightforward neurobiological interpretation of the obtained results. We do not stop short of considering the unresolved issues in voxel-based quantification currently implemented in the data processing and analysis framework of Statistical Parametric Mapping (SPM12). We conclude with an outlook to the future developments and perspectives in computational anatomy, particularly the need for integration of the available magnetic resonance imaging contrasts at the level of statistical analysis without hampering the interpretability of our findings.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Coffey CE, Wilkinson WE, Parashos IA, Soady SA, Sullivan RJ, Patterson LJ et al (1992) Quantitative cerebral anatomy of the aging human brain: a cross-sectional study using magnetic resonance imaging. Neurology 42(3 Pt 1):527–536
Coffey CE, Wilkinson WE, Weiner RD, Parashos IA, Djang WT, Webb MC et al (1993) Quantitative cerebral anatomy in depression. A controlled magnetic resonance imaging study. Arch Gen Psychiatry 50(1):7–16
Wright IC, McGuire PK, Poline JB, Travere JM, Murray RM, Frith CD et al (1995) A voxel-based method for the statistical analysis of gray and white matter density applied to schizophrenia. NeuroImage 2(4):244–252
Bullmore E, Brammer M, Rouleau G, Everitt B, Simmons A, Sharma T et al (1995) Computerized brain tissue classification of magnetic resonance images: a new approach to the problem of partial volume artifact. NeuroImage 2(2):133–147
Ashburner J, Friston KJ (2000) Voxel-based morphometry--the methods. NeuroImage 11(6 Pt 1):805–821
Ashburner J, Neelin P, Collins DL, Evans A, Friston K (1997) Incorporating prior knowledge into image registration. NeuroImage 6(4):344–352
Ashburner J, Andersson JL, Friston KJ (1999) High-dimensional image registration using symmetric priors. NeuroImage 9(6 Pt 1):619–628
Ashburner J, Friston KJ (1999) Nonlinear spatial normalization using basis functions. Hum Brain Mapp 7(4):254–266
Dale AM, Fischl B, Sereno MI (1999) Cortical surface-based analysis. I Segmentation and surface reconstruction. NeuroImage 9(2):179–194
Fischl B, Sereno MI, Dale AM (1999) Cortical surface-based analysis. II: Inflation, flattening, and a surface-based coordinate system. NeuroImage 9(2):195–207
Fischl B, Sereno MI, Tootell RB, Dale AM (1999) High-resolution intersubject averaging and a coordinate system for the cortical surface. Hum Brain Mapp 8(4):272–284
Fischl B, Dale AM (2000) Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc Natl Acad Sci 97(20):11050–11055
Hutton C, Draganski B, Ashburner J, Weiskopf N (2009) A comparison between voxel-based cortical thickness and voxel-based morphometry in normal aging. NeuroImage 48(2):371–380
Ashburner J, Csernansky JG, Davatzikos C, Fox NC, Frisoni GB, Thompson PM (2003) Computer-assisted imaging to assess brain structure in healthy and diseased brains. Lancet Neurol 2(2):79–88
Eickhoff SB, Constable RT, Yeo BTT (2018) Topographic organization of the cerebral cortex and brain cartography. NeuroImage 170:332–347
Eickhoff SB, Yeo BTT, Genon S (2018) Imaging-based parcellations of the human brain. Nat Rev Neurosci 19(11):672–686
Weber CJ, Carrillo MC, Jagust W, Jack CR, Shaw LM, Trojanowski JQ et al (2021) The Worldwide Alzheimer’s Disease Neuroimaging Initiative: ADNI-3 updates and global perspectives. Alzheimers Dement N Y N 7(1):e12226
