Abstract
The initial discovery of left-handed Z-DNA was met with great excitement as a dramatic alternative to the right-handed double-helical conformation of canonical B-DNA. In this chapter, we describe the workings of the program ZHUNT as a computational approach to mapping Z-DNA in genomic sequences using a rigorous thermodynamic model for the transition between the two conformations (the B–Z transition). The discussion starts with a brief summary of the structural properties that differentiate Z- from B-DNA, focusing on those properties that are particularly relevant to the B–Z transition and the junction that splices a left- to right-handed DNA duplex. We then derive the statistical mechanics (SM) analysis of the zipper model that describes the cooperative B–Z transition and show that this analysis very accurately simulates this behavior of naturally occurring sequences that are induced to undergo the B–Z transition through negative supercoiling. A description of the ZHUNT algorithm and its validation are presented, followed by how the program had been applied for genomic and phylogenomic analyses in the past and how a user can access the online version of the program. Finally, we present a new version of ZHUNT (called mZHUNT) that has been parameterized to analyze sequences that contain 5-methylcytosine bases and compare the results of the ZHUNT and mZHUNT analyses on native and methylated yeast chromosome 1.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
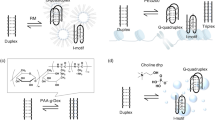
Similar content being viewed by others
References
Avery OT, MacLeod CM, McCarty M (1944) Studies on the chemical nature of the substance inducing transformation of pneumococcal types induction of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus type III. J Exp Med 79(2):137–158. https://doi.org/10.1084/jem.79.2.137
Hershey AD, Chase M (1952) Independent functions of viral protein and nucleic acid in growth of bacteriophage. J Gen Physiol 36(1):39–56. https://doi.org/10.1085/jgp.36.1.39
Watson JD, Crick FH (1953) Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171(4356):737–738
Wang AHJ, Quigley GJ, Kolpak FJ, Crawford JL, Vanboom JH, Vandermarel G et al (1979) Molecular-structure of a left-handed double helical DNA fragment at atomic resolution. Nature 282(5740):680–686. https://doi.org/10.1038/282680a0
Gannon HS, Zou T, Kiessling MK, Gao GF, Cai D, Choi PS et al (2018) Identification of ADAR1 adenosine deaminase dependency in a subset of cancer cells. Nat Comm 9:5450. https://doi.org/10.1038/s41467-018-07824-4
Suram A, Rao LKS, Latha KS, Viswamitra MA (2002) First evidence to show the topological change of DNA from B-DNA to Z-DNA conformation in the hippocampus of Alzheimer’s brain. NeuroMolecular Med 2(3):289–297. https://doi.org/10.1385/Nmm:2:3:289
Mitsui Y, Langridge R, Shortle BE, Cantor CR, Grant RC, Kodama M et al (1970) Physical and enzymatic studies on poly D(I-C) poly D(I-C), an unusual double-helical DNA. Nature 228(5277):1166. https://doi.org/10.1038/2281166a0
Pohl FM, Jovin TM (1972) Salt-induced cooperative conformational change of a synthetic DNA – equilibrium and kinetic studies with poly(dg-dc). J Mol Biol 67(3):375. https://doi.org/10.1016/0022-2836(72)90457-3
Wing R, Drew H, Takano T, Broka C, Tanaka S, Itakura K et al (1980) Crystal-structure analysis of a complete turn of B-DNA. Nature 287(5784):755–758. https://doi.org/10.1038/287755a0
Ho PS, Mooers BHM (1997) Z-DNA crystallography. Biopolymers 44(1):65–90. https://doi.org/10.1002/(Sici)1097-0282(1997)44:1<65::Aid-Bip5>3.0.Co;2-Y
Kagawa TF, Stoddard D, Zhou GW, Ho PS (1989) Quantitative analysis of DNA secondary structure from solvent-accessible surfaces: the B- to Z-DNA transition as a model. Biochemistry 28(16):6642–6651
Ho PS, Kagawa TF, Tseng K, Schroth GP, Zhou GW (1991) Prediction of a crystallization pathway for Z-DNA Hexanucleotides. Science 254(5034):1003–1006. https://doi.org/10.1126/science.1948069
