Abstract
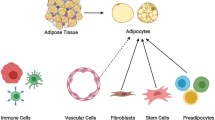
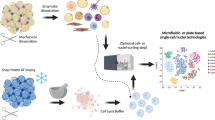
Adipose tissue is highly heterogeneous and plastic. Recent advances in single-cell/nucleus RNA sequencing technology have helped to study the cellular composition and dynamics of adipose tissue. In this protocol, I outline a typical workflow of analyzing single-cell/nucleus transcriptome data. Specifically, I show an example of how cellular populations are estimated and characterized from a single-nucleus RNAseq data set of frozen archived human adipose tissue.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Rosen ED, Spiegelman BM (2014) What we talk about when we talk about fat. Cell 156:20–44
Sun W, von Meyenn F, Peleg-Raibstein D, Wolfrum C (2019) Environmental and nutritional effects regulating adipose tissue function and metabolism across generations. Adv Sci 6:1900275
Hotamisligil GS, Shargill NS, Spiegelman BM (1993) Adipose expression of tumor necrosis factor-alpha: direct role in obesity-linked insulin resistance. Science 259:87–91
Zhang Y et al (1994) Positional cloning of the mouse obese gene and its human homologue. Nature 372:425–432
Scherer PE, Williams S, Fogliano M, Baldini G, Lodish HF (1995) A novel serum protein similar to C1q, produced exclusively in adipocytes (*). J Biol Chem 270:26746–26749
Steppan CM et al (2001) The hormone resistin links obesity to diabetes. Nature 409:307–312
Scheele C, Wolfrum C (2020) Brown adipose crosstalk in tissue plasticity and human metabolism. Endocr Rev 41:53–65
Cannon B, Nedergaard J (2004) Brown adipose tissue: function and physiological significance. Physiol Rev 84:277–359
Jung SM, Sanchez-Gurmaches J, Guertin DA (2019) Brown adipose tissue development and metabolism. In: Pfeifer A, Klingenspor M, Herzig S (eds.) Brown adipose tissue. Springer International Publishing, pp 3–36. https://doi.org/10.1007/164_2018_168
Sun W et al (2018) Cold-induced epigenetic programming of the sperm enhances brown adipose tissue activity in the offspring. Nat Med 24:1372–1383
Sun W et al (2020) snRNA-seq reveals a subpopulation of adipocytes that regulates thermogenesis. Nature 587:98–102
Burl RB et al (2018) Deconstructing Adipogenesis induced by β3-adrenergic receptor activation with single-cell expression profiling. Cell Metab 28:300–309.e4
Schwalie PC et al (2018) A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature 559:103–108
Merrick D et al (2019) Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science 364:eaav2501
Vijay J et al (2020) Single-cell analysis of human adipose tissue identifies depot- and disease-specific cell types. Nat Metab 2:97–109
Crewe C et al (2018) An endothelial-to-adipocyte extracellular vesicle axis governed by metabolic state. Cell 175:695–708.e13
Fischer AW et al (2021) Lysosomal lipoprotein processing in endothelial cells stimulates adipose tissue thermogenic adaptation. Cell Metab 33:547–564.e7
Wirsén C (1964) Adrenergic innervation of adipose tissue examined by fluorescence microscopy. Nature 202:913–913
Feuerer M et al (2009) Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nat Med 15:930–939
Hill DA et al (2018) Distinct macrophage populations direct inflammatory versus physiological changes in adipose tissue. Proc Natl Acad Sci U S A 115:E5096–E5105
Rajbhandari P et al (2019) Single cell analysis reveals immune cell-adipocyte crosstalk regulating the transcription of thermogenic adipocytes. Elife 8:e49501
Shinoda K et al (2015) Genetic and functional characterization of clonally derived adult human brown adipocytes. Nat Med 21:389–394
Xue R et al (2015) Clonal analyses and gene profiling identify genetic biomarkers of the thermogenic potential of human brown and white preadipocytes. Nat Med 21:760–768
Angueira AR et al (2021) Defining the lineage of thermogenic perivascular adipose tissue. Nat Metab 3:469–484
Watcham S, Kucinski I, Gottgens B (2019) New insights into hematopoietic differentiation landscapes from single-cell RNA sequencing. Blood 133:1415–1426
Stubbington MJT, Rozenblatt-Rosen O, Regev A, Teichmann SA (2017) Single-cell transcriptomics to explore the immune system in health and disease. Science 358:58–63
Slyper M et al (2020) A single-cell and single-nucleus RNA-Seq toolbox for fresh and frozen human tumors. Nat Med 26:792–802
Drokhlyansky E et al (2020) The human and mouse enteric nervous system at single-cell resolution. Cell 182:1606–1622.e23
Fleming SJ, Marioni JC, Babadi M (2019) CellBender remove-background: a deep generative model for unsupervised removal of background noise from scRNA-seq datasets. bioRxiv 791699. https://doi.org/10.1101/791699
Griffiths JA, Richard AC, Bach K, Lun ATL, Marioni JC (2018) Detection and removal of barcode swapping in single-cell RNA-seq data. Nat Commun 9:2667
Hao Y et al (2020)Integrated analysis of multimodal single-cell data. bioRxiv 2020.10.12.335331 https://doi.org/10.1101/2020.10.12.335331
Aran D et al (2019) Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20:163–172
Wolock SL, Lopez R, Klein AM (2019) Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst 8:281–291.e9
Browse < ArrayExpress < EMBL-EBI. https://www.ebi.ac.uk/arrayexpress/browse.html
Single-Library Analysis with Cell Ranger -Software -Single Cell Gene Expression -Official 10x Genomics Support. https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/using/count
Lun ATL et al (2019) EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol 20:63
Young MD, Behjati S (2020) SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9:giaa151
Ilicic T et al (2016) Classification of low quality cells from single-cell RNA-seq data. Genome Biol 17:29
Klein AM et al (2015) Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161:1187–1201
Macosko EZ et al (2015) Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161:1202–1214
Cao J et al (2017) Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357:661–667
McGinnis CS, Murrow LM, Gartner ZJ (2019) DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8:329–337.e4
Lun A, Griffiths J, McCarthy D (2021) DropletUtils: utilities for handling single-cell droplet data. (Bioconductor version: Release (3.13)). https://doi.org/10.18129/B9.bioc.DropletUtils
Price AL et al (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Acknowledgements
I am grateful to Christian Wolfrum and Ian Theo Mitchell for editing the manuscript. This work was supported by the Swiss National Science Foundation (P2EZP3 191874).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Sun, W. (2022). Analysis of Single-Cell/Nucleus Transcriptome Data in Adipose Tissue. In: Guertin, D.A., Wolfrum, C. (eds) Brown Adipose Tissue. Methods in Molecular Biology, vol 2448. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2087-8_19
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2087-8_19
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2086-1
Online ISBN: 978-1-0716-2087-8
eBook Packages: Springer Protocols