Abstract
Single-cell mRNA sequencing can dissect heterogeneous cell populations as it can identify cell types and cellular states based on their unique transcriptional signatures. We use fluorescence-activated cell sorting (FACS) to isolate individual cultured neurons derived from human-induced pluripotent stem cells (hiPSCs) followed by polyA-based Smart-Seq2 RNA sequencing to analyze the single-cell transcriptional profiles. We provide protocols and guidelines on dissociation, cell selection, and library preparation that can be readily adapted to other cell types or tissue samples.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Picelli S, Björklund ÅK, Faridani OR et al (2013) Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods 10:1096–1098. https://doi.org/10.1038/nmeth.2639
Picelli S (2017) Single-cell RNA-sequencing: the future of genome biology is now. RNA Biol 14:637–650. https://doi.org/10.1080/15476286.2016.1201618
Ding J, Adiconis X, Simmons SK et al (2020) Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol 38:737–746. https://doi.org/10.1038/s41587-020-0465-8
Nichterwitz S, Benitez JA, Hoogstraaten R et al (2018) LCM-Seq: a method for spatial transcriptomic profiling using laser capture microdissection coupled with PolyA-based RNA sequencing. Methods Mol Biol 1649:95–110. https://doi.org/10.1007/978-1-4939-7213-5_6
Picelli S, Faridani OR, Björklund AK et al (2014) Full-length RNA-seq from single cells using smart-seq2. Nat Protoc 9:171–181. https://doi.org/10.1038/nprot.2014.006
Nichterwitz S, Chen G, Aguila Benitez J et al (2016) Laser capture microscopy coupled with smart-seq2 for precise spatial transcriptomic profiling. Nat Commun 7:1–11. https://doi.org/10.1038/ncomms12139
Nijssen J, Aguila J, Hedlund E (2019) Axon-seq for in depth analysis of the RNA content of neuronal processes. Bio-Protocol 9. https://doi.org/10.21769/BioProtoc.3312
Di Giorgio FP, Boulting GL, Bobrowicz S, Eggan KC (2008) Human embryonic stem cell-derived motor neurons are sensitive to the toxic effect of glial cells carrying an ALS-causing mutation. Cell Stem Cell 3:637–648. https://doi.org/10.1016/j.stem.2008.09.017
Kapteyn J, He R, McDowell ET, Gang DR (2010) Incorporation of non-natural nucleotides into template-switching oligonucleotides reduces background and improves cDNA synthesis from very small RNA samples. BMC Genomics 11. https://doi.org/10.1186/1471-2164-11-413
Picelli S, Björklund AK, Reinius B et al (2014) Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res 24:2033–2040. https://doi.org/10.1101/gr.177881.114
Hennig BP, Velten L, Racke I et al (2018) Large-scale low-cost NGS library preparation using a robust Tn5 purification and Tagmentation protocol. G3#58; Genes Genomes Genetics 8:79–89. https://doi.org/10.1534/g3.117.300257
Rohland N, Reich D (2012) Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res 22:939–946. https://doi.org/10.1101/gr.128124.111
Acknowledgments
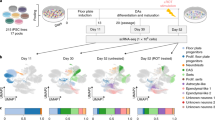
We thank Mattias Karlen for help with Fig. 1. Flow cytometry was performed in the Biomedicum Flowcytometry Core Facility with support of the Karolinska Institutet. We thank Juan Basile and Belinda Pannagel for assisting with flow cytometry. We are grateful for discussions with other members of the Hedlund laboratory. We are also thankful for discussions with and support from members of Rickard Sandberg’s lab. The work in the Hedlund laboratory is supported by grants from the Swedish Research Council (grant number 2020-01049), The Radala Foundation for ALS Research (Switzerland), Ulla-Carin Lindquists Foundation for ALS Research (Ulla-Carin Lindquists stiftelse för ALS forskning), Åhlén-stiftelsen (grant number 213051), Olav Thon Stiftelsen (Norway), The Swedish Brain Foundation (Hjärnfonden) (grant number FO2021-0145) and Parkinsonfonden (grant number 1328/21). C.S. was supported by an Early Postdoc.Mobility fellowship from the Swiss National Science Foundation (P2BEP3_172233).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Schweingruber, C., Nijssen, J., Benitez, J.A., Hedlund, E. (2023). Single-Cell mRNA-Seq of In Vitro-Derived Human Neurons Using Smart-Seq2. In: Song, Q., Tao, Z. (eds) Transcription Factor Regulatory Networks. Methods in Molecular Biology, vol 2594. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2815-7_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2815-7_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2814-0
Online ISBN: 978-1-0716-2815-7
eBook Packages: Springer Protocols