Abstract
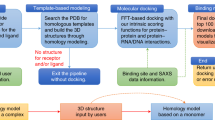
Protein–protein and protein–DNA/RNA interactions are involved in many cellular processes. Therefore, determining their complex structures at the atomic level is valuable to gain insights into these interactions. Because of the technical difficulties and high cost in experimental methods, computational approaches like molecular docking have been developed to predict the structures of macromolecular complexes in the last decades. To automatically integrate the available binding information from the PDB, we have developed HDOCK, a protein–protein/nucleic acid docking web server by combining template-based and free docking. In this chapter, we first briefly introduce our HDOCK server and then give a step-by-step description of docking bovine chymotrypsinogen A against its inhibitor (PDB ID: 1CGI). Two case studies of realistic examples are also discussed. The HDOCK server is freely available at http://hdock.phys.hust.edu.cn/.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Stumpf MP, Thorne T, de Silva E, Stewart R, An HJ, Lappe M, Wiuf C (2008) Estimating the size of the human interactome. Proc Natl Acad Sci U S A 105(19):6959–6964
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28(1):235–242
Chruszcz M, Domagalski M, Osinski T, Wlodawer A, Minor W (2010) Unmet challenges of structural genomics. Curr Opin Struct Biol 20(5):587–597
Rigden DJ, Rigden DJ (2009) From protein structure to function with bioinformatics. Springer, New York
Janin J, Henrick K, Moult J, Eyck LT, Sternberg MJE, Vajda S, Vakser I, Wodak SJ (2003) CAPRI: a critical assessment of PRedicted interactions. Proteins 52(1):2–9
Smith GR, Sternberg MJE (2002) Prediction of protein–protein interactions by docking methods. Curr Opin Struct Biol 12(1):28–35
Huang S-Y (2015) Exploring the potential of global protein–protein docking: an overview and critical assessment of current programs for automatic ab initio docking. Drug Discov Today 20(8):969–977
Ritchie DW (2008) Recent progress and future directions in protein-protein docking. Curr Protein Pept Sci 9(1):1–15
Yan Y, Wen Z, Wang X, Huang SY (2017) Addressing recent docking challenges: a hybrid strategy to integrate template-based and free protein–protein docking. Proteins 85(3):497–512
Vreven T, Hwang H, Pierce BG, Weng Z (2014) Evaluating template-based and template-free protein-protein complex structure prediction. Brief Bioinform 15(2):169–176
Szilagyi A, Zhang Y (2014) Template-based structure modeling of protein–protein interactions. Curr Opin Struct Biol 24:10–23
Kundrotas PJ, Zhu Z, Janin J, Vakser IA (2012) Templates are available to model nearly all complexes of structurally characterized proteins. Proc Natl Acad Sci U S A 109(24):9438–9441
Yan Y, Huang S-Y (2018) Protein–protein docking with improved shape complementarity. In: International conference on intelligent computing. Springer, New York, pp 600–605
Chen R, Weng Z (2003) A novel shape complementarity scoring function for protein–protein docking. Proteins 51(3):397–408
Katchalski-Katzir E, Shariv I, Eisenstein M, Friesem AA, Aflalo C, Vakser IA (1992) Molecular surface recognition: determination of geometric fit between proteins and their ligands by correlation techniques. Proc Natl Acad Sci U S A 89(6):2195–2199
Chen R, Li L, Weng Z (2003) ZDOCK: An initial-stage protein-docking algorithm. Proteins 52(1):80–87
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004) ClusPro: a fully automated algorithm for protein–protein docking. Nucleic Acids Res 32(suppl_2):W96–W99
Kozakov D, Brenke RComeau SR, Vajda S (2010) PIPER: an FFT-based protein docking program with pairwise potentials. Proteins 65(2):392–406
