Abstract
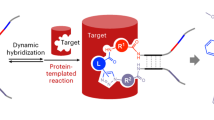
DNA-encoded library technologies have emerged as a powerful platform to rapidly screen for binders to a protein of interest. These technologies are underpinned by the ability to encode a rich diversity of small molecules. While large libraries are accessible by cycles of mix and split synthesis, libraries based on single chemistries tend to be redundant. Furthermore, the quality of libraries generally decreases with the number of synthetic transformations performed in its synthesis. An alternative approach is to use hybridization to program the combinatorial assembly of fragment pairs onto a library of DNA templates. A broad molecular diversity is more easily sampled since it arises from the pairing of diverse fragments. Upon identification of productive fragment pairs, a focused library covalently linking the fragments is prepared. This focused library includes linker of different length and geometry and offers the opportunity to enrich the selected fragment set with close neighbors. Herein we describe detailed protocols to covalently link diverse fragments and screen fragment-based libraries using commercially available microarray platform.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Erlanson DA, Fesik SW, Hubbard RE, Jahnke W, Jhoti H (2016) Twenty years on: the impact of fragments on drug discovery. Nat Rev Drug Discov 15:605–619. https://doi.org/10.1038/nrd.2016.109
Neri D, Lerner RA (2018) DNA-encoded chemical libraries: a selection system based on endowing organic compounds with amplifiable information. Annu Rev Biochem 87(87):479–502
Shi BB, Zhou Y, Huang YR, Zhang JF, Li XY (2017) Recent advances on the encoding and selection methods of DNA-encoded chemical library. Bioorg Med Chem Lett 27:361–369
Nielsen PE, Egholm M, Berg RH, Buchardt O (1991) Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 254:1497–1500
Egholm M, Buchardt O, Christensen L, Behrens C, Freier SM, Driver DA, Berg RH, Kim SK, Norden B, Nielsen PE (1993) PNA hybridizes to complementary oligonucleotides obeying the Watson-Crick hydrogen-bonding rules. Nature 365:566–568
Nielsen PE (1999) Peptide nucleic acid. A molecule with two identities. Acc Chem Res 32:624–630
Zambaldo C, Barluenga S, Winssinger N (2015) PNA-encoded chemical libraries. Curr Opin Chem Biol 26:8–15
Barluenga S, Winssinger N (2015) PNA as a biosupramolecular tag for programmable assemblies and reactions. Acc Chem Res 48:1319–1331
Daguer JP, Ciobanu M, Alvarez S, Barluenga S, Winssinger N (2011) DNA-templated combinatorial assembly of small molecule fragments amenable to selection/amplification cycles. Chem Sci 2:625–632. https://doi.org/10.1039/c0sc00574f
Pouchain D, Diaz-Mochon JJ, Bialy L, Bradley M (2007) A 10,000 member PNA-encoded peptide library for profiling tyrosine kinases. ACS Chem Biol 2:810–818. https://doi.org/10.1021/cb700199k
Debaene F, Mejias L, Harris JL, Winssinger N (2004) Synthesis of a PNA-encoded cysteine protease inhibitor library. Tetrahedron 60:8677–8690. https://doi.org/10.1016/j.tet.2004.05.107
Debaene F, Da Silva J, Pianowski Z, Duran F, Winssinger N (2007) Expanding the scope of PNA-encoded libraries: divergent synthesis of libraries targeting cysteine, serine and metalloproteases as well as tyrosine phosphatases. Tetrahedron 63:6577–6586
Daguer JP, Zambaldo C, Abegg D, Barluenga S, Tallant C, Muller S, Adibekian A, Winssinger N (2015) Identification of covalent bromodomain binders through DNA display of small molecules. Angew Chem Int Ed Engl 54:6057–6061. https://doi.org/10.1002/anie.201412276
Zambaldo C, Daguer JP, Saarbach J, Barluenga S, Winssinger N (2016) Screening for covalent inhibitors using DNA-display of small molecule libraries functionalized with cysteine reactive moieties. Med Chem Commun 7:1340–1351. https://doi.org/10.1039/c6md00242k
Ciobanu M, Huang KT, Daguer JP, Barluenga S, Chaloin O, Schaeffer E, Mueller CG, Mitchell DA, Winssinger N (2011) Selection of a synthetic glycan oligomer from a library of DNA-templated fragments against DC-SIGN and inhibition of HIV gp120 binding to dendritic cells. Chem Commun 47:9321–9323. https://doi.org/10.1039/c1cc13213j