Tafuri B, Lombardi A, Nigro S, Urso D, Monaco A, Pantaleo E et al (2022) The impact of harmonization on radiomic features in Parkinson’s disease and healthy controls: a multicenter study. Front Neurosci 16:1012287
Miller KL, Alfaro-Almagro F, Bangerter NK, Thomas DL, Yacoub E, Xu J et al (2016) Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nat Neurosci 19(11):1523
Mascarell Maričić L, Walter H, Rosenthal A, Ripke S, Quinlan EB, Banaschewski T et al (2020) The IMAGEN study: a decade of imaging genetics in adolescents. Mol Psychiatry 25(11):2648–2671
Thompson PM, Jahanshad N, Ching CRK, Salminen LE, Thomopoulos SI, Bright J et al (2020) ENIGMA and global neuroscience: a decade of large-scale studies of the brain in health and disease across more than 40 countries. Transl Psychiatry 10(1):100
Marek S, Tervo-Clemmens B, Calabro FJ, Montez DF, Kay BP, Hatoum AS et al (2022) Reproducible brain-wide association studies require thousands of individuals. Nature 603(7902):654–660
Rosas HD, Liu AK, Hersch S, Glessner M, Ferrante RJ, Salat DH et al (2002) Regional and progressive thinning of the cortical ribbon in Huntington’s disease. Neurology 58(5):695–701
Aylward EH, Brandt J, Codori AM, Mangus RS, Barta PE, Harris GJ (1994) Reduced basal ganglia volume associated with the gene for Huntington’s disease in asymptomatic at-risk persons. Neurology 44(5):823–828
Modenato C, Martin-Brevet S, Moreau CA, Rodriguez-Herreros B, Kumar K, Draganski B et al (2021) Lessons learned from neuroimaging studies of copy number variants: a systematic review. Biol Psychiatry 90(9):596–610
Kopal J, Kumar K, Saltoun K, Modenato C, Moreau CA, Martin-Brevet S et al (2022) Rare CNVs and phenome-wide profiling: a tale of brain-structural divergence and phenotypical convergence [Internet]. bioRxiv [cited 2022 Nov 27]. p. 2022.04.23.489093. Available from: https://www.biorxiv.org/content/10.1101/2022.04.23.489093v1
Kawasaki Y, Suzuki M, Kherif F, Takahashi T, Zhou SY, Nakamura K et al (2007) Multivariate voxel-based morphometry successfully differentiates schizophrenia patients from healthy controls. NeuroImage 34(1):235–242
Klöppel S, Stonnington CM, Chu C, Draganski B, Scahill RI, Rohrer JD et al (2008) Automatic classification of MR scans in Alzheimer’s disease. Brain J Neurol 131(Pt 3):681–689
Yang Z, Nasrallah IM, Shou H, Wen J, Doshi J, Habes M et al (2021) A deep learning framework identifies dimensional representations of Alzheimer’s Disease from brain structure. Nat Commun 12(1):7065
Draganski B, Gaser C, Busch V, Schuierer G, Bogdahn U, May A (2004) Neuroplasticity: changes in grey matter induced by training. Nature 427(6972):311–312
Sowell ER, Thompson PM, Leonard CM, Welcome SE, Kan E, Toga AW (2004) Longitudinal mapping of cortical thickness and brain growth in normal children. J Neurosci 24(38):8223–8231
Thompson PM, Vidal C, Giedd JN, Gochman P, Blumenthal J, Nicolson R et al (2001) Mapping adolescent brain change reveals dynamic wave of accelerated gray matter loss in very early-onset schizophrenia. Proc Natl Acad Sci U S A 98(20):11650–11655
Dicks E, Vermunt L, van der Flier WM, Visser PJ, Barkhof F, Scheltens P et al (2019) Modeling grey matter atrophy as a function of time, aging or cognitive decline show different anatomical patterns in Alzheimer’s disease. NeuroImage Clin 22:101786
Thomas C, Baker CI (2013) Teaching an adult brain new tricks: a critical review of evidence for training-dependent structural plasticity in humans. NeuroImage 73:225–236
