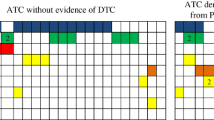
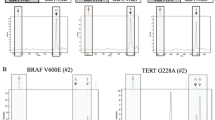
Abstract
The BRAF V600E mutation in papillary thyroid carcinoma is the major mutation in classical subtype of papillary thyroid carcinoma and other cancers. It is the most studied predictor of clinical and pathological characteristics as well as molecular targets for cancer therapy. On the other hand, there is potential for many more forms of activating mutation in BRAF that are not detectable by simple assays to detect V600E, or even simple polymerase chain reaction (PCR)-based sequencing for full-length BRAF. Such activating mutations could arise from larger-scale rearrangements which may apparently leave no sequence change to BRAF while causing increased expression or activation by unusual means, such as gene fusion. Detection of these kinds of changes can take place using a variety of methods, though capture-based sequencing can identify the existence of such forms of mutant BRAF without needing foreknowledge of the loci involved in these kinds of mutation. In this chapter, we detail a method for capture of specific DNA sequences and their amplification to prepare for massively parallel sequencing.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pakneshan S, Salajegheh A, Smith RA, Lam AK (2013) Clinicopathological relevance of BRAF mutations in human cancer. Pathology 45:346–356. https://doi.org/10.1097/PAT.0b013e328360b61d. PMID: 23594689
Ng JY, Lu CT, Lam AK (2019) BRAF mutation: current and future clinical pathological applications in colorectal carcinoma. Histol Histopathol 34:469–477. https://doi.org/10.14670/HH-18-079. Epub 2018 Dec 28. PMID: 30592501
Smith RA, Salajegheh A, Weinstein S, Nassiri M, Lam AK (2011) Correlation between BRAF mutation and the clinicopathological parameters in papillary thyroid carcinoma with particular reference to follicular variant. Hum Pathol 42:500–506. https://doi.org/10.1016/j.humpath.2009.09.023. PMID:21167555
Pillai S, Gopalan V, Smith RA, Lam AK (2015) Diffuse sclerosing variant of papillary thyroid carcinoma--an update of its clinicopathological features and molecular biology. Crit Rev Oncol Hematol 94:64–73. https://doi.org/10.1016/j.critrevonc.2014.12.001. PMID: 25577570
Liu X, Qu S, Liu R, Sheng C, Shi X, Zhu G, Murugan AK, Guan H, Yu H, Wang Y, Sun H, Shan Z, Teng W, Xing M (2014) TERT promoter mutations and their association with BRAF V600E mutation and aggressive clinicopathological characteristics of thyroid cancer. J Clin Endocrinol Metab 99:E1130–E1136. https://doi.org/10.1210/jc.2013-4048. PMID: 24617711
Rahman MA, Salajegheh A, Smith RA, Lam AK (2014) BRAF inhibitor therapy for melanoma, thyroid and colorectal cancers: development of resistance and future prospects. Curr Cancer Drug Targets 14:128–143. https://doi.org/10.2174/1568009614666140121150930. PMID: 24446739
Rahman MA, Salajegheh A, Smith RA, Lam AK (2015) Multiple proliferation-survival signalling pathways are simultaneously active in BRAF V600E mutated thyroid carcinomas. Exp Mol Pathol 99:492–497. https://doi.org/10.1016/j.yexmp.2015.09.006. PMID: 26403329
