Abstract
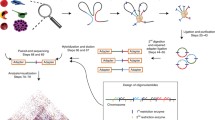
The chromosome conformation capture (3C) method has been invaluable in studying chromatin interactions in a population of cells at a resolution surpassing that of light microscopy, for example in the detection of functional contacts between enhancers and promoters. Recent developments in sequencing-based chromosomal contact mapping (Hi-C, 5C and 4C-Seq) have allowed researchers to interrogate pairwise chromatin interactions on a wider scale, shedding light on the three-dimensional organization of chromosomes. These methods present significant technical and bioinformatic challenges to consider at the start of the project. Here, we describe two alternative methods for Hi-C, depending on the size of the genome, and discuss the major computational approaches to convert the raw sequencing data into meaningful models of how genomes are organized.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lanctot C, Cheutin T, Cremer M et al (2007) Dynamic genome architecture in the nuclear space: regulation of gene expression in three dimensions. Nat Rev Genet 8:104–115
Dekker J, Rippe K, Dekker M et al (2002) Capturing chromosome conformation. Science 295:1306–1311
Thierry Forne’s chapter in this same book…
Dekker J, Marti-Renom MA, Mirny LA (2013) Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat Rev Genet 14:390–403
Lieberman-Aiden E, van Berkum NL, Williams L et al (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326:289–293
Dixon JR, Selvaraj S, Yue F et al (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485:376–380
Sexton T, Yaffe E, Kenigsberg E et al (2012) Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148:458–472
Seitan VC, Faure AJ, Zhan Y et al (2013) Cohesin-based chromatin interactions enable regulated gene expression within preexisting architectural compartments. Genome Res 23:2066–2077
Sofueva S, Yaffe E, Chan WC et al (2013) Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J 32:3119–3129
Zuin J, Dixon JR, van der Reijden MI et al (2014) Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc Natl Acad Sci U S A 111:996–1001
van de Werken HJ, Landan G, Holwerda SJ et al (2012) Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat Methods 9:969–972
Yaffe E, Tanay A (2011) Probabilistic modeling of Hi-C contact maps eliminates systematic biases to characterize global chromosomal architecture. Nat Genet 43:1059–1065
Schoenfelder S, Sexton T, Chakalova L et al (2010) Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat Genet 42:53–61
Raab JR, Chiu J, Zhu J et al (2012) Human tRNA genes function as chromatin insulators. EMBO J 31:330–350
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Vietri Rudan, M., Hadjur, S., Sexton, T. (2016). Detecting Spatial Chromatin Organization by Chromosome Conformation Capture II: Genome-Wide Profiling by Hi-C. In: Haggarty, P., Harrison, K. (eds) Population Epigenetics. Methods in Molecular Biology, vol 1589. Humana Press, New York, NY. https://doi.org/10.1007/7651_2015_261
Download citation
DOI: https://doi.org/10.1007/7651_2015_261
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6901-2
Online ISBN: 978-1-4939-6903-6
eBook Packages: Springer Protocols