Abstract
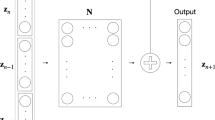
The dynamics of systems biological processes are usually modeled by a system of ordinary differential equations (ODEs) with many unknown parameters that need to be inferred from noisy and sparse measurements. Here, we introduce systems-biology-informed neural networks for parameter estimation by incorporating the system of ODEs into the neural networks. To complete the workflow of system identification, we also describe structural and practical identifiability analysis to analyze the identifiability of parameters. We use the ultradian endocrine model for glucose-insulin interaction as the example to demonstrate all these methods and their implementation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kitano H (2002) Systems biology: a brief overview. Science 295(5560):1662–1664
Sturis J, Polonsky KS, Mosekilde E, Van Cauter E (1991) Computer model for mechanisms underlying ultradian oscillations of insulin and glucose. Am J Physiol Endocrinol Metab 260(5):E801–E809
Albers DJ, Levine M, Gluckman B, Ginsberg H, Hripcsak G, Mamykina L (2017) Personalized glucose forecasting for type 2 diabetes using data assimilation. PLoS Comput Biol 13(4):e1005232
Albers DJ, Elhadad N, Tabak E, Perotte A, Hripcsak G (2014) Dynamical phenotyping: using temporal analysis of clinically collected physiologic data to stratify populations. PloS One 9(6):e96443
Dong R, Goodbrake C, Harrington HA, Pogudin G (2021) Differential elimination for dynamical models via projections with applications to structural identifiability. arXiv preprint arXiv:2111.00991
Yazdani A, Lu L, Raissi M, Karniadakis GE (2020) Systems biology informed deep learning for inferring parameters and hidden dynamics. PLoS Comput Biol 16(11):e1007575
Abadi M, Barham P, Chen J, Chen Z, Davis A, Dean J, Devin M, Ghemawat S, Irving G, Isard M et al (2016) Tensorflow: a system for large-scale machine learning. In: 12th {USENIX} symposium on operating systems design and implementation ({OSDI} 16), pp 265–283
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antiga L et al (2019) Pytorch: an imperative style, high-performance deep learning library. Adv Neural Inf Proces Syst 32:8026–8037
Lu L, Meng X, Mao Z, Karniadakis GE (2019) DeepXDE: a deep learning library for solving differential equations. arXiv preprint arXiv:1907.04502
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv preprint arXiv:1412.6980
Balsa-Canto E, Alonso AA, Banga JR (2008) Computational procedures for optimal experimental design in biological systems. IET Syst Biol 2(4):163–172
Foo J, Sindi S, Karniadakis GE (2009) Multi-element probabilistic collocation for sensitivity analysis in cellular signalling networks. IET Syst Biol 3(4):239–254
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendix
Appendix
1.1 A. Python
Python is the most common language for machine learning due to the plethora of libraries available for free. Learning python is fairly easy due to its popularity, and there are a number of free, high-quality videos and other tutorials on how to use python. We note that common software for installing Python is Anaconda, from which most common libraries have already been installed. The code we provide should remove the majority of the guesswork if solving similar problems to those stated in this tutorial; otherwise, the DeepXDE documentation https://deepxde.readthedocs.io should be of help.
1.2 B. Julia
Julia is another language used for machine learning. It is very similar to Python as far as the syntax is concerned. We recommend using the softwares Atom and the Juno IDE, though Jupiter notebook and similar programs will suffice. There are also a variety of online sources that provide free help for learning this language. If you understand the fundamentals of Python, you should be able to read our provided Julia code and use it for your own situation with only a few minor tweaks that do not involve a heavy amount of coding or even a thorough knowledge of Julia.
Rights and permissions
Copyright information
© 2023 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Daneker, M., Zhang, Z., Karniadakis, G.E., Lu, L. (2023). Systems Biology: Identifiability Analysis and Parameter Identification via Systems-Biology-Informed Neural Networks. In: Nguyen, L.K. (eds) Computational Modeling of Signaling Networks. Methods in Molecular Biology, vol 2634. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3008-2_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3008-2_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3007-5
Online ISBN: 978-1-0716-3008-2
eBook Packages: Springer Protocols