Abstract
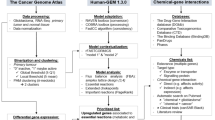
The drug development pipeline has stalled because of the difficulty in identifying new drug targets while minimizing off-target effects. Computational methods, such as the use of metabolic network reconstructions, may provide a cost-effective platform to test new hypotheses for drug targets and prevent off-target effects. Here, we summarize available methods to identify drug targets and off-target effects using either reaction-centric, gene-centric, or metabolite-centric approaches with genome-scale metabolic network reconstructions.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mougin F, Auber D, Bourqui R et al (2018) Visualizing omics and clinical data: Which challenges for dealing with their variety? Methods 132:3–18
Schuster D, Laggner C, Langer T (2005) Why drugs fail—a study on side effects in new chemical entities. Curr Pharm Des 11:3545–3559
Edwards IR, Aronson JK (2000) Adverse drug reactions: definitions, diagnosis, and management. Lancet 356:1255–1259
Sawada R, Iwata M, Tabei Y et al (2018) Predicting inhibitory and activatory drug targets by chemically and genetically perturbed transcriptome signatures. Sci Rep 8:156
Suthers PF, Zomorrodi A, Maranas CD (2009) Genome-scale gene/reaction essentiality and synthetic lethality analysis. Mol Syst Biol 5:301
Pey J, San José-Eneriz E, Ochoa MC et al (2017) In-silico gene essentiality analysis of polyamine biosynthesis reveals APRT as a potential target in cancer. Sci Rep 7:14358
Bordbar A, Lewis NE, Schellenberger J et al (2010) Insight into human alveolar macrophage and M. tuberculosis interactions via metabolic reconstructions. Mol Syst Biol 6:422
Orth JD, Palsson BØ, Fleming RMT (2010) Reconstruction and use of microbial metabolic networks: the core Escherichia coli metabolic model as an educational guide. EcoSal Plus 4:PMID: 26443778
Oberhardt MA, Puchalka J, Fryer KE et al (2008) Genome-scale metabolic network analysis of the opportunistic pathogen Pseudomonas aeruginosa PAO1. J Bacteriol 190:2790–2803
Förster J, Famili I, Fu P et al (2003) Genome-scale reconstruction of the Saccharomyces cerevisiae metabolic network. Genome Res 13:244–253
Thiele I, Swainston N, Fleming RMT et al (2013) A community-driven global reconstruction of human metabolism. Nat Biotechnol 31:419–425
Mardinoglu A, Agren R, Kampf C et al (2014) Genome-scale metabolic modelling of hepatocytes reveals serine deficiency in patients with non-alcoholic fatty liver disease. Nat Commun 5:3083
Blais EM, Rawls KD, Dougherty BV et al (2017) Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions. Nat Commun 8:14250
Yizhak K, Chaneton B, Gottlieb E et al (2015) Modeling cancer metabolism on a genome scale. Mol Syst Biol 11:817
Ghaffari P, Mardinoglu A, Asplund A et al (2015) Identifying anti-growth factors for human cancer cell lines through genome-scale metabolic modeling. Sci Rep 5:8183
Chang RL, Xie L, Xie L et al (2010) Drug off-target effects predicted using structural analysis in the context of a metabolic network model. PLoS Comput Biol 6:e1000938
Zielinski DC, Filipp FV, Bordbar A et al (2015) Pharmacogenomic and clinical data link non-pharmacokinetic metabolic dysregulation to drug side effect pathogenesis. Nat Commun 6:7101
Shaked I, Oberhardt MA, Atias N et al (2016) Metabolic network prediction of drug side effects. Cell Syst 2:209–213
Heirendt L, Arreckx S, Pfau T et al (2019) Creation and analysis of biochemical constraint-based models using the COBRA Toolbox v.3.0. Nat Protoc 14:639–702
Folger O, Jerby L, Frezza C et al (2011) Predicting selective drug targets in cancer through metabolic networks. Mol Syst Biol 7:501
Rawls KD, Dougherty BV, Blais EM et al (2019) A simplified metabolic network reconstruction to promote understanding and development of flux balance analysis tools. Comput Biol Med 105:64–71
Ma H, Sorokin A, Mazein A et al (2007) The Edinburgh human metabolic network reconstruction and its functional analysis. Mol Syst Biol 3:135
Swainston N, Smallbone K, Hefzi H et al (2016) Recon 2.2: from reconstruction to model of human metabolism. Metabolomics 12:109
Brunk E, Sahoo S, Zielinski DC et al (2018) Recon3D enables a three-dimensional view of gene variation in human metabolism. Nat Biotechnol 36:272–281
