Abstract
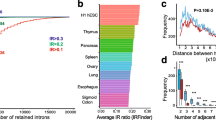
Intron retention (IR) regulates gene expression to control fundamental biological processes like metabolism, differentiation, and cell cycle. Despite a wide variety of genes controlled by IR, few techniques are available to identify regulators of IR in an unbiased manner. Here, we describe a CRISPR knockout screening method that can be applied to uncover regulators of IR. This method uses GFP reporter constructs containing a retained intron from a gene of interest such that GFP signal is regulated by IR in the same fashion as the endogenous gene. The GFP levels are then used as a readout for genome-wide CRISPR screening. We have successfully used this approach to identify novel regulator of IR of the MAT2A transcript and propose that similar screens will be broadly applicable for the identification of novel factors that control IR of specific transcripts.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Middleton R, Gao D, Thomas A, Singh B, Au A, Wong JJ, Bomane A, Cosson B, Eyras E, Rasko JE, Ritchie W (2017) IRFinder: assessing the impact of intron retention on mammalian gene expression. Genome Biol 18(1):51. https://doi.org/10.1186/s13059-017-1184-4
Gordon JM, Phizicky DV, Neugebauer KM (2021) Nuclear mechanisms of gene expression control: pre-mRNA splicing as a life or death decision. Curr Opin Genet Dev 67:67–76. https://doi.org/10.1016/j.gde.2020.11.002
Monteuuis G, Wong JJL, Bailey CG, Schmitz U, Rasko JEJ (2019) The changing paradigm of intron retention: regulation, ramifications and recipes. Nucleic Acids Res 47(22):11497–11513. https://doi.org/10.1093/nar/gkz1068
Jacob AG, Smith CWJ (2017) Intron retention as a component of regulated gene expression programs. Hum Genet 136(9):1043–1057. https://doi.org/10.1007/s00439-017-1791-x
Grabski DF, Broseus L, Kumari B, Rekosh D, Hammarskjold ML, Ritchie W (2021) Intron retention and its impact on gene expression and protein diversity: a review and a practical guide. Wiley Interdiscip Rev RNA 12(1):e1631. https://doi.org/10.1002/wrna.1631
Wegener M, Muller-McNicoll M (2018) Nuclear retention of mRNAs—quality control, gene regulation and human disease. Semin Cell Dev Biol 79:131–142. https://doi.org/10.1016/j.semcdb.2017.11.001
Wong JJ, Au AY, Ritchie W, Rasko JE (2016) Intron retention in mRNA: no longer nonsense: known and putative roles of intron retention in normal and disease biology. BioEssays 38(1):41–49. https://doi.org/10.1002/bies.201500117
Rekosh D, Hammarskjold ML (2018) Intron retention in viruses and cellular genes: detention, border controls and passports. Wiley Interdiscip Rev RNA 9(3):e1470. https://doi.org/10.1002/wrna.1470
Li Y, Bor YC, Misawa Y, Xue Y, Rekosh D, Hammarskjold ML (2006) An intron with a constitutive transport element is retained in a tap messenger RNA. Nature 443(7108):234–237. https://doi.org/10.1038/nature05107
Wong JJ, Ritchie W, Ebner OA, Selbach M, Wong JW, Huang Y, Gao D, Pinello N, Gonzalez M, Baidya K, Thoeng A, Khoo TL, Bailey CG, Holst J, Rasko JE (2013) Orchestrated intron retention regulates normal granulocyte differentiation. Cell 154(3):583–595. https://doi.org/10.1016/j.cell.2013.06.052
Yap K, Lim ZQ, Khandelia P, Friedman B, Makeyev EV (2012) Coordinated regulation of neuronal mRNA steady-state levels through developmentally controlled intron retention. Genes Dev 26(11):1209–1223. https://doi.org/10.1101/gad.188037.112
