Abstract
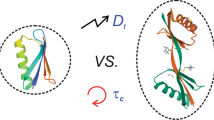
In the disordered state, a protein exhibits a high degree of structural freedom, in both space and time. For an ensemble of disordered or unfolded proteins, this means that the ensemble comprises a high diversity of structures, ranging from compact collapsed states to fully extended polypeptide chains. In addition, each chain is highly dynamic and undergoes conformational changes and local dynamics on both fast and slow timescales. The size properties of disordered proteins are thus best described as ensemble averages. A straightforward measure of the size is the hydrodynamic radius, RH, of the ensemble. Since the disordered state is conformationally fluid, the observed RH does not refer to a particular shape or fold. Instead, it should be interpreted as a measure for the average compaction of the structural ensemble. In addition to characterizing the disordered ensemble itself, RH can be used to, with good precision, monitor changes in the ensemble size properties upon functional interactions of the disordered protein, e.g., dimerization, ligand binding, and folding pathways. Here, we present a step-by-step protocol for diffusion measurements using pulsed field gradient nuclear magnetic resonance (PFG NMR) spectroscopy. We describe how to calibrate the magnetic field gradient and offer different schemes for sample preparation. Finally, we describe how to obtain RH directly from the diffusion coefficient as well as from using an internal standard as a reference.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wright PE, Dyson HJ (1999) Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 293(2):321–331. https://doi.org/10.1006/jmbi.1999.3110
Marsh JA, Forman-Kay JD (2012) Ensemble modeling of protein disordered states: experimental restraint contributions and validation. Proteins 80(2):556–572. https://doi.org/10.1002/prot.23220
Kohn JE, Millett IS, Jacob J et al (2004) Random-coil behavior and the dimensions of chemically unfolded proteins. Proc Natl Acad Sci U S A 101(34):12491–12496. https://doi.org/10.1073/pnas.0403643101
Schneider R, Huang JR, Yao M et al (2012) Towards a robust description of intrinsic protein disorder using nuclear magnetic resonance spectroscopy. Mol BioSyst 8(1):58–68. https://doi.org/10.1039/c1mb05291h
Flory PJ (1953) Principles of polymer chemistry. Cornell Univ. Press, Ithaca, NY
de Gennes PG (1979) Scaling concepts in polymer physics. Cornell Univ. Press, Ithaca, NY
Danielsson J, Jarvet J, Damberg P et al (2002) Translational diffusion measured by PFG-NMR on full length and fragments of the Alzheimer a beta(1-40) peptide. Determination of hydrodynamic radii of random coil peptides of varying length. Magn Reson Chem 40:S89–S97. https://doi.org/10.1002/mrc.1132
Hofmann H, Soranno A, Borgia A et al (2012) Polymer scaling laws of unfolded and intrinsically disordered proteins quantified with single-molecule spectroscopy. Proc Natl Acad Sci U S A 109(40):16155–16160. https://doi.org/10.1073/pnas.1207719109
Wilkins DK, Grimshaw SB, Receveur V et al (1999) Hydrodynamic radii of native and denatured proteins measured by pulse field gradient NMR techniques. Biochemistry 38(50):16424–16431. https://doi.org/10.1021/bi991765q
Ozenne V, Noel JK, Heidarsson PO et al (2014) Exploring the minimally frustrated energy landscape of unfolded ACBP. J Mol Biol 426(3):722–734. https://doi.org/10.1016/j.jmb.2013.10.031
Marsh JA, Forman-Kay JD (2010) Sequence determinants of compaction in intrinsically disordered proteins. Biophys J 98(10):2383–2390. https://doi.org/10.1016/j.bpj.2010.02.006
Grosberg AY, Kuznetsov DV (1992) Quantitative theory of the globule-to-coil transition .4. Comparison of theoretical results with experimental-data. Macromolecules 25(7):1996–2003. https://doi.org/10.1021/ma00033a025
Nygaard M, Kragelund BB, Papaleo E et al (2017) An efficient method for estimating the hydrodynamic radius of disordered protein conformations. Biophys J 113(3):550–557. https://doi.org/10.1016/j.bpj.2017.06.042
Lindorff-Larsen K, Kristjansdottir S, Teilum K et al (2004) Determination of an ensemble of structures representing the denatured state of the bovine acyl-coenzyme a binding protein. J Am Chem Soc 126(10):3291–3299. https://doi.org/10.1021/ja039250g
Penkett CJ, Redfield C, Dodd I et al (1997) NMR analysis of main-chain conformational preferences in an unfolded fibronectin-binding protein. J Mol Biol 274(2):152–159. https://doi.org/10.1006/jmbi.1997.1369
Jones JA, Wilkins DK, Smith LJ et al (1997) Characterisation of protein unfolding by NMR diffusion measurements. J Biomol NMR 10(2):199–203. https://doi.org/10.1023/A:1018304117895
Einstein A (1905) Über die von der molekularkinetischen Theorie der Wärme geforderte Bewegung von in ruhenden Flüssigkeiten suspendierten Teilchen. Ann Phys 322(8):549–560