Rahman MA, Salajegheh A, Smith RA, Lam AK (2015) MicroRNA-126 suppresses proliferation of undifferentiated (BRAF(V600E) and BRAF(WT)) thyroid carcinoma through targeting PIK3R2 gene and repressing PI3K-AKT proliferation-survival signalling pathway. Exp Cell Res 339:342–350. https://doi.org/10.1016/j.yexcr.2015.09.010. PMID: 26384552
Rahman MA, Salajegheh A, Smith RA, Lam AK (2016) Inhibition of BRAF kinase suppresses cellular proliferation, but not enough for complete growth arrest in BRAF V600E mutated papillary and undifferentiated thyroid carcinomas. Endocrine 54:129–138. https://doi.org/10.1007/s12020-016-0985-7. PMID:27179656
Xing M, Alzahrani AS, Carson KA, Viola D, Elisei R, Bendlova B, Yip L, Mian C, Vianello F, Tuttle RM, Robenshtok E, Fagin JA, Puxeddu E, Fugazzola L, Czarniecka A, Jarzab B, O’Neill CJ, Sywak MS, Lam AK, Riesco-Eizaguirre G, Santisteban P, Nakayama H, Tufano RP, Pai SI, Zeiger MA, Westra WH, Clark DP, Clifton-Bligh R, Sidransky D, Ladenson PW, Sykorova V (2013) Association between BRAF V600E mutation and mortality in patients with papillary thyroid cancer. JAMA 309:1493–1501. https://doi.org/10.1001/jama.2013.3190. PMID: 23571588
Xing M, Alzahrani AS, Carson KA, Shong YK, Kim TY, Viola D, Elisei R, Bendlová B, Yip L, Mian C, Vianello F, Tuttle RM, Robenshtok E, Fagin JA, Puxeddu E, Fugazzola L, Czarniecka A, Jarzab B, O’Neill CJ, Sywak MS, Lam AK, Riesco-Eizaguirre G, Santisteban P, Nakayama H, Clifton-Bligh R, Tallini G, Holt EH, Sýkorová V (2015) Association between BRAF V600E mutation and recurrence of papillary thyroid cancer. J Clin Oncol 33:42–50. https://doi.org/10.1200/JCO.2014.56.8253. Epub 2014 Oct 20. PMID: 25332244
Shen X, Zhu G, Liu R, Viola D, Elisei R, Puxeddu E, Fugazzola L, Colombo C, Jarzab B, Czarniecka A, Lam AK, Mian C, Vianello F, Yip L, Riesco-Eizaguirre G, Santisteban P, O’Neill CJ, Sywak MS, Clifton-Bligh R, Bendlova B, Sýkorová V, Xing M (2018) Patient age-associated mortality risk is differentiated by BRAF V600E status in papillary thyroid cancer. J Clin Oncol 36(5):438–445. https://doi.org/10.1200/JCO.2017.74.5497. Epub 2017 Dec 14. PMID: 29240540
Wang F, Zhao S, Shen X, Zhu G, Liu R, Viola D, Elisei R, Puxeddu E, Fugazzola L, Colombo C, Jarzab B, Czarniecka A, Lam AK, Mian C, Vianello F, Yip L, Riesco-Eizaguirre G, Santisteban P, O’Neill CJ, Sywak MS, Clifton-Bligh R, Bendlova B, Sýkorová V, Wang Y, Xing M (2018) BRAF V600E confers male sex disease-specific mortality risk in patients with papillary thyroid cancer. J Clin Oncol 36:2787–2795. https://doi.org/10.1200/JCO.2018.78.5097. PMID: 30070937
Tao Y, Wang F, Shen X, Zhu G, Liu R, Viola D, Elisei R, Puxeddu E, Fugazzola L, Colombo C, Jarzab B, Czarniecka A, Lam AK, Mian C, Vianello F, Yip L, Riesco-Eizaguirre G, Santisteban P, O’Neill CJ, Sywak MS, Clifton-Bligh R, Bendlova B, Sýkorová V, Zhao S, Wang Y, Xing M (2021) BRAF V600E status sharply differentiates lymph node metastasis-associated mortality risk in papillary thyroid cancer. J Clin Endocrinol Metab 106:3228–3238. https://doi.org/10.1210/clinem/dgab286. PMID: 34273152
Huang Y, Qu S, Zhu G, Wang F, Liu R, Shen X, Viola D, Elisei R, Puxeddu E, Fugazzola L, Colombo C, Jarzab B, Czarniecka A, Lam AK, Mian C, Vianello F, Yip L, Riesco-Eizaguirre G, Santisteban P, O’Neill CJ, Sywak MS, Clifton-Bligh R, Bendlova B, Sýkorová V, Xing M (2018) BRAF V600E mutation-assisted risk stratification of solitary intrathyroidal papillary thyroid cancer for precision treatment. J Natl Cancer Inst 110(4):362–370. https://doi.org/10.1093/jnci/djx227. PMID: 29165667
