Abstract
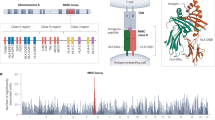
The only strategy to select individuals at increased risk for type 1 diabetes for primary prevention trials is through genetic risk assessment. While genome-wide association studies have identified more than 40 loci associated with type 1 diabetes, the single most important genetic determinants lie within the human leucocyte antigen gene family on chromosome 6.
In this chapter we describe a protocol for a straightforward, cheap strategy to determine HLA class II mediated risk of type 1 diabetes. This method has proved robust for genotyping whole-genome-amplified DNA as well as DNA extracted directly from human tissues.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Keywords:
1 Introduction
HLA class II presents self-peptides to the immune system and autoimmunity results from impaired tolerance to self-antigens. A fundamental role for HLA class II in susceptibility to type 1 diabetes (T1D) is therefore not surprising. An association between HLA and T1D was originally demonstrated in the 1970s [1, 2] with several major steps forward in the ensuing decades when the importance of HLA class II alleles was described [3] the relationship between age at onset of diabetes and HLA class II-mediated risk [4–6], and the hierarchy of HLA class II-mediated risk in type 1 diabetes [7].
The method described here has been adapted from the original “Phototyping” method for discrimination of HLA genotype by Bunce and colleagues [8]. It allows identification of all alleles of HLA DRB1 facilitating identification of homozygosity and heterozygosity as well as selected alleles from HLA DQA1 and DQB1. Overall the highest risk diplotype is HLA DRB1*03-DQB1*0201/DRB1*04-DQB1*0302.
2 Materials
2.1 DNA
DNA extracted from human tissues: This includes tissues extracted from fixed tissues using appropriate extraction kits and whole-genome-amplified DNA (Note 1 ).
2.2 Reagents and Supplies
PCR reaction
-
1.
Oligonucleotide primers specific for HLA DRB1, DQA1, and DQB1 as described in Tables 1 and 2 (Fig. 1). Make up to a standardized concentration of 100 pmol/μl and store in aliquots at −20 °C. A control primer set (any robust set will suffice) must be added to each allele specific primer set.
Table 1 HLA DRB1 primer sequences. DR2 has been split to DR15 and 16, DR5 to DR11 and 12, and DR6 to DR13 and 14 Table 2 HLA DQB1 primer sequences Fig. 1 A typical example of an HLA DQB1 genotyping result. In this case the sample is positive for HLA DQB1*02 (*0201) and DQB1*0302. Note that in lane 2 the target HLA DQB1*0201 allele is the larger band observed. The control band set used in this experiment is Human growth hormone; F5′CCAGCTCAAGGATCCCAA and R5′ CACCCATTACCCAAGAGCTTA
-
2.
Taq polymerase and PCR Buffer, MgCl2 (usually supplied together). Go Taq and buffer from Promega are used in the method described (Note 2 ).
-
3.
dNTPs commercially available
-
4.
96-Well PCR plates
-
5.
96-Well thermocycler
-
6.
10× TBE buffer: 108 g Tris and 55 g boric acid in 800 ml dH2O. Add 40 ml 0.5 M Na2EDTA (pH 8.0). Adjust volume to 1 l. Store at room temperature.
-
7.
Agarose gel electrophoresis tank, gel-forming tray, and powerpack
Agarose Gel Electrophoresis
-
1.
2 % agarose gel: 4 g Ultrapure Agarose, 200 ml 1× TBE buffer
-
2.
Ethidium bromide/Midori green or alternative
3 Methods
3.1 Set Up PCR Reaction
-
1.
Label a 96-well plate for the number of reactions to be carried out; for instance a full HLA DRB1 type requires 14 reactions per sample.
-
2.
Following the worksheet provided in Table 3; briefly add the DNA (Note 1 ) to the appropriate well. Ensure that positive and negative (dH2O) controls are included.
Table 3 Work sheet for typing HLA class II DRB1 and DQB1 -
3.
Make up a “cocktail” of all other reagents for the required number of reactions
-
4.
Vortex to mix
-
5.
Add the appropriate volume of “cocktail” to each well.
-
6.
Seal plate
-
7.
Spin down in an appropriate centrifuge or gently tap the plate to ensure that all reagents are mixed.
3.2 Thermal Cycler Program
Select the following touchdown PCR program on the thermal cycler
Steps | Temperature (°C) | Time (sec.) | Action |
|---|---|---|---|
1 cycle | 96 | 60 | Denaturation |
5 cycles | 96 | 25 | Denaturation |
70 | 50 | Annealing | |
72 | 45 | Extension | |
21 cycles | 96 | 25 | Denaturation |
65 | 50 | Annealing | |
72 | 45 | Extension | |
4 cycles | 96 | 25 | Denaturation |
55 | 60 | Annealing | |
72 | 120 | Extension | |
Hold | 4 | Specify time |
3.3 Agarose Gel Electrophoresis
-
1.
Weigh out 4 g of agarose into a conical flask. Add 200 mL of 1× TBE, and swirl to mix (Notes 3 and 4 ).
