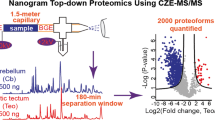
Abstract
Capillary zone electrophoresis (CZE) is a fundamentally simple and highly efficient separation technique based on differences in electrophoretic mobilities of analytes. CZE-mass spectrometry (MS) has become an important analytical tool in top-down proteomics which aims to delineate proteoforms in cells comprehensively, because of the improvement of capillary coatings, sample stacking methods, and CE-MS interfaces. Here, we present a CZE-MS/MS-based top-down proteomics procedure for the characterization of a standard protein mixture and an Escherichia coli (E. coli) cell lysate using linear polyacrylamide-coated capillaries, a dynamic pH junction sample stacking method, a commercialized electro-kinetically pumped sheath flow CE-MS interface and an Orbitrap mass spectrometer. CZE-MS/MS can identify hundreds of proteoforms routinely from the E. coli sample with a 1% proteoform-level false discovery rate (FDR).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Toby TK, Fornelli L, Kelleher NL (2016) Progress in top-down proteomics and the analysis of proteoforms. Annu Rev Anal Chem 9:499–519
Smith LM, Kelleher NL (2018) Proteoforms as the next proteomics currency. Science 359:1106–1107
McCool EN, Lubeckyj RA, Shen X, Chen D, Kou Q, Liu X, Sun L (2018) Deep top-down proteomics using capillary zone electrophoresis-tandem mass spectrometry: identification of 5700 proteoforms from the Eschericia coli proteome. Anal Chem 90:5529–5533
Lubeckyj RA, McCool EN, Shen X, Kou Q, Liu X, Sun L (2017) Single-shot top-down proteomics with capillary zone electrophoresis-electrospray ionization-tandem mass spectrometry for identification of nearly 600 Eschericia coli proteoforms. Anal Chem 89:12059–12067
Shen X, Yang Z, McCool EN, Lubeckyj RA, Chen D, Sun L (2019) Capillary zone electrophoresis-mass spectrometry for top-down proteomics. Trends Anal Chem 120:115644
Lubeckyj RA, Basharat AR, Shen X, Liu X, Sun L (2019) Large-scale qualitative and quantitative top-down using capillary zone electrophoresis-electrospray ionization-tandem mass spectrometry with Nanograms of proteome samples. J Am Soc Mass Spectrom 30:1435–1445
Han X, Wang Y, Aslanian A, Fonslow B, Graczyk B, Davis TN, Yates JR III (2014) In-line separation by capillary electrophoresis prior to analysis by top-down mass spectrometry enables sensitive characterization of protein complexes. J Proteome Res 13:6078–6086
McCool EN, Sun L (2019) Comparing nanoflow reversed-phase liquid chromatography-tandem mass spectrometry and capillary zone electrophoresis-tandem mass spectrometry for top-down proteomics. Se Pu 37:878–886
Li Y, Compton PD, Tran JC, Ntai I, Kelleher NL (2014) Optimizing capillary electrophoresis for top-down proteomics of 30–80 kDa proteins. Proteomics 14:1158–1164
Busnel JM, Schoenmaker B, Ramautar R, Carrasco-Pancorbo A, Ratnayake C, Feitelson JS, Chapman JD, Deelder AM, Mayboroda OA (2010) High capacity capillary electrophoresis-electrospray ionization mass spectrometry: coupling a porous sheathless interface with transient-isotachophoresis. Anal Chem 82:9476–9483
Zhu G, Sun L, Dovichi NJ (2016) Thermally-initiated free radical polymerization for reproducible production of stable linear polyacrylamide coated capillaries, and their application to proteomics analysis using capillary zone electrophoresis-mass spectrometry. Talanta 146:839–843
Haselberg R, de Jong GJ, Somsen GW (2013) Low-flow Sheathless capillary electrophoresis-mass spectrometry for sensitive Glycoform profiling of intact pharmaceutical proteins. Anal Chem 85:2289–2296