Kim KJ, Kim SG, Tan J, Shen X, Viola D, Elisei R, Puxeddu E, Fugazzola L, Colombo C, Jarzab B, Czarniecka A, Lam AK, Mian C, Vianello F, Yip L, Riesco-Eizaguirre G, Santisteban P, O’Neill CJ, Sywak MS, Clifton-Bligh R, Bendlova B, Sýkorová V, Xing M (2020) BRAF V600E status may facilitate decision-making on active surveillance of low-risk papillary thyroid microcarcinoma. Eur J Cancer 124:161–169. https://doi.org/10.1016/j.ejca.2019.10.017. PMID:31790974
Frisone D, Friedlaender A, Malapelle U, Banna G, Addeo A (2020) A BRAF new world. Crit Rev Oncol Hematol 152:103008. https://doi.org/10.1016/j.critrevonc.2020.103008. PMID: 32485528
Barbitoff YA, Polev DE, Glotov AS, Serebryakova EA, Shcherbakova IV, Kiselev AM, Kostareva AA, Glotov OS, Predeus AV (2020) Systematic dissection of biases in whole-exome and whole-genome sequencing reveals major determinants of coding sequence coverage. Sci Rep 10:2057. https://doi.org/10.1038/s41598-020-59026-y. PMID: 32029882
Jensen MR, Sigsgaard EE, Liu S, Manica A, Bach SS, Hansen MM, Møller PR, Thomsen PF (2021) Genome-scale target capture of mitochondrial and nuclear environmental DNA from water samples. Mol Ecol Resour 21:690–702. https://doi.org/10.1111/1755-0998.13293
Demidov VV, Bukanov NO, Frank-Kamenetskii D (2000) Duplex DNA capture. Curr Issues Mol Biol 2:31–35. PMID: 11464918
St John J, Quinn TW (2008) Rapid capture of DNA targets. BioTechniques 44:259–264. https://doi.org/10.2144/000112633. PMID: 18330355
Zhang X, Garnerone S, Simonetti M, Harbers L, Nicoś M, Mirzazadeh R, Venesio T, Sapino A, Hartman J, Marchiò C, Bienko M, Crosetto N (2019) CUTseq is a versatile method for preparing multiplexed DNA sequencing libraries from low-input samples. Nat Commun 10:4732. https://doi.org/10.1038/s41467-019-12570-2. PMID: 31628304
Sambrook J, Russell DW (2006) Fragmentation of DNA by sonication. CSH Protoc 2006:pdb.prot4538. https://doi.org/10.1101/pdb.prot4538. PMID: 22485919
Telenius H, Carter NP, Bebb CE, Nordenskjöld M, Ponder BA, Tunnacliffe A (1992) Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13:718–725. https://doi.org/10.1016/0888-7543(92)90147-k. PMID: 1639399
Blagodatskikh KA, Kramarov VM, Barsova EV, Garkovenko AV, Shcherbo DS, Shelenkov AA, Ustinova VV, Tokarenko MR, Baker SC, Kramarova TV, Ignatov KB (2017) Improved DOP-PCR (iDOP-PCR): a robust and simple WGA method for efficient amplification of low copy number genomic DNA. PLoS One 12:e0184507. https://doi.org/10.1371/journal.pone.0184507. PMID: 28892497
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Smith, R.A., Lam, A.K. (2022). BRAF Mutations in Papillary Thyroid Carcinoma: A Genomic Approach Using Probe-Based DNA Capture for Next-Generation Sequencing. In: Lam, A.K. (eds) Papillary Thyroid Carcinoma. Methods in Molecular Biology, vol 2534. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2505-7_12
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2505-7_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2504-0
Online ISBN: 978-1-0716-2505-7
eBook Packages: Springer Protocols