Huang K-T, Gorska K, Alvarez S, Barluenga S, Winssinger N (2011) Combinatorial self-assembly of glycan fragments into microarrays. Chembiochem 12:56–60. https://doi.org/10.1002/cbic.201000567
Novoa A, Eierhoff T, Topin J, Varrot A, Barluenga S, Imberty A, Romer W, Winssinger N (2014) A LecA ligand identified from a galactoside-conjugate array inhibits host cell invasion by Pseudomonas aeruginosa. Angew Chem Int Ed Engl 53:8885–8889. https://doi.org/10.1002/anie.201402831
Novoa A, Machida T, Barluenga S, Imberty A, Winssinger N (2014) PNA-encoded synthesis (PES) of a 10000-member hetero-glycoconjugate library and microarray analysis of diverse lectins. Chembiochem 15:2058–2065. https://doi.org/10.1002/cbic.201402280
Daguer JP, Zambaldo C, Ciobanu M, Morieux P, Barluenga S, Winssinger N (2015) DNA display of fragment pairs as a tool for the discovery of novel biologically active small molecules. Chem Sci 6:739–744. https://doi.org/10.1039/c4sc01654h
Barluenga S, Zambaldo C, Ioannidou HA, Ciobanu M, Morieux P, Daguer JP, Winssinger N (2016) Novel PTP1B inhibitors identified by DNA display of fragment pairs. Bioorg Med Chem Lett 26:1080–1085. https://doi.org/10.1016/j.bmcl.2015.11.102
Levitzki A (2003) Protein kinase inhibitors as a therapeutic modality. Acc Chem Res 36:462–469. https://doi.org/10.1021/ar0201207
Ferguson FM, Gray NS (2018) Kinase inhibitors: the road ahead. Nat Rev Drug Discov 17:353–376. https://doi.org/10.1038/nrd.2018.21
Garg R, Benedetti LG, Abera MB, Wang H, Abba M, Kazanietz MG (2014) Protein kinase C and cancer: what we know and what we do not. Oncogene 33:5225–5237. https://doi.org/10.1038/onc.2013.524
Chouikhi D, Ciobanu M, Zambaldo C, Duplan V, Barluenga S, Winssinger N (2012) Expanding the scope of PNA-encoded synthesis (PES): Mtt-protected PNA fully orthogonal to Fmoc chemistry and a broad array of robust diversity-generating reactions. Chem Eur J 18:12698–12704. https://doi.org/10.1002/chem.201201337
Winssinger N, Damoiseaux R, Tully DC, Geierstanger BH, Burdick K, Harris JL (2004) PNA-encoded protease substrate microarrays. Chem Biol 11:1351–1360. https://doi.org/10.1016/j.chembiol.2004.07.015
Harris JL, Winssinger N (2005) PNA encoding (PNA = peptide nucleic acid): from solution-based libraries to organized microarrays. Chem Eur J 11:6792–6801. https://doi.org/10.1002/chem.200500305
Urbina HD, Debaene F, Jost B, Bole-Feysot C, Mason DE, Kuzmic P, Harris JL, Winssinger N (2006) Self-assembled small-molecule microarrays for protease screening and profiling. Chembiochem 7:1790–1797. https://doi.org/10.1002/cbic.200600242
Gorska K, Huang KT, Chaloin O, Winssinger N (2009) DNA-templated homo- and heterodimerization of peptide nucleic acid encoded oligosaccharides that mimick the carbohydrate epitope of HIV. Angew Chem Int Ed Engl 48:7695–7700. https://doi.org/10.1002/anie.200903328
Saarbach J, Masi D, Zambaldo C, Winssinger N (2017) Facile access to modified and functionalized PNAs through Ugi-based solid phase oligomerization. Bioorg Med Chem 25:5171–5177
Svensen N, Diaz-Mochon JJ, Bradley M (2011) Decoding a PNA encoded peptide library by PCR: the discovery of new cell surface receptor ligands. Chem Biol 18:1284–1289. https://doi.org/10.1016/j.chembiol.2011.07.017
Li G, Zheng WL, Liu Y, Li XY (2015) Novel encoding methods for DNA-templated chemical libraries. Curr Opin Chem Biol 26:25–33
Melkko S, Scheuermann J, Dumelin CE, Neri D (2004) Encoded self-assembling chemical libraries. Nat Biotechnol 22:568–574
Acknowledgments
We gratefully acknowledge the funding agencies which have supported this work (Swiss National Science Foundation, NCCR Chemical Biology) and the collaborators that have contributed to the development of PNA-encoded technologies.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Saarbach, J., Barluenga, S., Winssinger, N. (2020). PNA-Encoded Synthesis (PES) and DNA Display of Small Molecule Libraries. In: Nielsen, P. (eds) Peptide Nucleic Acids. Methods in Molecular Biology, vol 2105. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0243-0_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0243-0_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0242-3
Online ISBN: 978-1-0716-0243-0
eBook Packages: Springer Protocols