Abstract
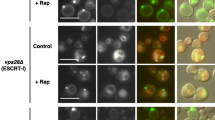
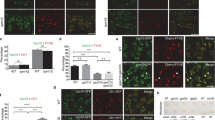
Protein–protein interactions are important for physiology performance. Green fluorescent protein (GFP) is a widely used protein tag to show protein localization in vivo. GFP binding protein (GBP) is a specific domain with high affinity to GFP. A novel technique with GBP fused protein X tagged with red fluorescence protein binding to GFP of GFP fused protein Y to establish a close association for proteins X and Y independently from other proteins has recently been developed. It is found that the interaction and colocalization between Snf7 and Atg17 is impaired in Saccharomyces cerevisiae vps21Δ cells, which are defective in autophagy. In order to determine whether the interaction between Snf7 and Atg17 is important for autophagy, we forced the interaction between Snf7 and Atg17 through GBP–GFP binding. Snf7-GBP-mCherry and/or GFP-Atg17 tagged wild-type and vps21Δ cells were compared for autophagy process under starvation by determining the maturation of proprotein of Ape1 (prApe1). Our results showed that the defective autophagy in vps21Δ cells was significantly suppressed when both Snf7-GBP-mCherry and GFP-Atg17 were installed. Our results indicate that the GBP–GFP nanotrap technique is a powerful tool to restore colocalization/interaction in vivo and the Snf7–Atg17 interaction is important for yeast autophagy.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chen YH, Wang GY, Hao HC et al (2017) Facile manipulation of protein localization in fission yeast through binding of GFP-binding protein to GFP. J Cell Sci 130(5):1003–1015
Chen X, Zaro JL, Shen WC (2013) Fusion protein linkers: property, design and functionality. Adv Drug Deliv Rev 65(10):1357–1369
Rothbauer U, Zolghadr K, Muyldermans S et al (2008) A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins. Mol Cell Proteomics 7(2):282–289
Huh WK, Falvo JV, Gerke LC et al (2003) Global analysis of protein localization in budding yeast. Nature 425(6959):686–691
Chen Y, Zhou F, Zou S et al (2014) A Vps21 endocytic module regulates autophagy. Mol Biol Cell 25(20):3166–3177
Zhou F, Wu Z, Zhao M et al (2019) Autophagosome closure by ESCRT: Vps21/RAB5-regulated ESCRT recruitment via an Atg17-Snf7 interaction. Autophagy 15(9):1653–1654
Gnügge R, Rudolf F (2017) Saccharomyces cerevisiae shuttle vectors. Yeast 34(5):205–221
Kunes S, Botstein D, Fox MS (1985) Transformation of yeast with linearized plasmid DNA. Formation of inverted dimers and recombinant plasmid products. J Mol Biol 184(3):375–387
Gietz RD, Schiestl RH (2007) High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat Protoc 2(1):31–34
Yamasaki A, Watanabe Y, Adachi W et al (2016) Structural basis for receptor-mediated selective autophagy of aminopeptidase I aggregates. Cell Rep 16(1):19–27
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Zhao, M., Liang, Y. (2021). Using GBP Nanotrap to Restore Autophagy in the Rab5/Vps21 Mutant by Forcing Snf7 and Atg17 Interaction. In: Li, G., Segev, N. (eds) Rab GTPases. Methods in Molecular Biology, vol 2293. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1346-7_12
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1346-7_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1345-0
Online ISBN: 978-1-0716-1346-7
eBook Packages: Springer Protocols