Draganski B, Kherif F (2013) In vivo assessment of use-dependent brain plasticity--beyond the ‘one trick pony’ imaging strategy. NeuroImage 73:255–259; discussion 265–267
Young AL, Oxtoby NP, Daga P, Cash DM, Fox NC, Ourselin S et al (2014) A data-driven model of biomarker changes in sporadic Alzheimer’s disease. Brain J Neurol 137(Pt 9):2564–2577
Fonteijn HM, Modat M, Clarkson MJ, Barnes J, Lehmann M, Hobbs NZ et al (2012) An event-based model for disease progression and its application in familial Alzheimer’s disease and Huntington’s disease. NeuroImage 60(3):1880–1889
Oxtoby NP, Young AL, Cash DM, Benzinger TLS, Fagan AM, Morris JC et al (2018) Data-driven models of dominantly-inherited Alzheimer’s disease progression. Brain J Neurol 141(5):1529–1544
Archetti D, Ingala S, Venkatraghavan V, Wottschel V, Young AL, Bellio M et al (2019) Multi-study validation of data-driven disease progression models to characterize evolution of biomarkers in Alzheimer’s disease. NeuroImage Clin 24:101954
Mechelli A, Friston KJ, Frackowiak RS, Price CJ (2005) Structural covariance in the human cortex. J Neurosci 25(36):8303–8310
Evans AC (2013) Networks of anatomical covariance. NeuroImage 80:489–504
Fox MD (2018) Mapping symptoms to brain networks with the human connectome. N Engl J Med 379(23):2237–2245
Fox MD, Buckner RL, Liu H, Chakravarty MM, Lozano AM, Pascual-Leone A (2014) Resting-state networks link invasive and noninvasive brain stimulation across diverse psychiatric and neurological diseases. Proc Natl Acad Sci U S A 111(41):E4367–E4375
Padmanabhan JL, Cooke D, Joutsa J, Siddiqi SH, Ferguson M, Darby RR et al (2019) A human depression circuit derived from focal brain lesions. Biol Psychiatry 86(10):749–758
Amunts K, Hawrylycz MJ, Van Essen DC, Van Horn JD, Harel N, Poline JB et al (2014) Interoperable atlases of the human brain. NeuroImage 99:525–532
Amunts K, Zilles K (2015) Architectonic mapping of the human brain beyond Brodmann. Neuron 88(6):1086–1107
Amunts K, Schleicher A, Zilles K (2007) Cytoarchitecture of the cerebral cortex--more than localization. NeuroImage 37(4):1061–1065; discussion 1066–1068
Annese J, Pitiot A, Dinov ID, Toga AW (2004) A myelo-architectonic method for the structural classification of cortical areas. NeuroImage 21(1):15–26
Lorio S, Kherif F, Ruef A, Melie-Garcia L, Frackowiak R, Ashburner J et al (2016) Neurobiological origin of spurious brain morphological changes: a quantitative MRI study. Hum Brain Mapp 37(5):1801–1815
Natu VS, Gomez J, Barnett M, Jeska B, Kirilina E, Jaeger C et al (2019) Apparent thinning of human visual cortex during childhood is associated with myelination. Proc Natl Acad Sci U S A
Helms G, Dathe H, Kallenberg K, Dechent P (2008) High-resolution maps of magnetization transfer with inherent correction for RF inhomogeneity and T1 relaxation obtained from 3D FLASH MRI. Magn Reson Med 60(6):1396–1407
Weiskopf N, Suckling J, Williams G, Correia MM, Inkster B, Tait R et al (2013) Quantitative multi-parameter mapping of R1, PD*, MT, and R2* at 3T: a multi-center validation. Front Neurosci 7:95
Draganski B, Ashburner J, Hutton C, Kherif F, Frackowiak RSJ, Helms G et al (2011) Regional specificity of MRI contrast parameter changes in normal ageing revealed by voxel-based quantification (VBQ). NeuroImage 55(4):1423–1434
Weiskopf N, Edwards LJ, Helms G, Mohammadi S, Kirilina E (2021) Quantitative magnetic resonance imaging of brain anatomy and in vivo histology. Nat Rev Phys [Internet] [cited 2021 Nov 1];3(8):570–88. Available from: https://www.nature.com/articles/s42254-021-00326-1
