Abstract
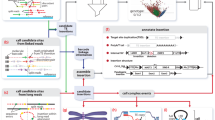
Pangenome graphs are flexible data structures that contain the genetic variation that exists in a population of genomes and describe the sequences of the many possible ensuing haplotypes. Here, we use such a pangenome graph to represent and genotype transposable element (TE) polymorphisms. By combining the transposable element annotation (Alus, L1s, and SVAs) of the human genome reference with novel transposable element insertions observed in two high-quality assemblies (HG002 and HG00733), we show how to create a transposable element pangenome that consists of ~1.2 million reference and 2939 non-reference transposable elements. We then demonstrate this approach by aligning short-read sequencing data and genotyping transposable element deletions and insertions with reasonable specificity and sensitivity (0.85 F1-score).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
van Dijk E, Auger H, Jaszczyszyn Y et al (2014) Ten years of next-generation sequencing technology. Trends Genet 30:418–426
Alser M, Rotman J, Deshpande D et al (2021) Technology dictates algorithms: recent developments in read alignment. Genome Biol 22:249
Mahmoud M, Gobet N, Cruz-Dávalos DI et al (2019) Structural variant calling: the long and the short of it. Genome Biol 20:246
Tattini L, D’Aurizio R, Magi A (2015) Detection of genomic structural variants from next-generation sequencing data. Front Bioeng Biotechnol 3:92
Garg S (2021) Computational methods for chromosome-scale haplotype reconstruction. Genome Biol 22:101
Sherman RM, Forman J, Antonescu V et al (2019) Assembly of a pan-genome from deep sequencing of 910 humans of African descent. Nat Genet 51:30–35
Paten B, Novak AM, Eizenga JM et al (2017) Genome graphs and the evolution of genome inference. Genome Res 27:665–676
Garrison E, Sirén J, Novak AM et al (2018) Variation graph toolkit improves read mapping by representing genetic variation in the reference. Nat Biotechnol 36:875
Martiniano R, Garrison E, Jones ER et al (2020) Removing reference bias and improving indel calling in ancient DNA data analysis by mapping to a sequence variation graph. Genome Biol 21:250
Li H, Feng X, Chu C (2020) The design and construction of reference pangenome graphs with minigraph. Genome Biol 21:265
Hickey G, Heller D, Monlong J et al (2020) Genotyping structural variants in pangenome graphs using the vg toolkit. Genome Biol 21:35
Cheng H, Concepcion GT, Feng X, et al (2020) Haplotype-resolved de novo assembly with phased assembly graphs
Groza C, Bourque G, and Goubert C (2022) Transposable element pangenome code and data, Zenodo
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinforma Oxf Engl 27:2156–2158
Li H (2018) Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34:3094–3100
Heller D, Vingron M (2020) SVIM-asm: structural variant detection from haploid and diploid genome assemblies. Bioinformatics 36:5519–5521
Jeffares DC, Jolly C, Hoti M et al (2017) Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat Commun 8:14061
Groza C, Bourque G, Goubert C (2022) Transposable element pangenome. https://doi.org/10.5281/zenodo.5898621
Jouni S, Jean M, Xian C et al (2021) Pangenomics enables genotyping of known structural variants in 5202 diverse genomes. Science 374:abg8871
Zook JM, Hansen NF, Olson ND et al (2020) A robust benchmark for detection of germline large deletions and insertions. Nat Biotechnol 38:1347–1355
Gardner EJ, Lam VK, Harris DN et al (2017) The Mobile Element Locator Tool (MELT): population-scale mobile element discovery and biology. Genome Res 27:1916–1929
Chen X, Li D (2019) ERVcaller: identifying polymorphic endogenous retrovirus and other transposable element insertions using whole-genome sequencing data. Bioinformatics 35:3913–3922
Chen S, Krusche P, Dolzhenko E et al (2019) Paragraph: a graph-based structural variant genotyper for short-read sequence data. Genome Biol 20:291
Rautiainen M, Marschall T (2020) GraphAligner: rapid and versatile sequence-to-graph alignment. Genome Biol 21:253
Ebler J, Clarke WE, Rausch T et al (2020) Pangenome-based genome inference. bioRxiv:2020.11.11.378133
Ivanov P, Bichsel B, Mustafa H et al (2020) AStarix: fast and optimal sequence-to-graph alignment. In: Schwartz R (ed) Research in computational molecular biology. Springer International Publishing, Cham, pp 104–119
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Groza, C., Bourque, G., Goubert, C. (2023). A Pangenome Approach to Detect and Genotype TE Insertion Polymorphisms. In: Branco, M.R., de Mendoza Soler, A. (eds) Transposable Elements. Methods in Molecular Biology, vol 2607. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2883-6_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2883-6_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2882-9
Online ISBN: 978-1-0716-2883-6
eBook Packages: Springer Protocols