Braunschweig U, Barbosa-Morais NL, Pan Q, Nachman EN, Alipanahi B, Gonatopoulos-Pournatzis T, Frey B, Irimia M, Blencowe BJ (2014) Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res 24(11):1774–1786. https://doi.org/10.1101/gr.177790.114
Boutz PL, Bhutkar A, Sharp PA (2015) Detained introns are a novel, widespread class of post-transcriptionally spliced introns. Genes Dev 29(1):63–80. https://doi.org/10.1101/gad.247361.114
Pendleton KE, Park SK, Hunter OV, Bresson SM, Conrad NK (2018) Balance between MAT2A intron detention and splicing is determined cotranscriptionally. RNA 24(6):778–786. https://doi.org/10.1261/rna.064899.117
Mauger O, Lemoine F, Scheiffele P (2016) Targeted intron retention and excision for rapid gene regulation in response to neuronal activity. Neuron 92(6):1266–1278. https://doi.org/10.1016/j.neuron.2016.11.032
Ninomiya K, Kataoka N, Hagiwara M (2011) Stress-responsive maturation of Clk1/4 pre-mRNAs promotes phosphorylation of SR splicing factor. J Cell Biol 195(1):27–40. https://doi.org/10.1083/jcb.201107093
Scarborough AM, Flaherty JN, Hunter OV, Liu K, Kumar A, Xing C, Tu BP, Conrad NK (2021) SAM homeostasis is regulated by CFIm-mediated splicing of MAT2A. Elife 10:e64930. https://doi.org/10.7554/eLife.64930
Horlbeck MA, Gilbert LA, Villalta JE, Adamson B, Pak RA, Chen Y, Fields AP, Park CY, Corn JE, Kampmann M, Weissman JS (2016) Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. Elife 5:e19760. https://doi.org/10.7554/eLife.19760
Gilbert LA, Horlbeck MA, Adamson B, Villalta JE, Chen Y, Whitehead EH, Guimaraes C, Panning B, Ploegh HL, Bassik MC, Qi LS, Kampmann M, Weissman JS (2014) Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159(3):647–661. https://doi.org/10.1016/j.cell.2014.09.029
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, Hsu PD, Habib N, Gootenberg JS, Nishimasu H, Nureki O, Zhang F (2015) Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517(7536):583–588. https://doi.org/10.1038/nature14136
Shalem O, Sanjana NE, Zhang F (2015) High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet 16(5):299–311. https://doi.org/10.1038/nrg3899
Moore MJ, Proudfoot NJ (2009) Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 136(4):688–700. https://doi.org/10.1016/j.cell.2009.02.001
Le Hir H, Nott A, Moore MJ (2003) How introns influence and enhance eukaryotic gene expression. Trends Biochem Sci 28(4):215–220. https://doi.org/10.1016/S0968-0004(03)00052-5
Muller-McNicoll M, Neugebauer KM (2013) How cells get the message: dynamic assembly and function of mRNA-protein complexes. Nat Rev Genet 14(4):275–287. https://doi.org/10.1038/nrg3434
Donnelly MLL, Luke G, Mehrotra A, Li X, Hughes LE, Gani D, Ryan MD (2001) Analysis of the aphthovirus 2A/2B polyprotein 'cleavage' mechanism indicates not a proteolytic reaction, but a novel translational effect: a putative ribosomal 'skip'. J Gen Virol 82(Pt 5):1013–1025. https://doi.org/10.1099/0022-1317-82-5-1013
Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, Conrad NK (2017) The U6 snRNA m(6)a methyltransferase METTL16 regulates SAM Synthetase intron retention. Cell 169 (5):824-835:e814. https://doi.org/10.1016/j.cell.2017.05.003
Park SK, Zhou X, Pendleton KE, Hunter OV, Kohler JJ, O'Donnell KA, Conrad NK (2017) A conserved splicing silencer dynamically regulates O-GlcNAc transferase intron retention and O-GlcNAc homeostasis. Cell Rep 20(5):1088–1099. https://doi.org/10.1016/j.celrep.2017.07.017