Robaina Estévez S, Nikoloski Z (2014) Generalized framework for context-specific metabolic model extraction methods. Front Plant Sci 5:491
Blazier AS, Papin JA (2012) Integration of expression data in genome-scale metabolic network reconstructions. Front Physiol 3:299
Uhlen M, Hallstro m BM, Lindskog C et al (2016) Transcriptomics resources of human tissues and organs. Mol Syst Biol 12:862–862
Schultz A, Qutub AA (2016) Reconstruction of tissue-specific metabolic networks using CORDA. PLoS Comput Biol 12:e1004808
Agren R, Bordel S, Mardinoglu A et al (2012) Reconstruction of genome-scale active metabolic networks for 69 human cell types and 16 cancer types using INIT. PLoS Comput Biol 8:e1002518
Zhang A-D, Dai S-X, Huang J-F Reconstruction and analysis of human kidney-specific metabolic network based on omics data. https://www.hindawi.com/journals/bmri/2013/187509/
Sohrabi-Jahromi S, Marashi S-A, Kalantari S (2016) A kidney-specific genome-scale metabolic network model for analyzing focal segmental glomerulosclerosis. Mamm Genome 27:158–167
Karlstädt A, Fliegner D, Kararigas G et al (2012) CardioNet: a human metabolic network suited for the study of cardiomyocyte metabolism. BMC Syst Biol 6:114
Zhao Y, Huang J (2011) Reconstruction and analysis of human heart-specific metabolic network based on transcriptome and proteome data. Biochem Biophys Res Commun 415:450–454
Bordbar A, Mo ML, Nakayasu ES et al (2012) Model-driven multi-omic data analysis elucidates metabolic immunomodulators of macrophage activation. Mol Syst Biol 8:558
Agren R, Mardinoglu A, Asplund A et al (2014) Identification of anticancer drugs for hepatocellular carcinoma through personalized genome-scale metabolic modeling. Mol Syst Biol 10:721
Thiele I, Palsson BØ (2010) A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat Protoc 5:93–121
Yizhak K, Gabay O, Cohen H et al (2013) Model-based identification of drug targets that revert disrupted metabolism and its application to ageing. Nat Commun 4:2632
Nogiec C, Burkart A, Dreyfuss JM et al (2015) Metabolic modeling of muscle metabolism identifies key reactions linked to insulin resistance phenotypes. Mol Metab 4:151–163
Yizhak K, Le Devedec SE, Rogkoti VM et al (2014) A computational study of the Warburg effect identifies metabolic targets inhibiting cancer migration. Mol Syst Biol 10:744–744
Stempler S, Yizhak K, Ruppin E (2014) Integrating transcriptomics with metabolic modeling predicts biomarkers and drug targets for Alzheimer’s disease. PLoS One 9:e105383
Wishart DS, Feunang YD, Guo AC, et al (2018) DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 46:D1074–D1082
Kuhn M, Campillos M, Letunic I et al (2010) A side effect resource to capture phenotypic effects of drugs. Mol Syst Biol 6:343
Kuhn M, Letunic I, Jensen LJ et al (2016) The SIDER database of drugs and side effects. Nucleic Acids Res 44:D1075–D1079
Rienksma RA, Suarez-Diez M, Spina L et al (2014) Systems-level modeling of mycobacterial metabolism for the identification of new (multi-)drug targets. Semin Immunol 26:610–622
Guarente L (1993) Synthetic enhancement in gene interaction: a genetic tool come of age. Trends Genet 9:362–366
Jerby L, Shlomi T, Ruppin E (2010) Computational reconstruction of tissue-specific metabolic models: application to human liver metabolism. Mol Syst Biol 6:401
Raškevičius V, Mikalayeva V, Antanavičiūtė I et al (2018) Genome scale metabolic models as tools for drug design and personalized medicine. PLoS One 13:e0190636
Longley DB, Harkin DP, Johnston PG (2003) 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 3:330–338
Mehrmohamadi M, Jeong SH, Locasale JW (2017) Molecular features that predict the response to antimetabolite chemotherapies. Cancer Metab 5:8
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Rawls, K., Dougherty, B.V., Papin, J. (2020). Metabolic Network Reconstructions to Predict Drug Targets and Off-Target Effects. In: Nagrath, D. (eds) Metabolic Flux Analysis in Eukaryotic Cells. Methods in Molecular Biology, vol 2088. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0159-4_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0159-4_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0158-7
Online ISBN: 978-1-0716-0159-4
eBook Packages: Springer Protocols