-
2.
Microwave for about 2 min to completely dissolve the agarose. Stop after 45 s to give agarose mix a swirl taking precautions not to allow agarose to boil over causing a burn.
-
3.
Ensure the agarose is completely dissolved. If not replace in the microwave until dissolved monitoring carefully.
-
4.
Cool the agarose by swirling the conical flask under a running cold tap until the gel is about 60 °C.
-
5.
Add 2 μL of ethidium bromide (10 mg/mL) or alternative DNA stain for instance Midori green and swirl to mix (Note 5 ).
-
6.
Pour the gel into a pre-prepared gel tank with appropriate sized gel combs on a level surface.
-
7.
Allow to set for 1 h.
-
8.
Pour 1× TBE buffer into the gel tank to submerge the gel (Note 6 ).
-
9.
Remove combs ensuring that wells are well formed
-
10.
Pipette 10 μl of PCR product into the gel and run at 100–115 V for approximately an hour (Note 7 ).
-
11.
Visualise the gel on an appropriate gel documentation system.
4 Notes
-
1.
This method works well on DNA from tissues or whole-genome-amplified DNA but ensure that whole-genome-amplified DNA is appropriately diluted.
-
2.
Go Taq (Promega) has been used in this protocol but other enzymes should also work. This enzyme comes with a coloured buffer with sufficient density that a gel loading dye is not required. An alternative to this addition of sucrose creosol red which acts as a density agent for gel electrophoresis but can be added to the PCR reaction without any negative affect. This saves time adding a loading buffer for electrophoresis. The recipe is as follows: 100 mM Creosol Red dye: dissolve 0.4 g creosol red in 10 ml ddH2O, vortex to mix. Before setting up the PCR reaction a loading dye can be generated as follows: 60 % Sucrose/1 mM Creosol Red (50 ml): Dissolve 30 g sucrose in 50 ml autoclaved deionized H2O, add 500 μl of 100 mM creosol red, vortex to mix. Aliquot 1 ml into 1.5 ml Eppendorfs, and store at −20 °C. Add 1.5 μl during the setup of a 10 μl reaction
-
3.
Use a large container, as long as it fits in the microwave, because the agarose boils over easily.
-
4.
The volume of gel can be scaled up depending on the number of samples to be analyzed and the gel equipment available.
-
5.
Ethidium bromide is mutagenic and should be handled with caution. Contaminated tips can be disposed of in a dedicated ethidium bromide waste container. Alternative DNA stains are increasingly used.
-
6.
The gel must be run in the same buffer as used to make up the gel.
-
7.
DNA is negatively charged and will run towards the anode.
References
Nerup J, Platz P, Andersen OO et al (1974) HL-A antigens and diabetes mellitus. Lancet 2:864–866
Cudworth AG, Woodrow JC (1975) Evidence for HL-A-linked genes in ‘juvenile’ diabetes mellitus. Br Med J 3:133–135
Todd JA, Bell JI, McDevitt HO (1987) HLA-DQ beta gene contributes to susceptibility and resistance to insulin-dependent diabetes mellitus. Nature 329:599–604
Caillat-Zucman S, Garchon HJ, Timsit J, Assan R, Boitard C, Djilali-Saiah I, Bougnères P, Bach JF (1992) Age-dependent HLA genetic heterogeneity of type 1 insulin-dependent diabetes mellitus. J Clin Invest 90:2242–2250
Gillespie KM, Gale EA, Bingley PJ (2002) High familial risk and genetic susceptibility in early-onset childhood diabetes. Diabetes 51:210–214
Gillespie KM, Aitken RJ, Wilson I et al (2014) Early onset of diabetes in the proband is the major determinant of risk in HLA DR3-DQ2/DR4-DQ8 siblings. Diabetes 63:1041–1047
Lambert AP, Gillespie KM, Thomson G et al (2004) Absolute risk of childhood-onset type 1 diabetes defined by human leukocyte antigen class II genotype: a population-based study in the United Kingdom. J Clin Endocrinol Metab 89:4037–4043
Bunce M, O’Neill CM, Barnardo MC et al (1995) Phototyping: comprehensive DNA typing for HLA-A, B, C, DRB1, DRB3, DRB4, DRB5 & DQB1 by PCR with 144 primer mixes utilizing sequence-specific primers (PCR-SSP). Tissue Antigens 46:355–367
Acknowledgement
This work was supported by Diabetes UK grants to KMG.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Aitken, R.J., Mortimer, G.L., Gillespie, K.M. (2015). Type 1 Diabetes High-Risk HLA Class II Determination by Polymerase Chain Reaction Sequence-Specific Primers. In: Gillespie, K. (eds) Type-1 Diabetes. Methods in Molecular Biology, vol 1433. Humana Press, New York, NY. https://doi.org/10.1007/7651_2015_307
Download citation
DOI: https://doi.org/10.1007/7651_2015_307
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3641-0
Online ISBN: 978-1-4939-3643-4
eBook Packages: Springer Protocols