Lorio S, Lutti A, Kherif F, Ruef A, Dukart J, Chowdhury R et al (2014) Disentangling in vivo the effects of iron content and atrophy on the ageing human brain. NeuroImage 103:280–289
Helms G, Draganski B, Frackowiak R, Ashburner J, Weiskopf N (2009) Improved segmentation of deep brain grey matter structures using magnetization transfer (MT) parameter maps. NeuroImage 47(1):194–198
Lorio S, Fresard S, Adaszewski S, Kherif F, Chowdhury R, Frackowiak RS et al (2016) New tissue priors for improved automated classification of subcortical brain structures on MRI. NeuroImage 130:157–166
Hallgren B, Sourander P (1958) The effect of age on the non-haemin iron in the human brain. J Neurochem 3(1):41–51
Morawski M, Kirilina E, Scherf N, Jäger C, Reimann K, Trampel R et al (2018) Developing 3D microscopy with CLARITY on human brain tissue: towards a tool for informing and validating MRI-based histology. NeuroImage 182:417–428
Kirilina E, Helbling S, Morawski M, Pine K, Reimann K, Jankuhn S et al (2020) Superficial white matter imaging: contrast mechanisms and whole-brain in vivo mapping. Sci Adv 6(41):eaaz9281
Brammerloh M, Morawski M, Friedrich I, Reinert T, Lange C, Pelicon P et al (2021) Measuring the iron content of dopaminergic neurons in substantia nigra with MRI relaxometry. NeuroImage 239:118255
Natu VS, Rosenke M, Wu H, Querdasi FR, Kular H, Lopez-Alvarez N et al (2021) Infants’ cortex undergoes microstructural growth coupled with myelination during development. Commun Biol 4(1):1191
Grotheer M, Yeatman J, Grill-Spector K (2021) White matter fascicles and cortical microstructure predict reading-related responses in human ventral temporal cortex. NeuroImage 227:117669
Grotheer M, Rosenke M, Wu H, Kular H, Querdasi FR, Natu VS et al (2022) White matter myelination during early infancy is linked to spatial gradients and myelin content at birth. Nat Commun 13(1):997
Ziegler G, Hauser TU, Moutoussis M, Bullmore ET, Goodyer IM, Fonagy P et al (2019) Compulsivity and impulsivity traits linked to attenuated developmental frontostriatal myelination trajectories. Nat Neurosci 22(6):992–999
Ziegler G, Moutoussis M, Hauser TU, Fearon P, Bullmore ET, Goodyer IM et al (2020) Childhood socio-economic disadvantage predicts reduced myelin growth across adolescence and young adulthood. Hum Brain Mapp 41(12):3392–3402
Granziera C, Wuerfel J, Barkhof F, Calabrese M, De Stefano N, Enzinger C et al (2021) Quantitative magnetic resonance imaging towards clinical application in multiple sclerosis. Brain J Neurol 144(5):1296–1311
Slater DA, Melie-Garcia L, Preisig M, Kherif F, Lutti A, Draganski B (2019) Evolution of white matter tract microstructure across the life span. Hum Brain Mapp 40(7):2252–2268
Taubert M, Roggenhofer E, Melie-Garcia L, Muller S, Lehmann N, Preisig M et al (2020) Converging patterns of aging-associated brain volume loss and tissue microstructure differences. Neurobiol Aging 88:108–118
Callaghan MF, Freund P, Draganski B, Anderson E, Cappelletti M, Chowdhury R et al (2014) Widespread age-related differences in the human brain microstructure revealed by quantitative magnetic resonance imaging. Neurobiol Aging 35(8):1862–1872
Trofimova O, Loued-Khenissi L, DiDomenicantonio G, Lutti A, Kliegel M, Stringhini S et al (2021) Brain tissue properties link cardio-vascular risk factors, mood and cognitive performance in the CoLaus|PsyCoLaus epidemiological cohort. Neurobiol Aging 102:50–63
