Abstract
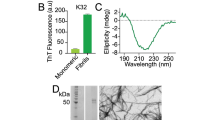
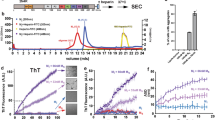
Protein assembly into beta-sheet-rich amyloids is a common phenomenon in neurodegenerative diseases including Alzheimer’s (AD) and Parkinson’s (PD). The proteins implicated in amyloid deposition are often intrinsically disordered proteins (IDPs) and are characterized by not folding into a defined globular conformation. The amyloidogenic properties of IDPs are determined by the presence of short sequence elements, referred to as amyloid motifs, that drive ordered aggregation (Thompson MJ, Sievers SA, Karanicolas J et al. Proc Natl Acad Sci USA 103(11):4074-8, 2006; Goldschmidt L, Teng PK, Riek R et al. Proc Natl Acad Sci USA 107(8):3487-92, 2010]. The microtubule-associated protein tau adopts amyloid assemblies in over 20 different diseases commonly referred to as tauopathies. However, native tau is aggregation-resistant despite encoding at least three amyloid motifs (Chen D, Drombosky KW, Hou Z et al. Nat Commun 10(1):2493, 2019). Recent cryogenic electron microscopy (cryo-EM) structures of tau amyloid fibrils isolated from patient brains showed the involvement of amyloid motifs in the fibril core (Fitzpatrick AWP, Falcon B, He S et al. Nature 547(7662):185–90, 2017; Falcon B, Zhang W, Murzin AG et al. Nature 561(7721):137-40, 2018; Zhang W, Tarutani A, Newell KL et al. Nature 580(7802):283-7, 2020). How does tau change from an aggregation-resistant state to an aggregation-prone state? Consistent with the fibril structures, we hypothesize that tau must change conformation to expose the amyloid motifs that allow self-association into beta-sheet-rich aggregates. This would suggest that the amyloid motifs are likely buried in natively folded tau to prevent self-assembly. We developed an approach that couples cross-linking mass spectrometry (XL-MS) with temperature denaturation to probe the loss of contacts as a proxy to measure protein unfolding with sequence resolution. Using this method, we demonstrated that disease-associated mutations in tau located near an amyloid motif disrupt the protective local structure, promote amyloid motif exposure, and thus lead to aggregation (Chen D, Drombosky KW, Hou Z et al. Nat Commun 10(1):2493, 2019). In this chapter, we describe the detailed protocol for this approach. We anticipate that our protocol can be generalized to other IDPs and will help discover critical structural elements to better understand important biological questions including protein aggregation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Thompson MJ, Sievers SA, Karanicolas J et al (2006) The 3D profile method for identifying fibril-forming segments of proteins. Proc Natl Acad Sci U S A 103(11):4074–4078
Goldschmidt L, Teng PK, Riek R et al (2010) Identifying the amylome, proteins capable of forming amyloid-like fibrils. Proc Natl Acad Sci U S A 107(8):3487–3492
Chen D, Drombosky KW, Hou Z et al (2019) Tau local structure shields an amyloid-forming motif and controls aggregation propensity. Nat Commun 10(1):2493
Fitzpatrick AWP, Falcon B, He S et al (2017) Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547(7662):185–190
Falcon B, Zhang W, Murzin AG et al (2018) Structures of filaments from Pick’s disease reveal a novel tau protein fold. Nature 561(7721):137–140
Zhang W, Tarutani A, Newell KL et al (2020) Novel tau filament fold in corticobasal degeneration. Nature 580(7802):283–287
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6(3):197–208
Nölting B, Golbik R, Neira JL et al (1997) The folding pathway of a protein at high resolution from microseconds to seconds. Proc Natl Acad Sci U S A 94(3):826–830
Holmes BB, Furman JL, Mahan TE et al (2014) Proteopathic tau seeding predicts tauopathy in vivo. Proc Natl Acad Sci 111(41):E4376–E4E85
Furman JL, Holmes BB, Diamond MI (2015) Sensitive detection of proteopathic seeding activity with FRET flow cytometry. J Vis Exp 106:e53205
Rinner O, Seebacher J, Walzthoeni T et al (2008) Identification of cross-linked peptides from large sequence databases. Nat Methods 5(4):315–318
Leitner A, Joachimiak LA, Unverdorben P et al (2014) Chemical cross-linking/mass spectrometry targeting acidic residues in proteins and protein complexes. Proc Natl Acad Sci U S A 111(26):9455–9460
Walzthoeni T, Claassen M, Leitner A et al (2012) False discovery rate estimation for cross-linked peptides identified by mass spectrometry. Nat Methods 9(9):901–903
Leitner A, Walzthoeni T, Aebersold R (2014) Lysine-specific chemical cross-linking of protein complexes and identification of cross-linking sites using LC-MS/MS and the xQuest/xProphet software pipeline. Nat Protocols 9(1):120–137
Acknowledgments
Work in the Joachimiak lab was supported by the Marie Effie Cain Endowed Scholarship, a Chan Zuckerberg Initiative Collaborative Science Award (2018-191983), and a Bright Focus Foundation grant (A2019060). We thank members of the Joachimiak lab for discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Chen, D., Joachimiak, L.A. (2023). Cross-Linking Mass Spectrometry Analysis of Metastable Compact Structures in Intrinsically Disordered Proteins. In: Cieplak, A.S. (eds) Protein Aggregation. Methods in Molecular Biology, vol 2551. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2597-2_13
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2597-2_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2596-5
Online ISBN: 978-1-0716-2597-2
eBook Packages: Springer Protocols