Chen R, Weng Z (2002) Docking unbound proteins using shape complementarity, desolvation, and electrostatics. Proteins 47(3):281–294
Mintseris J, Pierce B, Wiehe K, Anderson R, Chen R, Weng Z (2007) Integrating statistical pair potentials into protein complex prediction. Proteins 69(3):511–520
Huang S-Y, Zou X (2010) MDockPP: a hierarchical approach for protein-protein docking and its application to CAPRI rounds 15-19. Proteins 78(15):3096–3103
Yan Y, Zhang D, Zhou P, Li B, Huang S-Y (2017) HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res 45(W1):W365–W373
Huang SY, Zou X (2008) An iterative knowledge-based scoring function for protein–protein recognition. Proteins 72(2):557–579
Huang S-Y, Zou X (2014) A knowledge-based scoring function for protein-RNA interactions derived from a statistical mechanics-based iterative method. Nucleic Acids Res 42(7):e55–e55
Huang S-Y, Yan C, Grinter SZ, Chang S, Jiang L, Zou X (2013) Inclusion of the orientational entropic effect and low-resolution experimental information for protein-protein docking in critical assessment of PRedicted interactions (CAPRI). Proteins 81(12):2183–2191
Lensink MF, Velankar S, Wodak SJ (2017) Modeling protein–protein and protein–peptide complexes: CAPRI 6th edition. Proteins 85(3):359–377
Imran H, Manikandan PN, Prabhu D, Dharuman V, Jeyakanthan J, Hahn JH (2019) Ultra selective label free electrochemical detection of cancer prognostic p53-antibody at DNA functionalized graphene. Sens Biosensing Res 23:100261
Deep A, Kaundal S, Thakur KG, Tiwari P, Agarwal S, Kidwai S, Singh R (2018) Structural, functional and biological insights into the role of Mycobacterium tuberculosis VapBC11 toxin–antitoxin system: targeting a tRNase to tackle mycobacterial adaptation. Nucleic Acids Res 46(21):11639–11655
Yeh CC, Luo JL, Nhut Phan N, Cheng YC, Chow LP, Tsai MH, Chuang EY, Lai LC (2018) Different effects of long noncoding RNA NDRG1-OT1 fragments on NDRG1 transcription in breast cancer cells under hypoxia. RNA Biol 15(12):1487–1498
Wu H, Wang H, Jiang W, Lian Z (2018) The evolutionary characteristics and structural biology of Gallus toll-like receptor 21. J Mol Recognit 31(6):e2696
Soboleva SE, Zakharova OD, Sedykh SE, Ivanisenko NV, Buneva VN, Nevinsky GA (2019) DNase and RNase activities of fresh cow milk lactoferrin. J Mol Recognit 32:e2777
Hildebrand PW, Rose AS (2015) NGL viewer: a web application for molecular visualization. Nucleic Acids Res 43(W1):W576–W579
Prlić A, Bradley AR, Duarte JM, Rose PW, Rose AS, Valasatava Y (2018) NGL viewer: web-based molecular graphics for large complexes. Bioinformatics 34(21):3755–3758
Hecht HJ, Szardenings M, Collins J, Schomburg D (1991) Three-dimensional structure of the complexes between bovine chymotrypsinogen a and two recombinant variants of human pancreatic secretory trypsin inhibitor (Kazal-type). J Mol Biol 220(3):711–722
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612
Acknowledgments
This work is supported by the National Natural Science Foundation of China (grant No. 31670724), the National Key Research and Development Program of China (grant Nos. 2016YFC1305800 and 2016YFC1305805), and the startup grant of the Huazhong University of Science and Technology (grant No. 3004012104).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Yan, Y., Huang, SY. (2020). Modeling Protein–Protein or Protein–DNA/RNA Complexes Using the HDOCK Webserver. In: Kihara, D. (eds) Protein Structure Prediction. Methods in Molecular Biology, vol 2165. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0708-4_12
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0708-4_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0707-7
Online ISBN: 978-1-0716-0708-4
eBook Packages: Springer Protocols