David G, Pfyffer D, Vallotton K, Pfender N, Thompson A, Weiskopf N et al (2021) Longitudinal changes of spinal cord grey and white matter following spinal cord injury. J Neurol Neurosurg Psychiatry 92(11):1222–1230
Seif M, Leutritz T, Schading S, Emmengger T, Curt A, Weiskopf N et al (2022) Reliability of multi-parameter mapping (MPM) in the cervical cord: a multi-center multi-vendor quantitative MRI study. NeuroImage 264:119751
Gyger L, Ramponi C, Mall JF, Swierkosz-Lenart K, Stoyanov D, Lutti A et al (2021) Temporal trajectory of brain tissue property changes induced by electroconvulsive therapy. NeuroImage 232:117895
Lazari A, Salvan P, Verhagen L, Cottaar M, Papp D, van der Werf OJ et al (2022) A macroscopic link between interhemispheric tract myelination and cortico-cortical interactions during action reprogramming. Nat Commun 13(1):4253
Stikov N, Campbell JS, Stroh T, Lavelée M, Frey S, Novek J et al (2015) In vivo histology of the myelin g-ratio with magnetic resonance imaging. NeuroImage 118:397–405
Campbell JSW, Leppert IR, Narayanan S, Boudreau M, Duval T, Cohen-Adad J et al (2018) Promise and pitfalls of g-ratio estimation with MRI. NeuroImage 182:80–96
Clark IA, Mohammadi S, Callaghan MF, Maguire EA (2022) Conduction velocity along a key white matter tract is associated with autobiographical memory recall ability. elife 11:e79303
Gracien RM, Maiworm M, Brüche N, Shrestha M, Nöth U, Hattingen E et al (2020) How stable is quantitative MRI? - Assessment of intra- and inter-scanner-model reproducibility using identical acquisition sequences and data analysis programs. NeuroImage 207:116364
Karakuzu A, Biswas L, Cohen-Adad J, Stikov N (2022) Vendor-neutral sequences and fully transparent workflows improve inter-vendor reproducibility of quantitative MRI. Magn Reson Med 88(3):1212–1228
Tabelow K, Balteau E, Ashburner J, Callaghan MF, Draganski B, Helms G et al (2019) hMRI - a toolbox for quantitative MRI in neuroscience and clinical research. NeuroImage 194:191–210
Callaghan MF, Lutti A, Ashburner J, Balteau E, Corbin N, Draganski B et al (2019) Example dataset for the hMRI toolbox. Data Brief 25:104132
Depierreux F, Parmentier E, Mackels L, Baquero K, Degueldre C, Balteau E et al (2021) Parkinson’s disease multimodal imaging: F-DOPA PET, neuromelanin-sensitive and quantitative iron-sensitive MRI. NPJ Park Dis 7(1):57
Drori E, Berman S, Mezer AA (2022) Mapping microstructural gradients of the human striatum in normal aging and Parkinson’s disease. Sci Adv 8(28):eabm1971
Acknowledgments
BD is supported by the Swiss National Science Foundation (grants 32003B_159780, CRSK-3_190185, 324730_192755, and the Sinergia VascX project), the EU ERA-NET iSee project, the SPHN SACR project, the CLIMACT project, the Leenaards Foundation, and the HeadFirst Innosuisse project. LREN is grateful to the ROGER DE SPOELBERCH and Partridge Foundations for their generous financial support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Draganski, B., Paunova, R., Latypova, A., Kherif, F. (2023). Computational Anatomy Going Beyond Brain Morphometry. In: Stoyanov, D., Draganski, B., Brambilla, P., Lamm, C. (eds) Computational Neuroscience. Neuromethods, vol 199. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3230-7_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3230-7_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3229-1
Online ISBN: 978-1-0716-3230-7
eBook Packages: Springer Protocols