Britz-McKibbin P, Chen DD (2000) Selective focusing of catecholamines and weakly acidic compounds by capillary electrophoresis using a dynamic pH junction. Anal Chem 72:1242–1252
McCool EN, Lubeckyj RA, Shen X, Kou Q, Liu X, Sun L (2018) Large-scale top-down proteomics using capillary zone electrophoresis tandem mass spectrometry. J Vis Exp 140:e58644
Zhao Y, Sun L, Zhu G, Dovichi NJ (2016) Coupling capillary zone electrophoresis to a Q Exactive HF mass spectrometer for top-down proteomics: 580 proteoform identifications from yeast. J Proteome Res 15:3679–3685
Chen D, Shen X, Sun L (2017) Capillary zone electrophoresis-mass spectrometry with microliter-scale loading capacity, 140 min separation window and high peak capacity for bottom-up proteomics. Analyst 142:2118–2127
Han X, Wang Y, Aslanian A, Bern M, Lavallée-Adam M, Yates JR III (2014) Sheathless capillary electrophoresis-tandem mass spectrometry for top-down characterization of Pyrococcus furiosus proteins on a proteome scale. Anal Chem 86:11006–11012
Smith RD, Barinaga CJ, Udseth HR (1988) Improved electrospray ionization interface for capillary zone electrophoresis-mass spectrometry. Anal Chem 60:1948–1952
Maxwell EJ, Zhong X, Zhang H, van Zeijl N, Chen DD (2010) Decoupling CE and ESI for a more robust interface with MS. Electrophoresis 31:1130–1137
Moini M (2007) Simplifying CE-MS operation. 2. Interfacing low-flow separation techniques to mass spectrometry using a porous tip. Anal Chem 79:4241–4246
Wojcik R, Dada OO, Sadilek M, Dovichi NJ (2010) Simplified capillary electrophoresis nanospray sheath-flow interface for high efficiency and sensitive peptide analysis. Rapid Commun Mass Spectrom 24:2554–2560
Sun L, Zhu G, Zhang Z, Mou S, Dovichi NJ (2015) Third-generation Electrokinetically pumped sheath-flow Nanospray Interface with improved stability and sensitivity for automated capillary zone electrophoresis-mass spectrometry analysis of complex proteome digests. J Proteome Res 14:2312–2321
McCool EN, Lodge JM, Basharat AR, Liu X, Coon JJ, Sun L (2019) Capillary zone electrophoresis-tandem mass spectrometry with activated ion electron transfer dissociation for large-scale top-down proteomics. J Am Soc Mass Spectrom 30:2470–2479
Kou Q, Xun L, Liu X (2016) TopPIC: a software tool for top-down mass spectrometry-based proteoform identification and characterization. Bioinformatics 32:3495–3497
Donnelly DP, Rawlins CM, Dehart CJ, Fornelli L, Schachner LF, Lin Z et al (2019) Best practices and benchmarks for intact protein analysis for top-down mass spectrometry. Nat Methods 16:587–594
Yang Z, Shen X, Chen D, Sun L (2019) Improved Nanoflow RPLC-CZE-MS/MS system with high peak capacity and sensitivity for Nanogram bottom-up proteomics. J Proteome Res 18:4046–4054
Kessner D, Chambers M, Burke R, Agus D, Mallick P (2008) ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24:2534–2536
Elias JE, Gygi SP (2007) Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat Methods 4:207–214
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
McCool, E.N., Lubeckyj, R.A., Chen, D., Sun, L. (2022). Top-Down Proteomics by Capillary Zone Electrophoresis-Tandem Mass Spectrometry for Large-Scale Characterization of Proteoforms in Complex Samples. In: Neusüß, C., Jooß, K. (eds) Capillary Electrophoresis-Mass Spectrometry . Methods in Molecular Biology, vol 2531. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2493-7_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2493-7_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2492-0
Online ISBN: 978-1-0716-2493-7
eBook Packages: Springer Protocols