Roschke AV, Stover K, Tonon G, Schaffer AA, Kirsch IR (2002) Stable karyotypes in epithelial cancer cell lines despite high rates of ongoing structural and numerical chromosomal instability. Neoplasia 4(1):19–31. https://doi.org/10.1038/sj.neo.7900197
Oceguera-Yanez F, Kim SI, Matsumoto T, Tan GW, Xiang L, Hatani T, Kondo T, Ikeya M, Yoshida Y, Inoue H, Woltjen K (2016) Engineering the AAVS1 locus for consistent and scalable transgene expression in human iPSCs and their differentiated derivatives. Methods 101:43–55. https://doi.org/10.1016/j.ymeth.2015.12.012
Sanjana NE, Shalem O, Zhang F (2014) Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 11(8):783–784. https://doi.org/10.1038/nmeth.3047
Hockemeyer D, Soldner F, Beard C, Gao Q, Mitalipova M, DeKelver RC, Katibah GE, Amora R, Boydston EA, Zeitler B, Meng X, Miller JC, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Jaenisch R (2009) Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat Biotechnol 27(9):851–857. https://doi.org/10.1038/nbt.1562
Huang Y, Wimler KM, Carmichael GG (1999) Intronless mRNA transport elements may affect multiple steps of pre-mRNA processing. EMBO J 18(6):1642–1652. https://doi.org/10.1093/emboj/18.6.1642
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, Smith I, Tothova Z, Wilen C, Orchard R, Virgin HW, Listgarten J, Root DE (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34(2):184–191. https://doi.org/10.1038/nbt.3437
Golden RJ, Chen B, Li T, Braun J, Manjunath H, Chen X, Wu J, Schmid V, Chang TC, Kopp F, Ramirez-Martinez A, Tagliabracci VS, Chen ZJ, Xie Y, Mendell JT (2017) An Argonaute phosphorylation cycle promotes microRNA-mediated silencing. Nature 542(7640):197–202. https://doi.org/10.1038/nature21025
Largaespada DA, Collier LS (2008) Transposon-mediated mutagenesis in somatic cells: identification of transposon-genomic DNA junctions. Methods Mol Biol 435. https://doi.org/10.1007/978-1-59745-232-8_7
Richardson RB, Ohlson MB, Eitson JL, Kumar A, McDougal MB, Boys IN, Mar KB, De La Cruz-Rivera PC, Douglas C, Konopka G, Xing C, Schoggins JW (2018) A CRISPR screen identifies IFI6 as an ER-resident interferon effector that blocks flavivirus replication. Nat Microbiol 3(11):1214–1223. https://doi.org/10.1038/s41564-018-0244-1
Shalem O (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87. https://doi.org/10.1126/science.1247005
Broad Institute PCR of sgRNAs for Illumina sequencing protocol. https://media.addgene.org/cms/filer_public/61/16/611619f4-0926-4a07-b5c7-e286a8ecf7f5/broadgpp-sequencing-protocol.pdf
Li W, Xu H, Xiao T, Cong L, Love MI, Zhang F, Irizarry RA, Liu JS, Brown M, Liu XS (2014) MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol 15(12):554. https://doi.org/10.1186/s13059-014-0554-4
Acknowledgments
We would like to thank Dr. John Schoggins and Dr. Joshua Mendell and their lab members for their invaluable guidance in setting up CRISPR screens. This work was supported by the National Institutes of Health R01 GM127311, U01 CA242115, and T32 GM007062.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Scarborough, A.M., Govindan, A., Conrad, N.K. (2022). Genome-Wide CRISPR Screening to Identify Mammalian Factors that Regulate Intron Retention. In: Scheiffele, P., Mauger, O. (eds) Alternative Splicing. Methods in Molecular Biology, vol 2537. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2521-7_16
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2521-7_16
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2520-0
Online ISBN: 978-1-0716-2521-7
eBook Packages: Springer Protocols