Stejskal EO, Tanner JE (1965) Spin diffusion measurements: spin echoes in the presence of a time-dependent field gradient. J Chem Phys 42(1):288–292. https://doi.org/10.1063/1.1695690
Roding M, Nyden M (2015) Stejskal-tanner equation for three asymmetrical gradient pulse shapes used in diffusion NMR. Concept Magn Reson A 44(3):133–137. https://doi.org/10.1002/cmr.a.21338
Wu D, Chen AD, Johnson CS (1995) An improved diffusion-ordered spectroscopy experiment incorporating bipolar-gradient pulses. J Magn Reson Ser A 115:260–264. https://doi.org/10.1006/jmra.1995.1176
Chan SHS, Waudby CA, Cassaignau AME et al (2015) Increasing the sensitivity of NMR diffusion measurements by paramagnetic longitudinal relaxation enhancement, with application to ribosome-nascent chain complexes. J Biomol NMR 63(2):151–163. https://doi.org/10.1007/s10858-015-9968-x
Kärger J, Stallmach F (2005) PFG NMR studies of anomalous diffusion. In: Diffusion in condensed matter. Springer, Berlin, Heidelberg. https://doi.org/10.1063/1.1673336
Kuttner YY, Kozer N, Segal E et al (2005) Separating the contribution of translational and rotational diffusion to protein association. J Am Chem Soc 127(43):15138–15144. https://doi.org/10.1021/ja053681c
Dix JA, Verkman AS (2008) Crowding effects on diffusion in solutions and cells. Annu Rev Biophys 37:247–263. https://doi.org/10.1146/annurev.biophys.37.032807.125824
Danielsson J, Liljedahl L, Barany-Wallje E et al (2008) The intrinsically disordered RNR inhibitor Sml1 is a dynamic dimer. Biochemistry 47(50):13428–13437. https://doi.org/10.1021/bi801040b
Danielsson J, Jarvet J, Damberg P et al (2004) Two-site binding of beta-cyclodextrin to the Alzheimer Abeta(1-40) peptide measured with combined PFG-NMR diffusion and induced chemical shifts. Biochemistry 43(20):6261–6269. https://doi.org/10.1021/bi036254p
Abelein A, Graslund A, Danielsson J (2015) Zinc as chaperone-mimicking agent for retardation of amyloid beta peptide fibril formation. Proc Natl Acad Sci U S A 112(17):5407–5412. https://doi.org/10.1073/pnas.1421961112
Hrovat MI, Wade CG (1981) Nmr pulsed-gradient diffusion measurements .1. Spin-Echo stability and gradient calibration. J Magn Reson 44(1):62–75. https://doi.org/10.1016/0022-2364(81)90189-X
Damberg P, Jarvet J, Graslund A (2001) Accurate measurement of translational diffusion coefficients: a practical method to account for nonlinear gradients. J Magn Reson 148(2):343–348. https://doi.org/10.1006/jmre.2000.2260
Findeisen M, Brand T, Berger S (2007) A 1H-NMR thermometer suitable for cryoprobes. Magn Reson Chem 45(2):175–178. https://doi.org/10.1002/mrc.1941
Loening NM, Keeler J (1999) Measurement of convection and temperature profiles in liquid samples. J Magn Reson 139(2):334–341. https://doi.org/10.1006/jmre.1999.1777
Tanner JE (1970) Use of the stimulated Echo in NMR diffusion studies. J Chem Phys 52(5):2523–2526. https://doi.org/10.1063/1.1673336
Danielsson J, Andersson A, Jarvet J et al (2006) 15N relaxation study of the amyloid beta-peptide: structural propensities and persistence length. Magn Reson Chem 44:S114–S121. https://doi.org/10.1002/mrc.1814
Cho CH, Urquidi J, Singh S et al (1999) Thermal offset viscosities of liquid H2O, D2O, and T2O. J Phys Chem B 103(11):1991–1994. https://doi.org/10.1021/jp9842953
Ming LJ, Valentine JS (2014) Insights into SOD1-linked amyotrophic lateral sclerosis from NMR studies of Ni(2+)- and other metal-ion-substituted wild-type copper-zinc superoxide dismutases. J Biol Inorg Chem 19(4-5):647–657. https://doi.org/10.1007/s00775-014-1126-5
Dong A, Prestrelski SJ, Allison SD et al (1995) Infrared spectroscopic studies of lyophilization- and temperature-induced protein aggregation. J Pharm Sci 84(4):415–424. https://doi.org/10.1002/jps.2600840407
Wang CK, Northfield SE, Swedberg JE et al (2014) Translational diffusion of cyclic peptides measured using pulsed-field gradient NMR. J Phys Chem B 118(38):11129–11136. https://doi.org/10.1021/jp506678f
Danielsson J, Kurnik M, Lang L et al (2011) Cutting off functional loops from homodimeric enzyme superoxide dismutase 1 (SOD1) leaves monomeric beta-barrels. J Biol Chem 286(38):33070–33083. https://doi.org/10.1074/jbc.M111.251223
Acknowledgments
This work was funded by the Knut and Alice Wallenberg Foundation (2017-0041), the Swedish Research Council (2017-01517), and the Magnus Bergvall Foundation (2017-02228).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Leeb, S., Danielsson, J. (2020). Obtaining Hydrodynamic Radii of Intrinsically Disordered Protein Ensembles by Pulsed Field Gradient NMR Measurements. In: Kragelund, B.B., Skriver, K. (eds) Intrinsically Disordered Proteins. Methods in Molecular Biology, vol 2141. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0524-0_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0524-0_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0523-3
Online ISBN: 978-1-0716-0524-0
eBook Packages: Springer Protocols