Behe M, Felsenfeld G (1981) Effects of methylation on a synthetic polynucleotide – the B-Z transition in poly(dG-m5dC).Poly(dG-m5dC). Proc Natl Acad Sci USA 78(3):1619–1623. https://doi.org/10.1073/pnas.78.3.1619
Kagawa TF, Howell ML, Tseng K, Ho PS (1993) Effects of base substituents on the hydration of B- and Z-DNA: correlations to the B- to Z-DNA transition. Nucleic Acids Res 21(25):5978–5986. https://doi.org/10.1093/nar/21.25.5978
Howell ML, Schroth GP, Ho PS (1996) Sequence-dependent effects of spermine on the thermodynamics of the B-DNA to Z-DNA transition. Biochemistry 35(48):15373–15382. https://doi.org/10.1021/bi961881i
Peck LJ, Nordheim A, Rich A, Wang JC (1982) Flipping of cloned D(Pcpg)N.D(Pcpg)N DNA-sequences from right-handed to left-handed helical structure by salt, co(iii), or negative supercoiling. Proc Natl Acad Sci, USA 79(15):4560–4564. https://doi.org/10.1073/pnas.79.15.4560
Nordheim A, Peck LJ, Lafer EM, Stollar BD, Wang JC, Rich A (1983) Supercoiling and left-handed Z-DNA. Cold Spring Harb Symp Quant Biol 47(Pt 1):93–100
Peck LJ, Wang JC (1983) Energetics of B-to-Z transition in DNA. Proc Natl Acad Sci U S A 80(20):6206–6210. https://doi.org/10.1073/pnas.80.20.6206
Wang JC, Peck LJ, Becherer K (1983) DNA supercoiling and its effects on DNA structure and function. Cold Spring Harb Symp Quant Biol 47(Pt 1):85–91
Nickol J, Behe M, Felsenfeld G (1982) Effect of the B--Z transition in poly(dG-m5dC). poly(dG-m5dC) on nucleosome formation. Proc Natl Acad Sci U S A 79(6):1771–1775
Ausio J, Zhou G, van Holde K (1987) A reexamination of the reported B----Z DNA transition in nucleosomes reconstituted with poly(dG-m5dC).poly(dG-m5dC). Biochemistry 26(18):5595–5599
Liu LF, Wang JC (1987) Supercoiling of the DNA-template during transcription. Proc Natl Acad Sci U S A 84(20):7024–7027. https://doi.org/10.1073/pnas.84.20.7024
Ellison MJ, Kelleher RJ 3rd, Wang AH, Habener JF, Rich A (1985) Sequence-dependent energetics of the B-Z transition in supercoiled DNA containing nonalternating purine-pyrimidine sequences. Proc Natl Acad Sci U S A 82(24):8320–8324
Ellison MJ, Feigon J, Kelleher RJ 3rd, Wang AH, Habener JF, Rich A (1986) An assessment of the Z-DNA forming potential of alternating dA-dT stretches in supercoiled plasmids. Biochemistry 25(12):3648–3655
Dickerson RE (1992) DNA-structure from A to Z. Method Enzymol 211:67–111
Ha SC, Lowenhaupt K, Rich A, Kim YG, Kim KK (2005) Crystal structure of a junction between B-DNA and Z-DNA reveals two extruded bases. Nature 437(7062):1183–1186
Zimm BH, Bragg JK (1959) Theory of the phase transition between helix and random coil in polypeptide chains. J Chem Phys 31(2):526–535. https://doi.org/10.1063/1.1730390
Nordheim A, Lafer EM, Peck LJ, Wang JC, Stollar BD, Rich A (1982) Negatively supercoiled plasmids contain left-handed Z-DNA segments as detected by specific antibody-binding. Cell 31(2):309–318. https://doi.org/10.1016/0092-8674(82)90124-6
Pulleyblank DE, Haniford DB, Morgan AR (1985) A structural basis for S1 nuclease sensitivity of double-stranded DNA. Cell 42(1):271–280. https://doi.org/10.1016/S0092-8674(85)80122-7
Dicapua E, Stasiak A, Koller T, Brahms S, Thomae R, Pohl FM (1983) Torsional stress induces left-handed helical stretches in DNA of Natural Base sequence - circular-dichroism and antibody-binding. EMBO J 2(9):1531–1535. https://doi.org/10.1002/j.1460-2075.1983.tb01619.x
Revet B, Zarling DA, Jovin TM, Delain E (1984) Different Z DNA forming sequences are revealed in phi-X174 Rfi by high-resolution dark-field Immuno-electron microscopy. EMBO J 3(13):3353–3358. https://doi.org/10.1002/j.1460-2075.1984.tb02303.x
Ho PS, Ellison MJ, Quigley GJ, Rich A (1986) A computer aided thermodynamic approach for predicting the formation of Z-DNA in naturally occurring sequences. EMBO J 5(10):2737–2744
Ho PS (2009) Thermogenomics: thermodynamic-based approaches to genomic analyses of DNA structure. Methods 47(3):159–167. https://doi.org/10.1016/j.ymeth.2008.09.007
Kladde MP, Kohwi Y, Kohwi-Shigematsu T, Gorski J (1994) The non-B-DNA structure of d(CA/TG)n differs from that of Z-DNA. Proc Natl Acad Sci U S A 91(5):1898–1902
Ho PS (1994) The non-B-DNA structure of d(CA/TG)n does not differ from that of Z-DNA. Proc Natl Acad Sci U S A 91(20):9549–9553
Collins FS (1991) The genome project and human health. FASEB J 5(1):77
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG et al (2001) The sequence of the human genome. Science 291(5507):1304–1351
Szustakowski J, Consor IHGS (2001) Initial sequencing and analysis of the human genome. Nature 409(6822):860–921. 411(6838):720
Schroth GP, Chou PJ, Ho PS (1992) Mapping Z-DNA in the human genome. Computer-aided mapping reveals a nonrandom distribution of potential Z-DNA-forming sequences in human genes. J Biol Chem 267(17):11846–11855
Liu R, Liu H, Chen X, Kirby M, Brown PO, Zhao K (2001) Regulation of CSF1 promoter by the SWI/SNF-like BAF complex. Cell 106(3):309–318
Champ PC, Maurice S, Vargason JM, Camp T, Ho PS (2004) Distributions of Z-DNA and nuclear factor I in human chromosome 22: a model for coupled transcriptional regulation. Nucleic Acids Res 32(22):6501–6510
Collins FS, Lander ES, Rogers J, Waterston RH, Conso IHGS (2004) Finishing the euchromatic sequence of the human genome. Nature 431(7011):931–945. https://doi.org/10.1038/nature03001
Khuu P, Sandor M, DeYoung J, Ho PS (2007) Phylogenomic analysis of the emergence of GC-rich transcription elements. Proc Natl Acad Sci U S A 104(42):16528–16533
Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ et al (2016) A new view of the tree of life. Nat Microbiol 1:16048. https://doi.org/10.1038/nmicrobiol.2016.48
Mulligan CJ (2018) Insights from epigenetic studies on human health and evolution. Curr Opin Genet Dev 53:36–42. https://doi.org/10.1016/j.gde.2018.06.008
Ho PS, Quigley GJ, Tilton RF Jr, Rich A (1988) Hydration of methylated and nonmethylated B-DNA and Z-DNA. J Phys Chem 92:939–945
Wang YH, Wang AQ, Liu ZJ, Thurman AL, Powers LS, Zou M et al (2019) Single-molecule long-read sequencing reveals the chromatin basis of gene expression. Genome Res 29(8):1329–1342. https://doi.org/10.1101/gr.251116.119
Zhabinskaya D, Benham CJ (2011) Theoretical analysis of the stress induced B-Z transition in superhelical DNA. PLoS Comput Biol 7(1):e1001051. https://doi.org/10.1371/journal.pcbi.1001051
Li H, Xiao J, Li JM, Lu L, Feng S, Droge P (2009) Human genomic Z-DNA segments probed by the Z domain of ADAR1. Nucleic Acids Res 37(8):2737–2746. https://doi.org/10.1093/nar/gkp124
Beknazarov N, Jin S, Poptsova M (2020) Deep learning approach for predicting functional Z-DNA regions using omics data. Sci Rep UK 10(1):19134. https://doi.org/10.1038/s41598-020-76203-1
Shin SI, Ham S, Park J, Seo SH, Lim CH, Jeon H et al (2016) Z-DNA-forming sites identified by ChIP-Seq are associated with actively transcribed regions in the human genome. DNA Res 23(5):477–486. https://doi.org/10.1093/dnares/dsw031
Drew HR, Wing RM, Takano T, Broka C, Tanaka S, Itakura K et al (1981) Structure of a B-DNA dodecamer: conformation and dynamics. Proc Natl Acad Sci U S A 78:2179–2183
Luo ZP, Dauter M, Dauter Z (2014) Phosphates in the Z-DNA dodecamer are flexible, but their P-SAD signal is sufficient for structure solution. Acta Crystallogr Sect D Struct Biol 70:1790–1800. https://doi.org/10.1107/S1399004714004684
Carter M, Ho PS (2011) DNA structure: alphabet soup for the cellular soul. In: Seligmann H (ed) DNA replication – current advances. InTech, London, pp 3–28
Acknowledgments
The studies in Ho laboratory were supported by a grant from the National Science Foundation (CHE-1905328 and MCB-2124202). We thank A. N. Ho for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Czarny, R.S., Ho, P.S. (2023). Thermogenomic Analysis of Left-Handed Z-DNA Propensities in Genomes. In: Kim, K.K., Subramani, V.K. (eds) Z-DNA. Methods in Molecular Biology, vol 2651. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3084-6_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3084-6_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3083-9
Online ISBN: 978-1-0716-3084-6
eBook Packages: Springer Protocols